25-9326-42
antibody from Invitrogen Antibodies
Targeting: EPCAM
17-1A, 323/A3, CD326, CO-17A, EGP-2, EGP34, EGP40, Ep-CAM, ESA, GA733-2, HEA125, KS1/4, KSA, Ly74, M4S1, MH99, MIC18, MK-1, MOC31, TACST-1, TACSTD1, TROP1
Antibody data
- Antibody Data
- Antigen structure
- References [20]
- Comments [0]
- Validations
- Flow cytometry [1]
- Other assay [13]
Submit
Validation data
Reference
Comment
Report error
- Product number
- 25-9326-42 - Provider product page

- Provider
- Invitrogen Antibodies
- Product name
- CD326 (EpCAM) Monoclonal Antibody (1B7), PE-Cyanine7, eBioscience™
- Antibody type
- Monoclonal
- Antigen
- Other
- Description
- Description: EpCAM (Epithelial cell adhesion molecule, CD326, KSA, TROP1) is a 40 kD cell-surface adhesion molecule participating in homophilic, calcium-independent cell-cell interactions. EpCAM is a type-I transmembrane protein, and is expressed primarily on the basolateral surface of most epithelia. Although normal epithelia express low levels of EpCAM, increased expression has been correlated with increased proliferation and progression to a mesenchymal phenotype. EpCAM has also been used as a diagnostic marker on circulating metastatic carcinoma cells, while cancer cells of non-epithelial origin do not express EpCAM.
- Antibody clone number
- 1B7
- Concentration
- 5 µL/Test
Submitted references The sodium/glucose cotransporters as potential therapeutic targets for CF lung diseases revealed by human lung organoid swelling assay.
EMP3 negatively modulates breast cancer cell DNA replication, DNA damage repair, and stem-like properties.
A molecular cell atlas of the human lung from single-cell RNA sequencing.
Chemotherapeutic Stress Influences Epithelial-Mesenchymal Transition and Stemness in Cancer Stem Cells of Triple-Negative Breast Cancer.
FZD5 contributes to TNBC proliferation, DNA damage repair and stemness.
Profiling human breast epithelial cells using single cell RNA sequencing identifies cell diversity.
The hypoxic tumor microenvironment in vivo selects the cancer stem cell fate of breast cancer cells.
Characterization of a switchable chimeric antigen receptor platform in a pre-clinical solid tumor model.
Detection of Circulating Tumour Cells in Urothelial Cancers and Clinical Correlations: Comparison of Two Methods.
Evidence of In Vitro Preservation of Human Nephrogenesis at the Single-Cell Level.
TGF-β1 secreted by Tregs in lymph nodes promotes breast cancer malignancy via up-regulation of IL-17RB.
The isolation and characterization of CTC subsets related to breast cancer dormancy.
NDY1/KDM2B functions as a master regulator of polycomb complexes and controls self-renewal of breast cancer stem cells.
GATA3 suppresses metastasis and modulates the tumour microenvironment by regulating microRNA-29b expression.
Stem-like epithelial cells are concentrated in the distal end of the fallopian tube: a site for injury and serous cancer initiation.
Slow-cycling therapy-resistant cancer cells.
Exploration of target molecules for prostate cancer gene therapy.
Expression of epithelial cell adhesion molecule in carcinoma cells present in blood and primary and metastatic tumors.
Evidence for a role of the epithelial glycoprotein 40 (Ep-CAM) in epithelial cell-cell adhesion.
Ep-CAM: a human epithelial antigen is a homophilic cell-cell adhesion molecule.
Hirai H, Liang X, Sun Y, Zhang Y, Zhang J, Chen YE, Mou H, Zhao Y, Xu J
Molecular therapy. Methods & clinical development 2022 Mar 10;24:11-19
Molecular therapy. Methods & clinical development 2022 Mar 10;24:11-19
EMP3 negatively modulates breast cancer cell DNA replication, DNA damage repair, and stem-like properties.
Zhou K, Sun Y, Dong D, Zhao C, Wang W
Cell death & disease 2021 Sep 12;12(9):844
Cell death & disease 2021 Sep 12;12(9):844
A molecular cell atlas of the human lung from single-cell RNA sequencing.
