Antibody data
- Antibody Data
- Antigen structure
- References [0]
- Comments [0]
- Validations
- Western blot [7]
- Immunocytochemistry [4]
- Immunohistochemistry [3]
- Flow cytometry [4]
- Other assay [21]
Submit
Validation data
Reference
Comment
Report error
- Product number
- MA5-15738-BTIN - Provider product page

- Provider
- Invitrogen Antibodies
- Product name
- GAPDH Loading Control Monoclonal Antibody (GA1R), Biotin
- Antibody type
- Monoclonal
- Antigen
- Recombinant full-length protein
- Description
- MA5-15738-BTIN has successfully been used for Western blot, FACS, ELISA, IHC (P) and ICC/IF applications.
- Reactivity
- Human, Mouse, Rat, Bacteria, Chicken/Avian, Hamster, Rabbit, Yeast
- Host
- Mouse
- Conjugate
- Biotin
- Isotype
- IgG
- Antibody clone number
- GA1R
- Vial size
- 50 µL
- Concentration
- 1 mg/mL
- Storage
- 4° C, do not freeze
No comments: Submit comment
Supportive validation
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
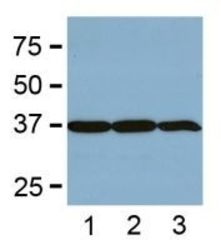
- Experimental details
- WB from BL-21 bacteria (1), Sf9 insect (2), and Saccharomyces cerevisiae (3).
- Conjugate
- Biotin
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
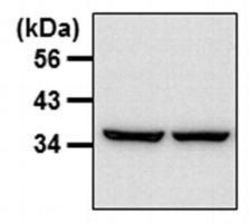
- Experimental details
- Western blot analysis of GAPDH was performed by loading 20 µg of THP-1 whole cell lysates per well onto an SDS-PAGE gel. Proteins were transferred to a PVDF membrane and blocked with 5% milk in TBST. The membrane was probed with a GAPDH loading control monoclonal antibody (Product # MA5-15738) at a dilution of 1:5000 overnight at 4ºC, washed in TBST, and probed with a HRP-conjugated goat anti-mouse IgG at a dilution of 1:40,000 for 1 hr at room temperature. Detection was performed using ECL substrate. Data courtesy of the Innovators Program.
- Conjugate
- Biotin
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- Western blot analysis of GAPDH was performed by loading 20 µg of THP-1 whole cell lysates per well onto an SDS-PAGE gel. Proteins were transferred to a PVDF membrane and blocked with 5% milk in TBST. The membrane was probed with a GAPDH loading control monoclonal antibody (Product # MA5-15738) at a dilution of 1:5000 overnight at 4ºC, washed in TBST, and probed with a HRP-conjugated goat anti-mouse IgG at a dilution of 1:40,000 for 1 hr at room temperature. Detection was performed using ECL substrate. Data courtesy of the Innovators Program.
- Conjugate
- Biotin
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- 1:2000 (0.5 µg/mL) Ab dilution used in WB of 5 µg/lane tissue lysates from human (1), mouse (2), rat (3), rabbit (4), chicken (5), and hamster (6).
- Conjugate
- Biotin
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
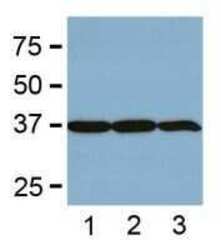

- Experimental details
- WB from BL-21 bacteria (1), Sf9 insect (2), and Saccharomyces cerevisiae (3).
- Conjugate
- Biotin
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- Western blot analysis of GAPDH was performed by loading 50 µg of various cell lysates per well onto a 4-20% Tris-HCl polyacrylamide gel. Proteins were transferred to a PVDF membrane and blocked with 5% BSA/TBST for at least 1 hour. The membrane was probed with a biotinylated GAPDH monoclonal antibody (Product # MA5-15738-BTIN) at a dilution of 1:1000 for 1 hour at room temperature, followed by Streptavidin-HRP (Product # 21126) at a dilution of 1:20,000 for 30 minutes at room temperature. Chemiluminescent detection was performed using SuperSignal West Pico (Product # 34078).
