13-9700
antibody from Invitrogen Antibodies
Targeting: CTNNA2
CAP-R, CT114
 Western blot
Western blot ELISA
ELISA Immunocytochemistry
Immunocytochemistry Immunoprecipitation
Immunoprecipitation Immunohistochemistry
Immunohistochemistry Flow cytometry
Flow cytometry Other assay
Other assayAntibody data
- Antibody Data
- Antigen structure
- References [0]
- Comments [0]
- Validations
- Western blot [4]
- Immunocytochemistry [2]
- Other assay [9]
Submit
Validation data
Reference
Comment
Report error
- Product number
- 13-9700 - Provider product page

- Provider
- Invitrogen Antibodies
- Product name
- alpha Catenin Monoclonal Antibody (alpha-CAT-7A4)
- Antibody type
- Monoclonal
- Antigen
- Synthetic peptide
- Reactivity
- Human, Mouse, Rat, Chicken/Avian, Xenopus
- Host
- Mouse
- Isotype
- IgG
- Antibody clone number
- alpha-CAT-7A4
- Vial size
- 100 µg
- Concentration
- 0.5 mg/mL
- Storage
- -20°C
No comments: Submit comment
Supportive validation
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- Western blot analysis of HeLa cell lysates using Ms anti-a-Catenin (Product # 13-9700).
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- Knockout of Catenin alpha-1 was achieved by CRISPR-Cas9 genome editing using LentiArray™ Lentiviral sgRNA (Product # A32042, Assay ID CRISPR787281_LV) and LentiArray Cas9 Lentivirus (Product # A32064). Western blot analysis of Catenin alpha-1 was performed by loading 30 µg of HeLa Wild Type (Lane 1), HeLa Cas9 (Lane 2) andHeLa Catenin alpha-1 KO (Lane 3) whole cell extracts. The samples were electrophoresed using NuPAGE™ Novex™ 4-12% Bis-Tris Protein Gel (Product # NP0322BOX). Resolved proteins were then transferred onto a nitrocellulose membrane (Product # IB23001) by iBlot® 2 Dry Blotting System (Product # IB21001). The blot was probed with Anti-alpha Catenin Monoclonal Antibody (alpha-CAT-7A4) (Product # 13-9700, 1:250 dilution) and Goat anti-Mouse IgG (H+L) Superclonal™ Recombinant Secondary Antibody, HRP (Product # A28177, 1:5,000 dilution) using the iBright FL 1000 (Product # A32752). Chemiluminescent detection was performed using Novex® ECL Chemiluminescent Substrate Reagent Kit (Product # WP20005). Loss of signal upon CRISPR mediated knockout (KO) using the LentiArray™ CRISPR product line confirms that antibody is specific to Catenin alpha-1.
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- Western blot analysis was performed on membrane enriched extracts (30 µg) of A549 (Lane 1), HeLa (Lane 2), MDA-MB-231 (Lane 3), MCF7 (Lane 4), NIH/3T3 (Lane 5) and A-431 (Lane 6).The blots were probed with Anti-alpha Catenin Mouse Monoclonal Antibody (Product # 13-9700, 1:250 dilution) and detected by chemiluminescence using Goat anti-Mouse IgG (H+L) Superclonal™ Secondary Antibody, HRP conjugate (Product # A28177, 0.4 µg/mL, 1:2500 dilution). A ~100 kDa band corresponding alpha Catenin was observed across cell lines tested. Known quantity of protein samples were electrophoresed using Novex® NuPAGE® 4-12 % Bis-Tris gel (Product # NP0321BOX), XCell SureLock™ Electrophoresis System (Product # EI0002) and Novex® Sharp Pre-Stained Protein Standard (Product # LC5800). Resolved proteins were then transferred onto a nitrocellulose membrane by Pierce™ Power Blotter System (22834). The membrane was probed with the relevant primary and secondary Antibody using iBind™ Flex Western Starter Kit (Product # SLF2000S). Chemiluminescent detection was performed using Pierce™ ECL Western Blotting Substrate (Product # 32106).
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- Western blot was performed using Anti-alpha Catenin Monoclonal Antibody (alpha-CAT-7A4)(Product # 13-9700) and a 100kDa band corresponding to alpha Catenin was observed across cell lines and tissue extracts tested except PC-3 and Daudi which is reported to be negative. Whole cell extracts (30 µg lysate) of A-431 (Lane 1), HeLa (Lane 2), MCF7 (Lane 3), LNCaP (Lane 4), PC-3 (Lane 5), Daudi (Lane 6), NIH/3T3 (Lane 7) and tissue extracts of Mouse Lung (Lane 8) and Rat Heart (Lane 9) were electrophoresed using NuPAGE™ 4-12% Bis-Tris Protein Gel (Product # NP0322BOX). Resolved proteins were then transferred onto a Nitrocellulose membrane (Product # IB23001) by iBlot® 2 Dry Blotting System (Product # IB21001). The blot was probed with the primary antibody (1:250 dilution) and detected by chemiluminescence with Goat anti-Mouse IgG (H+L) Superclonal™ Recombinant Secondary Antibody, HRP (Product # A28177, 1:4000 dilution) using the iBright FL 1000 (Product # A32752). Chemiluminescent detection was performed using SuperSignal™ West Dura Extended Duration Substrate (Product # 34076).
