Antibody data
- Antibody Data
- Antigen structure
- References [7]
- Comments [0]
- Validations
- Flow cytometry [1]
- Other assay [2]
Submit
Validation data
Reference
Comment
Report error
- Product number
- 25-9488-41 - Provider product page

- Provider
- Invitrogen Antibodies
- Product name
- KLRG1 Monoclonal Antibody (13F12F2), PE-Cyanine7, eBioscience™
- Antibody type
- Monoclonal
- Antigen
- Other
- Description
- Description: This 13F12F2 monoclonal antibody reacts with human killer cell lectin-like receptor subfamily G, member 1 (KLRG1), a type II transmembrane inhibitor receptor of the C-type lectin superfamily that contains an ITIM domain. This inhibitory receptor is expressed on subsets of gamma-delta T cells, NK (CD56dim), CD8+ and CD4+ T Cells. KLRG1 is expressed primarily by cells with an effector/memory phenotype that are short-lived, but capable of immediate effector cell function. Cadherin-E, -N, and -R are ligands for KLRG1. Cadherin/KLRG1 interaction inhibits cytolytic activity and proliferation. The percentage of KLRG1 positive cells can vary considerably, depending on antigen experience.
- Antibody clone number
- 13F12F2
- Concentration
- 5 µL/Test
Submitted references Isolation of tumor-resident CD8(+) T cells from human lung tumors.
PD-1-Expressing SARS-CoV-2-Specific CD8(+) T Cells Are Not Exhausted, but Functional in Patients with COVID-19.
IL-1β, IL-23, and TGF-β drive plasticity of human ILC2s towards IL-17-producing ILCs in nasal inflammation.
Diverse roles for T-bet in the effector responses required for resistance to infection.
Autophagy is a critical regulator of memory CD8(+) T cell formation.
Heterologous immunity triggered by a single, latent virus in Mus musculus: combined costimulation- and adhesion- blockade decrease rejection.
Breakdown in repression of IFN-γ mRNA leads to accumulation of self-reactive effector CD8+ T cells.
Corgnac S, Lecluse Y, Mami-Chouaib F
STAR protocols 2021 Mar 19;2(1):100267
STAR protocols 2021 Mar 19;2(1):100267
PD-1-Expressing SARS-CoV-2-Specific CD8(+) T Cells Are Not Exhausted, but Functional in Patients with COVID-19.
Rha MS, Jeong HW, Ko JH, Choi SJ, Seo IH, Lee JS, Sa M, Kim AR, Joo EJ, Ahn JY, Kim JH, Song KH, Kim ES, Oh DH, Ahn MY, Choi HK, Jeon JH, Choi JP, Kim HB, Kim YK, Park SH, Choi WS, Choi JY, Peck KR, Shin EC
Immunity 2021 Jan 12;54(1):44-52.e3
Immunity 2021 Jan 12;54(1):44-52.e3
IL-1β, IL-23, and TGF-β drive plasticity of human ILC2s towards IL-17-producing ILCs in nasal inflammation.
Golebski K, Ros XR, Nagasawa M, van Tol S, Heesters BA, Aglmous H, Kradolfer CMA, Shikhagaie MM, Seys S, Hellings PW, van Drunen CM, Fokkens WJ, Spits H, Bal SM
Nature communications 2019 May 14;10(1):2162
Nature communications 2019 May 14;10(1):2162
Diverse roles for T-bet in the effector responses required for resistance to infection.
Harms Pritchard G, Hall AO, Christian DA, Wagage S, Fang Q, Muallem G, John B, Glatman Zaretsky A, Dunn WG, Perrigoue J, Reiner SL, Hunter CA
Journal of immunology (Baltimore, Md. : 1950) 2015 Feb 1;194(3):1131-40
Journal of immunology (Baltimore, Md. : 1950) 2015 Feb 1;194(3):1131-40
Autophagy is a critical regulator of memory CD8(+) T cell formation.
Puleston DJ, Zhang H, Powell TJ, Lipina E, Sims S, Panse I, Watson AS, Cerundolo V, Townsend AR, Klenerman P, Simon AK
eLife 2014 Nov 11;3
eLife 2014 Nov 11;3
Heterologous immunity triggered by a single, latent virus in Mus musculus: combined costimulation- and adhesion- blockade decrease rejection.
Beus JM, Hashmi SS, Selvaraj SA, Duan D, Stempora LL, Monday SA, Cheeseman JA, Hamby KM, Speck SH, Larsen CP, Kirk AD, Kean LS
PloS one 2013;8(8):e71221
PloS one 2013;8(8):e71221
Breakdown in repression of IFN-γ mRNA leads to accumulation of self-reactive effector CD8+ T cells.
Chang PP, Lee SK, Hu X, Davey G, Duan G, Cho JH, Karupiah G, Sprent J, Heath WR, Bertram EM, Vinuesa CG
Journal of immunology (Baltimore, Md. : 1950) 2012 Jul 15;189(2):701-10
Journal of immunology (Baltimore, Md. : 1950) 2012 Jul 15;189(2):701-10
No comments: Submit comment
Supportive validation
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
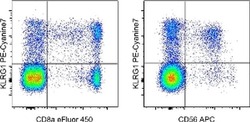
- LEFT: Staining of normal human peripheral blood cells with Anti-Human CD8a eFluor® 450 (Product # 48-0087-42) and Anti-Human KLRG1 PE-Cyanine7. Cells in the lymphocyte gate were used for analysis. RIGHT: Staining of normal human peripheral blood cells with Anti-Human CD56 (NCAM) APC (Product # 17-0567-42) and Anti-Human KLRG1 PE-Cyanine7. Cells in the lymphocyte gate were used for analysis.
Supportive validation
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
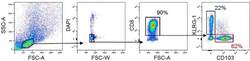

- Experimental details
- Figure 2 Gating strategy for sorting CD8 + CD103 + and CD8 + KLRG1 + cells by FACS
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
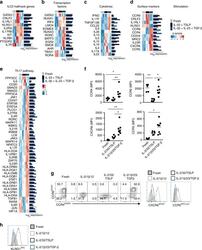
- Fig. 6 ILC2 stimulations result in unique expression patterns. a Heatmap of genes related to ILC2 signature. b Heatmap of genes related to ILC-related transcription factors. c Heatmap of genes related to ILC-related cytokines. d Heatmap of genes encoding significantly differentially expressed surface markers d . Both z-scores (row-normalized) and absolute expression values are shown. e Heatmap of genes significantly different expressed in ILC2s upon stimulation with IL-2, IL-1beta, IL-23, and TGF-beta as compared with ILC2s cultured with IL-2, IL-33, and TSLP that are related to the Th17 KEGG pathway. f Quantification of expression of CCR4, CCR6, CCR5 and CXCR6 on freshly isolated blood ILC2s, and upon culture for 5 days with IL-2, IL-33 and TSLP or IL-2, IL-1beta, IL-23, and TGF-beta ( n = 5-8). g Representative flow cytometric analysis of CCR4, CCR6, CCR5, and CXCR6 upon culture as in f . h Representative flow cytometric analysis of KLRG1 expression upon culture as in f . Data are presented as individual values with mean of two to four independent experiments ( f ). * p < 0.05, ** p < 0.01 as determined by one way ANOVA. Source data
 Explore
Explore Validate
Validate Learn
Learn Flow cytometry
Flow cytometry