- Conjugate
- Biotin
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
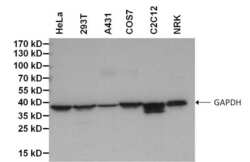
- Experimental details
- Western blot analysis of GAPDH was performed by loading 50 µg of various cell lysates per well onto a 4-20% Tris-HCl polyacrylamide gel. Proteins were transferred to a PVDF membrane and blocked with 5% BSA/TBST for at least 1 hour. The membrane was probed with a biotinylated GAPDH monoclonal antibody (Product # MA5-15738-BTIN) at a dilution of 1:1000 for 1 hour at room temperature, followed by Streptavidin-HRP (Product # 21126) at a dilution of 1:20,000 for 30 minutes at room temperature. Chemiluminescent detection was performed using SuperSignal West Pico (Product # 34078).
- Conjugate
- Biotin
Supportive validation
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
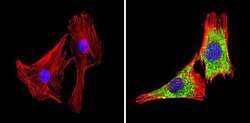
- Experimental details
- Immunofluorescent analysis of GAPDH (green) showing staining in the in the cytoplasm of C2C12 cells (right) compared to a negative control without primary antibody (left). Formalin-fixed cells were permeabilized with 0.1% Triton X-100 in TBS for 5-10 minutes and blocked with 3% BSA-PBS for 30 minutes at room temperature. Cells were probed with a GAPDH loading control antibody (Product # MA5-15738) in 3% BSA-PBS at a dilution of 1:100 and incubated overnight at 4ºC in a humidified chamber. Cells were washed with PBST and incubated with a DyLight-conjugated secondary antibody in PBS at room temperature in the dark. F-actin (red) was stained with a fluorescent red phalloidin and nuclei (blue) were stained with Hoechst or DAPI. Images were taken at a magnification of 60x.
- Conjugate
- Biotin
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
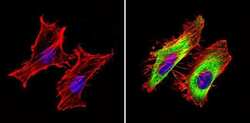
- Immunofluorescent analysis of GAPDH (green) showing staining in the in the cytoplasm of Hela cells (right) compared to a negative control without primary antibody (left). Formalin-fixed cells were permeabilized with 0.1% Triton X-100 in TBS for 5-10 minutes and blocked with 3% BSA-PBS for 30 minutes at room temperature. Cells were probed with a GAPDH loading control antibody (Product # MA5-15738) in 3% BSA-PBS at a dilution of 1:100 and incubated overnight at 4ºC in a humidified chamber. Cells were washed with PBST and incubated with a DyLight-conjugated secondary antibody in PBS at room temperature in the dark. F-actin (red) was stained with a fluorescent red phalloidin and nuclei (blue) were stained with Hoechst or DAPI. Images were taken at a magnification of 60x.
- Conjugate
- Biotin
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
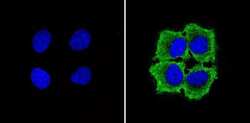
- Immunofluorescent analysis of GAPDH (green) showing staining in the in the cytoplasm of MCF-7 cells (right) compared to a negative control without primary antibody (left). Formalin-fixed cells were permeabilized with 0.1% Triton X-100 in TBS for 5-10 minutes and blocked with 3% BSA-PBS for 30 minutes at room temperature. Cells were probed with a GAPDH loading control antibody (Product # MA5-15738) in 3% BSA-PBS at a dilution of 1:100 and incubated overnight at 4ºC in a humidified chamber. Cells were washed with PBST and incubated with a DyLight-conjugated secondary antibody in PBS at room temperature in the dark. Nuclei (blue) were stained with Hoechst or DAPI. Images were taken at a magnification of 60x.
- Conjugate
- Biotin
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
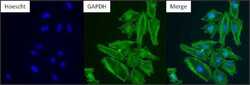
- Immunofluorescent analysis of GAPDH (green) in HeLa. Formalin-fixed cells were permeabilized with 0.1% Triton X-100 in TBS for 10 minutes at room temperature and blocked with 5% normal goat serum (Product # 31873) for 15 minutes at room temperature. Cells were probed with a GAPDH monoclonal antibody (Product # MA5-15738) at a dilution of 1:100 for at least 1 hour at room temperature, washed with PBS, and incubated with DyLight 488 goat anti-mouse IgG secondary antibody (Product # 35502) at a dilution of 1:400 for 30 minutes at room temperature. Nuclei (blue) were stained with Hoechst 33342 dye (Product # 62249). Images were taken on a Thermo Scientific ArrayScan or ToxInsight Instrument at 20X magnification.