Supportive validation
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- Immunofluorescent analysis of alpha Catenin was performed using 90% confluent log phase MCF-7 cells. The cells were fixed with 4% paraformaldehyde for 10 minutes, permeabilized with 0.1% Triton™ X-100 for 10 minutes, and blocked with 1% BSA for 1 hour at room temperature. The cells were labeled with alpha Catenin (alpha-CAT-7A4) Mouse Monoclonal Antibody (Product # 13-9700) at 1:250 dilution in0.1% BSA and incubated for 3 hours at room temperature and then labeled with Goat anti-Mouse IgG (H+L) Superclonal™ Secondary Antibody, Alexa Fluor® 488 conjugate (Product # A28175) a dilution of 1:2000 for 45 minutes at room temperature (Panel a: green). Nuclei (Panel b: blue) were stained with SlowFade® Gold Antifade Mountant with DAPI (Product # S36938). F-actin (Panel c: red) was stained with Rhodamine Phalloidin (Product # R415, 1:300). Panel d represents the merged image showing cell junctional localization. Panel e shows the no primary antibody control. The images were captured at 60X magnification.
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
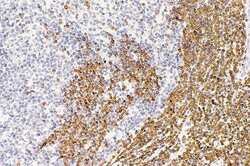
- Experimental details
- Immunocytochemical analysis of Jurkat cell lysates using Mouse anti-Bcl-2 monoclonal antibody (Product # 13-8800)
Supportive validation
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- FIGURE 5: Identification of protein domains involved in cross-linking of p120 and alpha-catenin by BM[PEO]3. (A) A cysteine map of p120-F. The protein consists of three major domains--the central arm-repeat domain (individual arm repeats are depicted by open boxes, numbered) flanked by the amino-terminal (N) and carboxy-terminal (C) domains. Note that all 10 cysteines of p120 (indicated by numbered arrows) are located at the arm-repeat domain, one of which, Cys-9, is present in a large loop separating arm repeats 5 and 6. This loop may contain a small insert encoding by the alternative exon C (filled circle). FLAG epitope (F) is indicated by an open circle. (B) Recruitment of the p120-F cysteine mutants into the cadherin-catenin complex. The p120-F mutants were expressed in A-431 cells and immunoprecipitated with anti-FLAG agarose. The immunoprecipitates were probed for the p120 mutants (p120 mutants; the number of the mutant is indicated above the lanes) and for alpha-catenin (alpha-cat) using anti-FLAG or anti-alpha-catenin antibodies. Note that except for Cys-4, all Cys mutants interact with alpha-catenin. (C) Digitonin cell lysates of A-431 cells expressing p120-F (left) and its different cysteine mutants (right; the mutant's numbers are indicated above the lanes) were cross-linked by BM[PEO]3 and analyzed by Western blotting for p120-F or its mutants (p120F, p120F-mutants) and for alpha-catenin (alpha-cat). Note that the addition of the cross-link (+) resulted in ap
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- Figure 3 Down-regulation of zonula occludens protein 2 (ZO-2) protein expression in myocardium of cyanotic compared with acyanotic patients. Myocardium biopsies were lysed to isolate protein content and western blot analysis performed probing for ZO-2, ZO-1, N-cadherin, beta -catenin, and GAPDH. ZO-2 protein expression was significantly down-regulated in cyanotic biopsies compared with acyanotic, whereas no difference in any other adherens junction expressed proteins was detected. All results were normalized to glyceraldehyde 3-phosphate dehydrogenase (GAPDH) levels. Data are mean +- SEM * = P < 0.05 ( n = 5).
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- Figure 4 Down-regulation of zonula occludens protein 2 (ZO-2) levels in rat cardiomyocytes following 24 h hypoxic treatment. Cells were treated with hypoxic conditions by incubating in a chamber with 5% CO 2 / 0.2% O 2 . After 24 h, cells were lysed, and western blot analysis performed probing for ZO-2, ZO-1, N-cadherin, beta -catenin, and beta -actin, normalizing all results to glyceraldehyde 3-phosphate dehydrogenase (GAPDH) levels. ZO-2 protein expression was significantly down-regulated in hypoxic treated cells compared with untreated cells (control), whereas no difference in ZO-1, N-cadherin, beta -catenin, or beta -actin levels was detected between treated and untreated cells. Data are mean +- SEM ** = P < 0.01 ( n = 4).