Travaglini KJ, Nabhan AN, Penland L, Sinha R, Gillich A, Sit RV, Chang S, Conley SD, Mori Y, Seita J, Berry GJ, Shrager JB, Metzger RJ, Kuo CS, Neff N, Weissman IL, Quake SR, Krasnow MA
Nature 2020 Nov;587(7835):619-625
Nature 2020 Nov;587(7835):619-625
Chemotherapeutic Stress Influences Epithelial-Mesenchymal Transition and Stemness in Cancer Stem Cells of Triple-Negative Breast Cancer.
Li X, Strietz J, Bleilevens A, Stickeler E, Maurer J
International journal of molecular sciences 2020 Jan 8;21(2)
International journal of molecular sciences 2020 Jan 8;21(2)
FZD5 contributes to TNBC proliferation, DNA damage repair and stemness.
Sun Y, Wang Z, Na L, Dong D, Wang W, Zhao C
Cell death & disease 2020 Dec 12;11(12):1060
Cell death & disease 2020 Dec 12;11(12):1060
Profiling human breast epithelial cells using single cell RNA sequencing identifies cell diversity.
Nguyen QH, Pervolarakis N, Blake K, Ma D, Davis RT, James N, Phung AT, Willey E, Kumar R, Jabart E, Driver I, Rock J, Goga A, Khan SA, Lawson DA, Werb Z, Kessenbrock K
Nature communications 2018 May 23;9(1):2028
Nature communications 2018 May 23;9(1):2028
The hypoxic tumor microenvironment in vivo selects the cancer stem cell fate of breast cancer cells.
Kim H, Lin Q, Glazer PM, Yun Z
Breast cancer research : BCR 2018 Mar 6;20(1):16
Breast cancer research : BCR 2018 Mar 6;20(1):16
Characterization of a switchable chimeric antigen receptor platform in a pre-clinical solid tumor model.
Pishali Bejestani E, Cartellieri M, Bergmann R, Ehninger A, Loff S, Kramer M, Spehr J, Dietrich A, Feldmann A, Albert S, Wermke M, Baumann M, Krause M, Bornhäuser M, Ehninger G, Bachmann M, von Bonin M
Oncoimmunology 2017;6(10):e1342909
Oncoimmunology 2017;6(10):e1342909
Detection of Circulating Tumour Cells in Urothelial Cancers and Clinical Correlations: Comparison of Two Methods.
Fina E, Necchi A, Bottelli S, Reduzzi C, Pizzamiglio S, Iacona C, Daidone MG, Verderio P, Cappelletti V
Disease markers 2017;2017:3414910
Disease markers 2017;2017:3414910
Evidence of In Vitro Preservation of Human Nephrogenesis at the Single-Cell Level.
Pode-Shakked N, Gershon R, Tam G, Omer D, Gnatek Y, Kanter I, Oriel S, Katz G, Harari-Steinberg O, Kalisky T, Dekel B
Stem cell reports 2017 Jul 11;9(1):279-291
Stem cell reports 2017 Jul 11;9(1):279-291
TGF-β1 secreted by Tregs in lymph nodes promotes breast cancer malignancy via up-regulation of IL-17RB.
Huang SC, Wei PC, Hwang-Verslues WW, Kuo WH, Jeng YM, Hu CM, Shew JY, Huang CS, Chang KJ, Lee EY, Lee WH
EMBO molecular medicine 2017 Dec;9(12):1660-1680
EMBO molecular medicine 2017 Dec;9(12):1660-1680
The isolation and characterization of CTC subsets related to breast cancer dormancy.
Vishnoi M, Peddibhotla S, Yin W, T Scamardo A, George GC, Hong DS, Marchetti D
Scientific reports 2015 Dec 3;5:17533
Scientific reports 2015 Dec 3;5:17533
NDY1/KDM2B functions as a master regulator of polycomb complexes and controls self-renewal of breast cancer stem cells.
Kottakis F, Foltopoulou P, Sanidas I, Keller P, Wronski A, Dake BT, Ezell SA, Shen Z, Naber SP, Hinds PW, McNiel E, Kuperwasser C, Tsichlis PN
Cancer research 2014 Jul 15;74(14):3935-46
Cancer research 2014 Jul 15;74(14):3935-46
GATA3 suppresses metastasis and modulates the tumour microenvironment by regulating microRNA-29b expression.