- Conjugate
- Biotin
Supportive validation
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
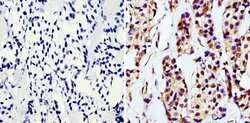
- Immunohistochemistry analysis of GAPDH showing staining in the nucleus and cytoplasm of paraffin-embedded human breast carcinoma (right) compared with a negative control without primary antibody (left). To expose target proteins, antigen retrieval was performed using 10mM sodium citrate (pH 6.0), microwaved for 8-15 min. Following antigen retrieval, tissues were blocked in 3% H2O2-methanol for 15 min at room temperature, washed with ddH2O and PBS, and then probed with a GAPDH monoclonal antibody (Product # MA5-15738) diluted in 3% BSA-PBS at a dilution of 1:200 overnight at 4°C in a humidified chamber. Tissues were washed extensively in PBST and detection was performed using an HRP-conjugated secondary antibody followed by colorimetric detection using a DAB kit. Tissues were counterstained with hematoxylin and dehydrated with ethanol and xylene to prep for mounting.
- Conjugate
- Biotin
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
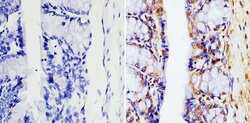
- Immunohistochemistry analysis of GAPDH showing staining in the nucleus and cytoplasm of paraffin-embedded mouse colon tissue (right) compared with a negative control without primary antibody (left). To expose target proteins, antigen retrieval was performed using 10mM sodium citrate (pH 6.0), microwaved for 8-15 min. Following antigen retrieval, tissues were blocked in 3% H2O2-methanol for 15 min at room temperature, washed with ddH2O and PBS, and then probed with a GAPDH monoclonal antibody (Product # MA5-15738) diluted in 3% BSA-PBS at a dilution of 1:100 overnight at 4°C in a humidified chamber. Tissues were washed extensively in PBST and detection was performed using an HRP-conjugated secondary antibody followed by colorimetric detection using a DAB kit. Tissues were counterstained with hematoxylin and dehydrated with ethanol and xylene to prep for mounting.
- Conjugate
- Biotin
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
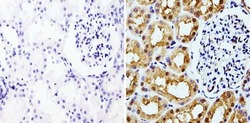
- Immunohistochemistry analysis of GAPDH showing staining in the nucleus and cytoplasm of paraffin-embedded human kidney tissue (right) compared with a negative control without primary antibody (left). To expose target proteins, antigen retrieval was performed using 10mM sodium citrate (pH 6.0), microwaved for 8-15 min. Following antigen retrieval, tissues were blocked in 3% H2O2-methanol for 15 min at room temperature, washed with ddH2O and PBS, and then probed with a GAPDH monoclonal antibody (Product # MA5-15738) diluted in 3% BSA-PBS at a dilution of 1:100 overnight at 4°C in a humidified chamber. Tissues were washed extensively in PBST and detection was performed using an HRP-conjugated secondary antibody followed by colorimetric detection using a DAB kit. Tissues were counterstained with hematoxylin and dehydrated with ethanol and xylene to prep for mounting.
- Conjugate
- Biotin
Supportive validation
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
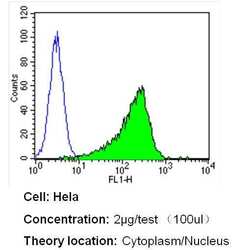
- Flow cytometry analysis of GAPDH in Hela cells (green) compared to an isotype control (blue). Cells were harvested, adjusted to a concentration of 1-5x10^6 cells/mL, fixed with 2% paraformaldehyde and washed with PBS. Cells were blocked with a 2% solution of BSA-PBS for 30 min at room temperature and incubated with a GAPDH loading control antibody (Product # MA5-15738) at a dilution of 2 µg/test for 40 min at room temperature. Cells were then incubated for 40 min at room temperature in the dark using a Dylight 488-conjugated secondary antibody and re-suspended in PBS for FACS analysis.