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
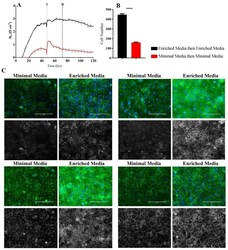
- Figure 6 Immunocytochemistry of brain endothelial junctional space following growth in either Enriched Media or Minimal Media. Cells were seeded at time 0 h at 20,000 cells per 0.3 cm 2 . ( A ) ECIS resistance and R b measurements over 120 h. Cells were grown in either Enriched Media or Minimal Media from T = 0 h, with media changed at T = 48 h (I). Data shown is the mean +- SD (n = 3 wells) of one independent experiment representative of three experimental repeats; ( B ) Cell number count of Hoechst stained nuclei at time point II, obtained through Image J Software analysis. Data shown is the mean +- SEM (n = 18 wells) of 1 independent experiment representative of 3 experimental repeats; ( C ) The junctional space following growth in either Enriched Media or Minimal Media 72 h post seeding. The time point of fixation corresponds to the second vertical dotted line (II) shown on the ECIS traces. Representative GFP/DAPI and GFP monochrome images for the junctional proteins CD144, ZO-1, beta-catenin, and alpha-catenin are shown. Each panel shows cells grown in Minimal Media on the left and cells grown in Enriched Media on the right. The corresponding Alexa Fluor 488 (GFP) monochrome image is shown below each merged image. Green--junctional protein, blue--nuclei. Scale bar is 200 um. Immunocytochemistry data show one representative image from one independent experiment, which is representative of three experimental repeats. Graphical representations of p values are * p
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- Figure 7 Immunocytochemistry of brain endothelial junctional space following growth in either Enriched Media or Minimal Media. Cells were seeded at time 0 h at 20,000 cells per 0.3 cm 2 . ( A ) ECIS resistance and R b measurements over 120 h. Cells were grown in either Enriched Media or Minimal Media from T = 0 h, with media changed at T = 48 h (I). Data shown is the mean +- S.D. (n = 3 wells) of 1 independent experiment representative of 3 experimental repeats; ( B ) Cell number count of Hoechst stained nuclei at time point II, obtained through Image J Software analysis. Data shown is the mean +- SEM (n = 18 wells) of 1 independent experiment representative of 3 experimental repeats; ( C ) The junctional space following growth in either Enriched Media or Minimal Media 72 h post seeding. The time point of fixation corresponds to the second vertical dotted line (II) shown on the ECIS traces. Representative GFP/DAPI and GFP monochrome images for the junctional proteins CD144, ZO-1, beta-catenin and alpha-catenin are shown. Each panel shows cells grown in Minimal Media on the left and cells grown in Enriched Media on the right. The corresponding Alexa Fluor 488 (GFP) monochrome image is shown below each merged image. Green--junctional protein, blue--nuclei. Scale bar is 200 um. Immunocytochemistry data shows 1 representative image from 1 independent experiment, which is representative of 3 experimental repeats. Graphical representations of p values are * p
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- Endothelial and specialized epithelial-specific proteins. Representative indirect fluorescent digital photomicrographs of cryostat axial sections of normal adult sural nerves show UEA-1-positive endothelial cells (green; A, E, I, M, Q) with expression of CAV1, GJA1, TJP1, ZYX, and CTNNA1 (red; B, F, J, N, R, respectively) restricted to endoneurial microvessels, fenestrated epineurial macrovessels and the perineurium, shown in the merged images at lower and higher magnifications (yellow/orange for endothelial cells). Expression of these proteins by both microvascular and macrovascular endothelial cells and the perineurium suggests non-restrictive barrier, but specialized endothelial and epithelial cell functions in the normal adult human peripheral nerves. White arrows demonstrate positively staining endoneurial microvessels. Blue (4, 6-diamidino-2-phenylindole) staining indicates nuclei. Scale bar 500 mum for A-C, E-G, I-K, M-O, and Q-S and 125 mum for D, H, L, P, and T
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- Figure 5 Knockdown of Rai14 in the Sertoli cell epithelium with an established TJ-permeability barrier in vitro by RNAi disrupts actin filament organization and the TJ barrier. (A) Sertoli cells cultured alone on Matrigel-coated 12-well dishes for 2-day with an established TJ-permeability barrier were transfected with Rai14 siRNA duplexes (Rai14 RNAi) versus non-targeting control duplexes (Ctrl RNAi) at 100 nM using Ribojuice transfection medium for 24 hr, thereafter, cells were washed twice and cultured in F12/DMEM for 12 hr to allow recovery. Thereafter, cells were transfected again under the same conditions for another 24 hr. Thereafter, cells were rinsed with fresh F12/DMEM and cultured for an additional 12 hr before termination, and used to prepare lysates for immunoblotting using antibodies against several BTB-associated constituent or regulatory proteins. A knockdown of Rai14 by ~50% was noted in which the control was arbitrarily set at 1 against which statistical comparison was performed (B) without any apparent off-target effects (A). The findings shown herein are the results of 3 independent experiments excluding pilot experiments which were used to establish optimal experimental conditions, such as different concentrations of siRNA duplexes and Ribojuice. It was noted that we achieved only ~50-60% knockdown of Rai14 in several pilot experiments, unlike other target genes [ e.g ., Scribble, beta1-integrin, and P-glycoprotein] wherein we could silence the target gene
 Explore
Explore Validate
Validate Learn
Learn