Antibody data
- Antibody Data
- Antigen structure
- References [0]
- Comments [0]
- Validations
- Western blot [3]
- Immunocytochemistry [3]
- Flow cytometry [1]
- Other assay [5]
Submit
Validation data
Reference
Comment
Report error
- Product number
- A21994 - Provider product page

- Provider
- Invitrogen Antibodies
- Product name
- GAPDH Monoclonal Antibody (3E8AD9)
- Antibody type
- Monoclonal
- Antigen
- Purifed from natural sources
- Reactivity
- Human
- Host
- Mouse
- Isotype
- IgG
- Antibody clone number
- 3E8AD9
- Vial size
- 100 µg
- Concentration
- 1 mg/mL
- Storage
- 4° C
No comments: Submit comment
Supportive validation
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- Knockdown of GAPDH was achieved by transfecting HeLa cells with GAPDH specific validated siRNAs (Silencer® select Product # s5572, s5573 ). Western blot analysis (Fig. a) was performed using whole cell extracts from the GAPDH knockdown cells (lane 3), non-specific scrambled siRNA transfected cells (lane 2) and untransfected cells (lane 1). The blots were probed with GAPDH Monoclonal Antibody (Product # A21994, 1:4000 dilution) and Goat anti-Mouse IgG (H+L) Superclonal™ Secondary Antibody, HRP conjugate (Product # A28177, 0.25 µg/mL, 1:4000 dilution). Densitometric analysis of this western blot is shown in histogram (Fig. b). Decrease in signal upon siRNA mediated knock down confirms that antibody is specific to GAPDH.
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- Samples were separated by SDS-PAGE (gradient gel 10-20%). Bands were transferred to a PVDF membrane which was incubated with the primary antibody at the recommended working concentration. AP-conjugated GAM secondary antibodies were used at a 1:3000 dilution for detection and signal was developed with AP development kit.
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- Western blot analysis of GAPDH was performed by loading 20 µg of A549 (lane1), MCF7 (lane2), U-87 MG (lane3) and HeLa (lane4) cell lysate using Novex®NuPAGE®4-12 % Bis-Tris gel (Product # NP0321BOX), XCell SureLock Electrophoresis System (Product # EI0002), Novex® Sharp Pre-Stained Protein Standard (LC5800), and iBlot® Dry Blotting System (IB21001). Proteins were transferred to a nitrocellulose membrane and blocked with 5 % skim milk for 1 hour at room temperature. GAPDH was detected at ~37 kDa using GAPDH Mouse Monoclonal Antibody (Product # A21994) at 0.1-0.2 µg/mL in 2.5 % skim milk at 4°C overnight on a rocking platform. Goat Anti-Mouse IgG - HRP Secondary Antibody (Product # 62-6520) at 1:4000 dilution was used and chemiluminescent detection was performed using Pierce™ ECL Western Blotting Substrate (Product # 32106).
Supportive validation
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- Immunofluorescence analysis of GAPDH was performed using 70% confluent log phase HeLa cells. The cells were fixed with 4% paraformaldehyde for 10 minutes, permeabilized with 0.1% Triton™ X-100 for 10 minutes, and blocked with 1% BSA for 1 hour at room temperature. The cells were labeled with GAPDH Polyclonal Antibody (Product # A21994) at 5µg/mL in 0.1% BSA and then labeled with Goat anti-Mouse IgG (H+L) Superclonal™ Secondary Antibody, Alexa Fluor® 488 conjugate (Product # A28175) at a dilution of 1:2000 for 45 minutes at room temperature(Panel a: green). Nuclei (Panel b: blue) were stained with SlowFade® Gold Antifade Mountant with DAPI (Product # S36938). F-actin (Panel c: red) was stained with Rhodamine Phalloidin (Product # R415, 1:300) at a dilution of 1:300 for 45 minutes at room temperature. Panel d represents the merged image showing cytoplasmic localization. Panel e shows no primary antibody control. The images were captured at 60X magnification.
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- Human HDFn cells were paraformaldehyde fixed (4%, 20 min) and Triton X-100 permeabilized (0.1%, 15 min). The cells were incubated with the antibody (3E8AD9, 0.5µg/mL) for 2h at room temperature or over night at 4°C. The secondary antibody was (red) Alexa Fluor® 594 goat anti-mouse IgG (H+L) used at a 1/1000 dilution for 1h. 10% Goat serum was used as the blocking agent for all blocking steps. DAPI was used to stain the cell nuclei (blue). Target protein locates mainly in cytoplasm.
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
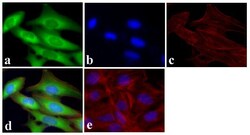
- Immunofluorescent analysis of GAPDH was done on 70% confluent log phase U2OS cells. The cells were fixed with 4% paraformaldehyde for 15 minutes, permeabilized with 0.25% Triton™ X-100 for 10 minutes, and blocked with 5% BSA for 1 hour at room temperature. The cells were labeled with GAPDH Mouse Monoclonal Antibody (Product # A21994) at 1 µg/mL and incubated for 3 hours at room temperature and then labeled with Alexa Fluor 488 Rabbit Anti-Mouse IgG Secondary Antibody (Product # A-11059) at a dilution of 1:400 for 30 minutes at room temperature (Panel a: green). Nuclei (Panel b: blue) were stained with SlowFade® Gold Antifade Mountant with DAPI (Product # S36938). F-actin (Panel c: red) was stained with Alexa Fluor 594 Phalloidin (Product # A12381). Panel d is a merged image showing cytoplasmic localization. Panel e shows no primary antibody control. The images were captured at 20X magnification.