- Conjugate
- Biotin
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
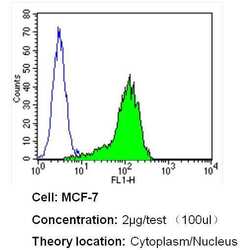
- Flow cytometry analysis of GAPDH in MCF-7 cells (green) compared to an isotype control (blue). Cells were harvested, adjusted to a concentration of 1-5x10^6 cells/mL, fixed with 2% paraformaldehyde and washed with PBS. Cells were blocked with a 2% solution of BSA-PBS for 30 min at room temperature and incubated with a GAPDH loading control antibody (Product # MA5-15738) at a dilution of 2 µg/test for 40 min at room temperature. Cells were then incubated for 40 min at room temperature in the dark using a Dylight 488-conjugated secondary antibody and re-suspended in PBS for FACS analysis.
- Conjugate
- Biotin
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
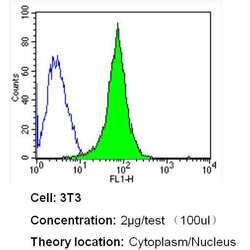
- Flow cytometry analysis of GAPDH in NIH-3T3 cells (green) compared to an isotype control (blue). Cells were harvested, adjusted to a concentration of 1-5x10^6 cells/mL, fixed with 2% paraformaldehyde and washed with PBS. Cells were blocked with a 2% solution of BSA-PBS for 30 min at room temperature and incubated with a GAPDH loading control antibody (Product # MA5-15738) at a dilution of 2 µg/test for 40 min at room temperature. Cells were then incubated for 40 min at room temperature in the dark using a Dylight 488-conjugated secondary antibody and re-suspended in PBS for FACS analysis.
- Conjugate
- Biotin
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
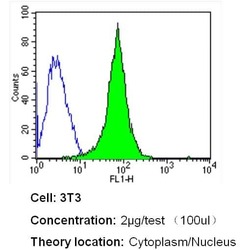
- Flow cytometry analysis of GAPDH in NIH-3T3 cells (green) compared to an isotype control (blue). Cells were harvested, adjusted to a concentration of 1-5x10^6 cells/mL, fixed with 2% paraformaldehyde and washed with PBS. Cells were blocked with a 2% solution of BSA-PBS for 30 min at room temperature and incubated with a GAPDH loading control antibody (Product # MA5-15738) at a dilution of 2 µg/test for 40 min at room temperature. Cells were then incubated for 40 min at room temperature in the dark using a Dylight 488-conjugated secondary antibody and re-suspended in PBS for FACS analysis.
- Conjugate
- Biotin
Supportive validation
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
- NULL
- Conjugate
- Biotin
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
- NULL
- Conjugate
- Biotin
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
- NULL
- Conjugate
- Biotin
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
- NULL
- Conjugate
- Biotin
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
- NULL
- Conjugate
- Biotin
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
- NULL
- Conjugate
- Biotin
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
- NULL
- Conjugate
- Biotin
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
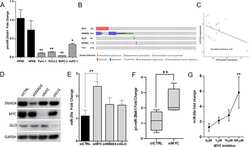
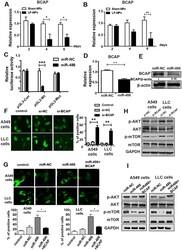
- Figure 6 miR-486 directly regulate post-transcriptional expression of BCAP. ( A , B ) A549 and LLC cells were treated as described in Fig. 2C . Expression levels of BCAP were detected by qRT-PCR. ( C ) Luciferase reporters containing wild-type (WT) or mutant (MUT) miR-486 binding sites in the BCAP 3'-UTR were co-transfected into 293 T cells along with control luciferase reporter plasmid, miR-NC, miR-486 mimic for 48 hours. ( D - E ) qRT-PCR analysis of BCAP mRNA levels and Western blot analysis of BCAP protein levels in A549 cell treated with miR-486 mimic. ( F ) A549 and LLC cells were co-transfected with GFP-LC3 plasmid, siNC or siBCAP for 48 h. Number of autophagosomes was quantified. ( G ) A549 and LLC cells were transfected with GFP-LC3 plasmid, miR-NC, miR-486 mimic or co-transfected with miR-486 mimic and BCAP plasmid. Number of autophagosomes was quantified. ( H ) A549 and LLC cells transfected with siNC or siBCAP for 48 h. Western blot analysis of phospho-AKT, AKT, phospho-mTOR, mTOR and GAPDH expression. ( I ) A549 or LLC cells transfected with miR-NC, miR-486 mimic or co-transfected with miR-486 and BCAP plasmid. Western blot analysis of phospho-AKT, AKT, phospho-mTOR, mTOR and GAPDH expression. All error bars indicate mean +- SEM. Experiments were repeated three times independently. ** P < 0.01.