Chou J, Lin JH, Brenot A, Kim JW, Provot S, Werb Z
Nature cell biology 2013 Feb;15(2):201-13
Nature cell biology 2013 Feb;15(2):201-13
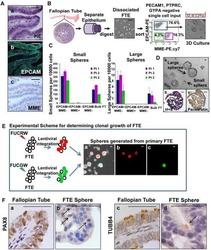
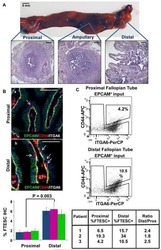
Stem-like epithelial cells are concentrated in the distal end of the fallopian tube: a site for injury and serous cancer initiation.
Paik DY, Janzen DM, Schafenacker AM, Velasco VS, Shung MS, Cheng D, Huang J, Witte ON, Memarzadeh S
Stem cells (Dayton, Ohio) 2012 Nov;30(11):2487-97
Stem cells (Dayton, Ohio) 2012 Nov;30(11):2487-97
Slow-cycling therapy-resistant cancer cells.
Moore N, Houghton J, Lyle S
Stem cells and development 2012 Jul 1;21(10):1822-30
Stem cells and development 2012 Jul 1;21(10):1822-30
Exploration of target molecules for prostate cancer gene therapy.
Suzuki K, Nakamura K, Kato K, Hamada H, Tsukamoto T
The Prostate 2007 Aug 1;67(11):1163-73
The Prostate 2007 Aug 1;67(11):1163-73
Expression of epithelial cell adhesion molecule in carcinoma cells present in blood and primary and metastatic tumors.
Rao CG, Chianese D, Doyle GV, Miller MC, Russell T, Sanders RA Jr, Terstappen LW
International journal of oncology 2005 Jul;27(1):49-57
International journal of oncology 2005 Jul;27(1):49-57
Evidence for a role of the epithelial glycoprotein 40 (Ep-CAM) in epithelial cell-cell adhesion.
Litvinov SV, Bakker HA, Gourevitch MM, Velders MP, Warnaar SO
Cell adhesion and communication 1994 Oct;2(5):417-28
Cell adhesion and communication 1994 Oct;2(5):417-28
Ep-CAM: a human epithelial antigen is a homophilic cell-cell adhesion molecule.
Litvinov SV, Velders MP, Bakker HA, Fleuren GJ, Warnaar SO
The Journal of cell biology 1994 Apr;125(2):437-46
The Journal of cell biology 1994 Apr;125(2):437-46
No comments: Submit comment
Supportive validation
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
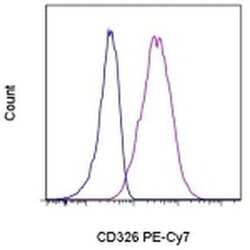
- Staining of A549 cells with Mouse IgG1 K Isotype Control PE-Cyanine7 (Product # 25-4714-80) (blue histogram) or Anti-Human CD326 (EpCAM) PE-Cyanine7 (purple histogram). Total viable cells, as determined by Fixable Viability Dye eFluor® 450 (Product # 65-0863-14), were used for analysis.
Supportive validation
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
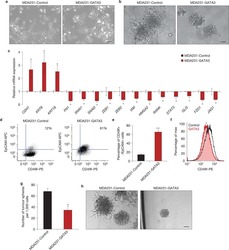
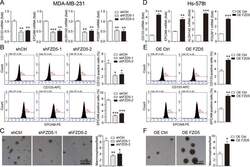
- Fig. 5 FZD5 maintains stem cell-like properties. A CD133, EPCAM, ALDH1A2, and POU5F1 expression in MDA-MB-231 cells was detected by real-time PCR. B Fractions of CD133-positive and EPCAM-positive MDA-MB-231 cells were detected by flow cytometry. C Mammosphere formation of MDA-MB-231 cells was shown. Scale bar: 500 mum, mean +- SD, n = 3. * P < 0.05, ** P < 0.01, *** P < 0.001, vs. shCtrl. D CD133, EPCAM, ALDH1A2, and POU5F1 expression in Hs-578t cells was detected by real-time PCR. E Fractions of CD133-positive and E PCAM-positive Hs-578t cells were detected by flow cytometry. F Mammosphere formation of Hs-578t cells was shown. Scale bar: 500 mum, mean +- SD, n = 3. * P < 0.05, ** P < 0.01, *** P < 0.001, vs. OE Ctrl.