- Conjugate
- Biotin
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
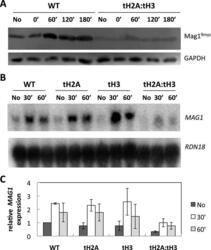
- Figure 3. Mag1 protein expression and MAG1 mRNA expression are reduced in tH2A:tH3 cells. WT and the N-tail deletion mutants were treated with 0.2% MMS for 10 min and further analyzed at the indicated time points. ( A ) Mag1 9myc and GAPDH were detected via western blotting. ( B ) A northern blot detecting MAG1 and reprobed for RDN18 . [We note that the small band shift between the tH3(60') sample and the tH2A:tH3 samples is due to a gel running artifact]. ( C ) Quantification of different gels, such as shown in (B) by normalizing MAG1 levels to that of WT in the absence of MMS. Error bars indicate standard deviations derived from three independent experiments.
- Conjugate
- Biotin
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
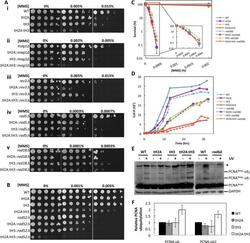
- Figure 6. The N-tails of H2A and H3 are epistatic to RAD18 . ( A ) Epistasis studies of (i) N-tail deletion mutants combinatorially deleted for (ii) MAG1 , (iii) REV3 , (iv) RAD5 and (v) RAD18 . ( B ) Epistasis study of N-tail deletion mutants combinatorially deleted for RAD52 . ( C ) MMS survival assay of WT and N-tail mutants (solid lines) and the N-tail mutants combinatorially deleted for RAD18 (dashed lines). ( D ) Growth curves of the cells in ( C ) in the absence of MMS. ( E ) Western blot of PCNA 9myc in WT and N-tail mutants before UV exposure and after a 1 h recovery in YPD following 100 J/m 2 of UV irradiation. GAPDH is used as the loading control. rad6Delta serves as a negative control for PCNA ubiquitylation. ( F ) Quantifications of (E) by normalizing (PCNA 9myc -ub)/(PCNA 9myc -total) and (PCNA 9myc -ub 2 )/(PCNA 9myc -total) to WT. Error bars indicate standard deviations from three independent experiments.
- Conjugate
- Biotin
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
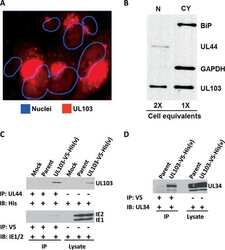
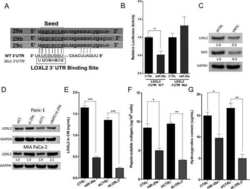
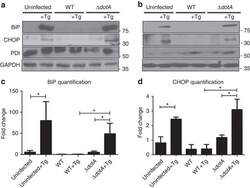
- Figure 1 Expression of two critical components of the UPR, BiP and CHOP are blocked by L. pneumophila ( a ) Caspase 1/11 (-/-) bone marrow derived macrophages or ( b ) HEK-293 FCgamma cells-expressing Fc gamma (HEK-293 FCgamma) were either left uninfected or infected with WT or DeltadotA strains of L. pneumophila at an MOI of 150. Cells were then left untreated or treated with thapsigargin (Tg; 1 mug mul -1 ) for 6 h. BiP, CHOP and PDI protein expression were monitored via immunoblot. GAPDH was used as a loading control. Full blots are provided for Fig. 1a,b in Supplementary Fig. 5 . ( c , d ) BiP and CHOP levels were quantified respectively from three biological replicates using the same experimental conditions in HEK-293 FCgamma cells. Data are depicted as the fold change normalized to uninfected levels. Data represent three independent experiments. Values in all graphs are means+-s.e.m. * P
- Conjugate
- Biotin
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
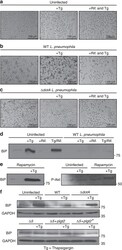
- Figure 3 Suppression of ER stress is an effector driven process that is independent of the mTOR pathway. ( a - c ) HEK-293 FCgamma cells were either left uninfected or infected with WT L. pneumophila or DeltadotA L. pneumophila at an MOI of 150. Cells were then either left untreated or treated with thapsigargin (Tg; 1 mug mul -1 ) or treated with a combination of Tg and the antibiotic Rifampicin (Rif.; 2 mug mul -1 ) for 6 h. After 6 h phase contrast images were taken. ( d ) HEK-293 FCgamma cells were infected at an MOI of 150 with WT L. pneumophila , or left uninfected. Cells were then treated with Rif. alone (2 mug mul -1 ), Rif. with Tg (1 mug mul -1 ), Tg (1 mug mul -1 ) alone or left untreated for 6 h. BiP protein levels were monitored via immunoblot. ( e ) HEK-293 FCgamma cells were untreated or treated with rapamycin (10 nM) or rapamycin with Tg (1 mug mul -1 ) for 6 h. BiP protein levels were then monitored via immunoblot and levels of phosphorylated Akt at serine 473 (P-Akt) were monitored from these lysates via immunoblot. ( f ) RAW 264.7 cells were infected with WT L. pneumophila, dotA , or Delta 5 strains of L. pneumophila at an MOI of 150. BiP protein expression was monitored via immunoblot. GAPDH was used as a loading control. Full blots for Fig. 3d-f are provided in Supplementary Fig. 5 .
- Conjugate
- Biotin
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
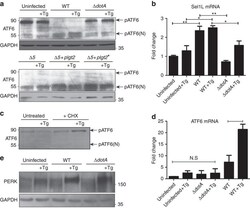
- Figure 6 ATF6 is processed during L. pneumophila infection. ( a ) RAW 264.7 cells were left uninfected or infected with either WT L. pneumophila , DeltadotA , B. Delta5, Delta5+plgt2, or Delta5 plgt2* (catalytically dead lgt2) at an MOI of 150. Cells were then left untreated or treated with thapsigargin (Tg; 1 mug mul -1 ) for 6 h. Full-length ATF6 (pATF6) and cleaved pATF6(N) were monitored via immunoblot. Full blots for Fig. 6a are provided in Supplementary Fig. 5 . ( b ) HEK-293 FCgamma cells infected with either WT L. pneumophila or DeltadotA and then treated for 6 h with Tg (1 mug mul -1 ). Sel1L mRNA levels were then monitored via qRT-PCR, with GAPDH serving as an endogenous control. Two biological replicates were used each with three technical replicates using the identical experimental conditions. Values in all graphs are means+-s.e.m. * P
- Conjugate
- Biotin
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
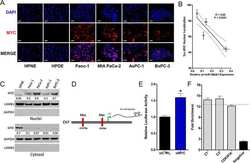
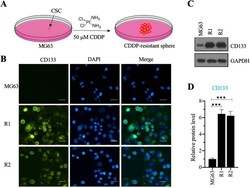
- Figure 1 Screening cisplatin (CDDP)-resistant osteosarcoma cancer stem cells (CSCs) and detecting CD133 level in these cells. (A) A schematic diagram of the screening procedure for CDDP-resistant osteosarcoma CSCs. (B) Immunofluorescence staining for CD133. An osteosarcoma CSC marker CD133 was stained using its specific antibody in MG63, MG63-R1, and MG63-R2 cells. The same cells were also counterstained with DAPI. Bars=25 um. (C and D) Protein level of CD133. Total cell lysates from MG63, MG63-R1, and MG63-R2 cells were subjected to western blotting to determine CD133 and GAPDH (loading control) levels (C). The CD133 protein signals in different cells were quantified and normalized to their corresponding GAPDH (D). *** P < 0.001.