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
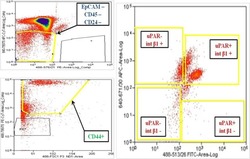
- Figure 1 Multiparametric flow cytometry of PBMCs capturing uPAR/int beta1 CTC subsets. Breast cancer PBMCs were first sorted applying gating parameters to select for DAPI - (4', 6-diamidino-2-phenylindole)/EpCAM - /CD45 - /CD44 + /CD24 - cells. Cells were then subsequently sorted to obtain uPAR/int beta1 subsets containing combinatorial expression of these markers. Antibodies used for flow cytometry and cell sorting were: anti-human CD45-APC-Cy7 (Biolegend, cat # 304015, 1:50 dilution), mouse anti-human EpCAM-PE CD326 (eBiosciences, cat # 12-9326-71, 1:40 dilution), anti-human CD24-PE ML5 (Biolegend, cat # 311106, 1:20 dilution), anti-human CD44-PE-Cy7 IM7 (Biolegend, cat # 103030, 1:20 dilution), mouse anti-human uPAR (CD87)-FITC (AbD Serotec cat # MCA2506FT, 1:10 dilution), anti-human int beta1 (CD29)-ApC TS2/16 (Biolegend, cat # 3030008, 1:50 dilution). Cells were confirmed to be CTCs by performing RT-PCR, immunoflurescence and genotyping arrays. Representative images are shown.
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
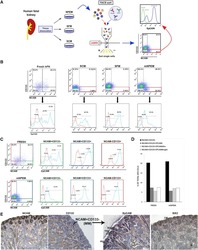
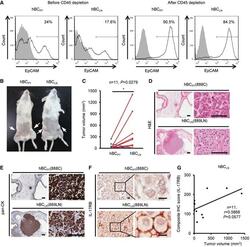
- Figure 8 Elevated IL-17RB expression in cancer cells derived from tumor-draining lymph nodes in breast cancer patients correlates with aggressive growth nature A Freshly collected human breast cancer (hBC) specimens (PT, primary tumors; LN, paired LN metastasis from the same breast cancer patient) were digested with collagenase-containing buffer overnight. Depletion of hematopoietic cells by CD45 Dynabeads (Invitrogen) enriched the EpCAM + cells for following xenografts. Percentages of EpCAM + cells in human primary breast cancer cells were confirmed by FACS analysis (left two panels: before enrichment; right two panels: after enrichment). All of the histograms were shown that they were EpCAM expressing (black line), compared to isotype control (gray filled). B Tumorigenesis assay was determined in NOD/SCID/IL2Rgamma mice orthotopically injected with hBC PT and hBC LN cells. Representative data on day 180 were shown. White arrow: Orthotopic injection site. C Dot plot illustrated the tumor volumes of hBC PT and hBC LN cell-injected NOD/SCID/IL2Rgamma mice on day 180 ( n = 11). All tumors were confirmed by pan-keratin IHC staining of fat pad sections. D, E H&E stainings and IHC stainings of pan-keratin were shown in the representative cases of fat pad sections from (B). Scale bar: 1 mm (left panels), 100 mum (right panels). F IHC stainings of IL-17RB were shown in the representative cases of fat pad sections from (B). Scale bar: 10 mum. G The correlatio
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
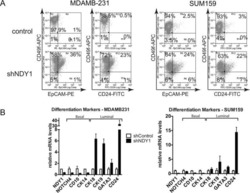
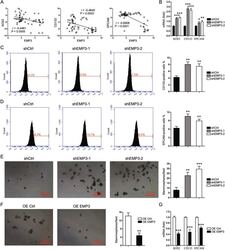
- Fig. 4 EMP3 interferes with stem-like properties. A CCLE database was interrogated for EMP3, SOX2, CD133, and EPCAM expression. Correlation between two genes in a total of 57 breast cancer cell lines was analyzed by Pearson statistics. B SOX2, CD133, and EPCAM expression in MDA-MB-231 cells were detected by real-time PCR. Mean +- SD, n = 3. * P < 0.05, ** P < 0.01, *** P < 0.001, vs. shCtrl. C Fractions of CD133-positive MDA-MB-231 cells were detected by flow cytometry. Mean +- SD, n = 3. ** P < 0.01, vs. shCtrl. D Fractions of EPCAM-positive MDA-MB-231 cells were detected by flow cytometry. Mean +- SD, n = 3. ** P < 0.01, vs. shCtrl. E Mammosphere formation of MDA-MB-231 cells was shown. Mean +- SD, n = 3. ** P < 0.01, *** P < 0.001, vs. shCtrl. F Mammosphere formation of MCF7 cells was shown. Mean +- SD, n = 3. ** P < 0.01, vs. OE Ctrl. G SOX2, CD133, and EPCAM expression in MCF7 cells were detected by real-time PCR. Mean +- SD, n = 3. *** P < 0.001, vs. OE Ctrl.