- Conjugate
- Biotin
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
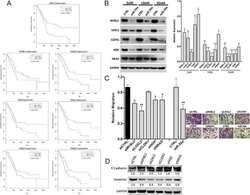
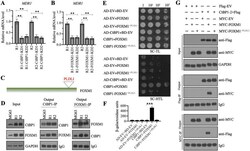
- Figure 4 CtBP1 directly interacted with FOXM1 in vivo and in vitro. (A and B) Knockdown of CtBP1 or FOXM1 significantly decreased the expression of MDR1 . Total RNA samples from MG63-R1, MG63-R2, and two independent knockdown cell lines of CtBP1-KD (A), and two independent knockdown cell lines of FOXM1-KD (B) underwent RT-qPCR analyses to examine MDR1 expression. ** P < 0.01. (C) The FOXM1 protein contained a PLDLI motif. (D) CtBP1 directly interacted with FOXM1 in vivo . Total cell lysates from MG63, MG63-R1, and MG63-R2 cells were subjected to IP assays using anti-CtBP1-coupled and anti-FOXM1-coupled Protein G beads. The input and output proteins were subjected to western blotting to detect CtBP1 and FOXM1 protein levels. GAPDH and IgG were the loading controls for input and output proteins, respectively. (E) CtBP1 directly interacted with FOXM1 in yeast cells. Yeast cells expressing different plasmids were subjected to plate-dotting assays on SC-TL and SC-HTL media with different dilution folds (1, 10 1 , 10 2 , and 10 3 ). (F) beta-galactosidase activity. Yeast cells used in (E) were applied to measure beta-galactosidase activity. *** P
- Conjugate
- Biotin
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
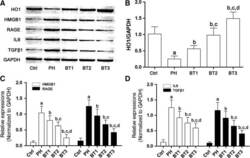
- Figure 4 Effects of baicalin on the HO 1- HMGB 1- RAGE pathway in infant rats with PH . (A) Immunoblots of HO 1, HMGB 1, RAGE , IL 6, TGF beta1 and GAPDH in lung tissue harvested from infant rats with PH . (B) The relative expression level of HO 1 in the Ctrl, PH , BT 1, BT 2 and BT 3 groups. (C) The relative expression of HMGB 1 (white columns) and RAGE (black columns) in the Ctrl, PH , BT 1, BT 2 and BT 3 groups. (D) The relative expression of IL 6 (white columns) and TGF beta1 (black columns) in the Ctrl, PH , BT 1, BT 2 and BT 3 groups. a-d: differences were significant when compared with Ctrl (a), PH (b), BT 1 (c) and BT 2 (d).
- Conjugate
- Biotin
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- Figure 2 Overexpression of HIPK2 caused the phosphorylation and degradation of CtBP1. (A-C) The mRNA levels of HIPK2 , CtBP1 and CtBP2 in HIPK2-OE cells. RNA samples from hFOB1.19+pCDNA3 (hFOB1.19), hFOB1.19+pCDNA3-HIPK2 (hFOB1.19+HIPK2), MG63+pCDNA3 (MG63), MG63+pCDNA3-HIPK2 (MG63+HIPK2), Saos2+pCDNA3 (Saos2), and Saos2+pCDNA3-HIPK2 (Saos2+HIPK2) cells were subjected to RT-qPCR analysis to determine the mRNA levels of HIPK2 (A), CtBP1 (B) and CtBP2 (C). *** P
- Conjugate
- Biotin
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
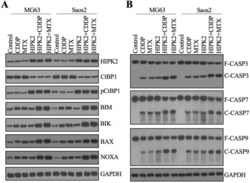
- Figure 5 Overexpression of HIPK2 induced apoptosis. (A) The protein levels of proapoptotic proteins. Total cell extracts from MG63, MG63+CDDP, MG63+MTX, MG63+HIPK2, MG63+HIPK2+CDDP, MG63+HIPK2+MTX, Saos2, Saos2+CDDP, Saos2+MTX, Saos2+HIPK2, Saos2+HIPK2+CDDP, and Saos2+HIPK2+MTX cells were subjected to western blotting to examine protein levels of HIPK2, CtBP1, pCtBP1, BIM, BIK, VAX, NOXA, and GAPDH (loading control). (B) The protein levels of caspases. The same protein samples used in (A) were subjected to western blotting to examine the protein levels of CSAP3, CASP7, CASP9, and GAPDH (loading control).
- Conjugate
- Biotin
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
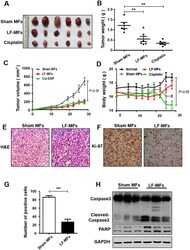
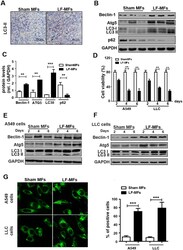
- Figure 1 LF-MFs inhibit tumor growth in LLC mice model. Mice (n = 15, each group) were injected subcutaneously with Lewis lung cancer cells (1 x 10 6 cells) and were treated with LF-MFs (0.4T, 7.5 Hz, 4 h/day) or Sham MFs for 35 days. ( A ) Representative tumors excised from mice treated with LF-MFs or Sham MFs. Cisplatin was used as a positive control. ( B ) LF-MFs treatment reduced the weight of LLC tumors. ( C ) LF-MFs treatment significantly inhibited the growth of LLC tumors. ( D ) LF-MFs treatment didn't affect the body weight of mice. ( E ) H&E staining of tumor tissues in sham MFs or LF-MF treated mice. ( F ) Immunohistochemical staining for proliferation marker (Ki-67) in tumor tissues. ( G ) Ki-67 positive cells were quantified in tumors tissues. ( H ) Protein expression of Caspase3, cleaved-caspase3, and PARP were detected in tumor tissues from sham MFs group (n = 3) and LF-MFs group (n = 3) by Western blotting. Data from one representative experiment performed in triplicate. ** P < 0.01 and *** P < 0.001.
- Conjugate
- Biotin
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
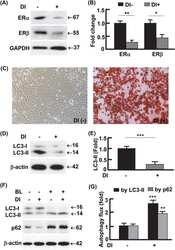
- Figure 2 LF-MFs induce autophagy in tumor tissues and lung cancer cells. ( A ) Mice were treated as described in Fig. 1 and tumor tissues were acquired from each mouse. Expression of LC3II was detected using immunohistochemistry staining. ( B , C ) Protein expression of Beclin-1, ATG5, LC3B and p62 were detected in tumor tissues from sham MFs group (n = 3) and LF-MFs group (n = 3) by Western blotting. ( D ) A549 and LLC cells were treated with sham MFs or LF-MFs for different time intervals (2, 4, 6, days, 4 h/day). Cell activity was analyzed by using a cell counting kit-8 assay. ( E , F ) A549 and LLC cells were treated with sham MFs or LF-MFs for different time intervals (2, 4, 6, days, 4 h/day). ( G ) Fluorescent images of A549 and LLC cells transfected with the GFP-LC3 plasmids and treated with sham MFs or LF-MFs for 6d (original magnification x 1000). The images showed GFP-LC3 dots formation in LF-MFs-treated cells. All error bars indicate mean +- SEM. Experiments were repeated three times independently. *** P < 0.001.
- Conjugate
- Biotin
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
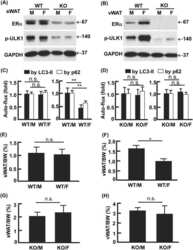
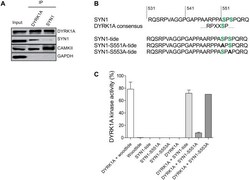
- Fig. 9. Direct interaction of DYRK1A and SYN1, phosphorylation of SYN1 by DYRK1A. (A) DYRK1A and SYN1 were immunoblotted following immunoprecipitation from wt mice brain extracts. DYRK1A or SYN1 present in the starting material (Input) were recovered in the IPs. SYN1 (74 kDa) was present in the DYRK1A IP and DYRK1A (85 kDa) was detected in the SYN1 IP, suggesting that these two proteins interact directly. Positive control of the SYN1 IP was performed using an anti-CAMKII antibody. As expected, CAMKII (50 kDa) was present in the SYN1 IP. DYRK1A IP also brought down CAMKII, suggesting complexes between SYN1, CAMKII and DYRK1A. (B) Sequence of SYN1 in the vicinity of Ser551 matches with the consensus DYRK1A phosphorylation site. Based on this sequence, three peptides were synthesized and used as potential substrates: SYN1, SYN1-S551A and SYN1-S553A. (C) Kinase activity of recombinant DYRK1A towards the three different SYN1peptides. SYN1 and SYN1-S553A peptides were phosphorylated at the same level as Woodtide by recombinant DYRK1A (71.7%+-5.2%, 70.1% and 78.4%+-11.4%, respectively). No significant catalytic activity was measured with the SYN1-S551A peptide (7.9%+-1.2%).
- Conjugate
- Biotin
 Explore
Explore Validate
Validate Learn
Learn Western blot
Western blot ELISA
ELISA