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
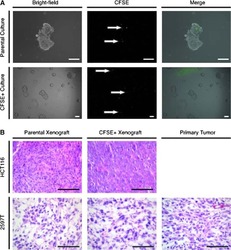
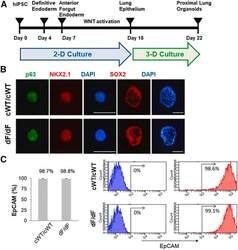
- Figure 1 Generation of proximal lung organoids from cystic fibrosis patient iPSCs (A) Schematic representation of lung organoid differentiation protocol from human iPSCs (hiPSCs). (B) Representative immunofluorescence staining for p63 (green), NKX2.1 (red), SOX2 (red), and nuclear protein expression with DNA stain (DAPI; blue) in HLOs on day 20 (p63 and NKX2.1) and day 22 (SOX2). Scale bars represent 100 mum. (C) Flow cytometry analysis of EpCAM expression in organoid cells on day 23. Left: summary percentage of EpCAM-positive cells. Three independent HLO plates were used for the experiment. Data are presented as mean +- standard error of the mean (SEM). Right: a representative flow cytometry result. The x axis indicates the count of cells, and the y axis indicates EpCAM signal level.
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
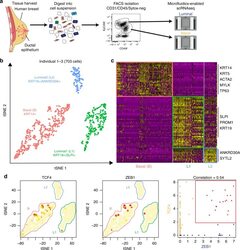
- Fig. 1 Identification of three major epithelial cell types and their markers using scRNAseq. a Overview of scRNAseq approach using primary human breast tissue samples that were processed into single cell suspension, followed by FACS isolation of basal (CD49f-hi, EPCAM+) and luminal (CD49f+, EPCAM-hi), and scRNAseq analysis using the microfluidics-enabled scRNAseq. b Combined tSNE projection of cells from all three microfluidics-enabled scRNAseq datasets. The major basal cluster is highlighted in red; Luminal1 (L1) in green; Luminal2 (L2) in blue. c Heatmap displaying the scaled expression patterns of top marker genes within each cell type with selected marker genes highlighted; yellow indicating high expression of a particular gene, and purple indicating low expression. d Feature plots showing the scaled expression of TCF4 and ZEB1 marking a subpopulation of basal cells and gene plot showing co-expression of TCF4 and ZEB1 in the same cells. See Supplementary Fig. 1 capture site imaging, gene detection, individual principal component analysis, tSNE plot colored by individual-derived cells and feature plots of cell type-specific markers
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
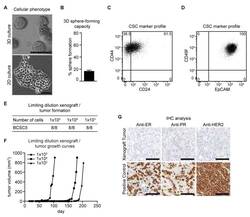
- Figure 1 Characterization of BCSC5 in vitro and in vivo. ( A ) Representative pictures of BCSC5 cultured in 3D and 2D conditions, scale bar 100 mum. ( B ) Sphere-forming capacity of BCSC5 cells in an anchorage-independent growth assay ( n = 3). Data represent means + SEM. ( C , D ) Expression patterns of CD24 and CD44 ( C ) as well as EpCAM and CD49f ( D ) in BCSC5 cells analyzed by flow cytometry. ( E ) Tumor formation in limiting dilution xenografts of BCSC5. ( F ) Representative growth curves for limiting dilution assay of BCSC5 xenografts in immunocompromised NOD/SCID mice. ( G ) Immunohistochemical (IHC) analysis of ER, PR, and HER2 on sections of the BCSC5 xenograft tumors, scale bar 100 mum.
 Explore
Explore Validate
Validate Learn
Learn Flow cytometry
Flow cytometry