Supportive validation
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
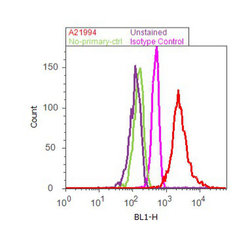
- Flow cytometry analysis of GAPDH was done on HeLa cells. Cells were fixed with 70% ethanol for 10 minutes, permeabilized with 0.25% Tritonª X-100 for 20 minutes, and blocked with 5% BSA for 30 minutes at room temperature. Cells were labeled with GAPDH Mouse Monoclonal Antibody (A21994, red histogram) or with mouse isotype control (pink histogram) at 3-5 µg/million cells in 2.5% BSA. After incubation at room temperature for 2 hours, the cells were labeled with Alexa Fluor¨ 488 Rabbit Anti-Mouse Secondary Antibody (A11059) at a dilution of 1:400 for 30 minutes at room temperature. The representative 10,000 cells were acquired and analyzed for each sample using an Attune¨ Acoustic Focusing Cytometer. The purple histogram represents unstained control cells and the green histogram represents no-primary-antibody control.
Supportive validation
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- Figure 3 delta-Opioid receptor (DOPr) overexpression and activation impairs keratinocyte differentiation. Cells were grown to confluence and differentiation induced by removal of growth factors under the influence of methionine 5 (Met)-enkephalin for up to 10 days. Differentiation marker gene expression was analyzed by quantitative real-time PCR. Data are represented after normalization to RPL13a expression as mean +-SEM ( n =3 keratin intermediate filament (KRT10), n =4 KRT1, n =5 involucrin (IVL), loricrin (LOR), and filaggrin (FLG)). Expression levels of ( a ) KRT10, ( b ) KRT1, ( c ) IVL, ( d ) LOR, and ( e ) FLG are shown after normalization to vehicle-treated control cells. Two-way analysis of variance, *** P
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
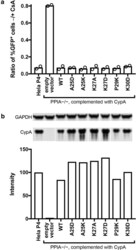
- Fig. 5 Functional analysis of CypA mutants in Hela-P4 cells. The PPIA gene was disrupted in Hela-P4 cells with CRISPR technology. The cloned cell line was transduced with a lentiviral vector encoding the indicated CypA mutant cDNAs. a Analysis of the extent of HIV-1 infection of the indicated cell lines by the A92E CA mutant in the presence and absence of CsA. A higher ratio indicates less enhancement of infection by CsA. Shown are the results of two independent determinations. b Quantitative analysis of CypA expression by immunoblotting relative to wild type HeLa-P4 cells. All samples were separated on the same gel and CypA and GAPDH were stained on the same blot membrane. CypA signals were normalized by the corresponding GAPDH signals
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- Fig. 6 Functional analysis of selected CypA mutants in Jurkat cells. Jurkat PPIA -/- cells were transduced with lent viral vectors encoding the indicated wild type (WT) and mutant CypA proteins containing the indicated amino acid substitutions. a Analysis of the extent of infection of the indicated cell lines by HIV-1 in the presence and absence of cyclosporin A (CsA). The results shown are from two independent assays. b Analysis of CypA expression by immunoblotting relative to wild type Jurkat cells. Signals were normalized by the corresponding GAPDH signals from the same blot. All samples were separated on the same gel and CypA and GAPDH were stained on the same blot membrane. CypA signals were normalized by the corresponding GAPDH signals
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
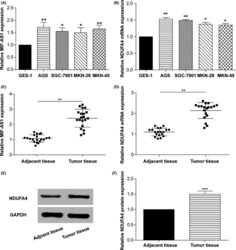
- Figure 2 Long non-coding (lnc) RNA MIF - AS 1 and NDUFA 4 expression levels were significantly upregulated in gastric cancer. A,B, qRT - PCR results showing the mRNA levels of (A) lnc MIF - AS 1 and (B) NDUFA 4 in normal human gastric epithelium cell line ( GES -1) and four human gastric cancer cell lines ( AGS , SGC -7901, MKN -28, MKN -45). * P < .05, ** P < .01. C,D, qRT - PCR was used to detect the expression level of (C) lnc MIF - AS 1 and (D) NDUFA 4 in gastric cancer and adjacent non-cancerous tissues. ** P < .01. E,F, Western blotting assay shows the protein levels of NDUFA 4 in gastric cancer and adjacent non-cancerous tissues. GAPDH was used as a loading control. *** P < .001
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- Figure 3 Silencing hsa_circRNA_002178 contributed to suppression in the proliferation, energy metabolism and angiogenesis of breast cancer cells. MDA-MB-231 cells were transfected with either si-hsa_circRNA_002178-1 or si-hsa_circRNA_002178-2 to silence its expression, with si-NC used as control. A, the expression of hsa_circRNA_002178 determined by RT-qPCR. B, the viability of breast cancer cells assessed by CCK-8. C, PCNA and Ki67 protein expression normalized to GAPDH measured by Western blot assay. D, ECAR and OCR levels in MDA-MB-231 cells measured by Seahorse XF24 analyzer. E, energy metabolism of breast cancer cells measured by HPLC. F, angiogenesis of MDA-MB-231 cells (x100). G, the tube length and the number of nodes. * P < .05 vs the si-NC group (MDA-MB-231 cells transfected with si-NC). The measurement data were expressed as mean +- standard deviation. Data among multiple groups were analysed using one-way ANOVA. The cell experiment was repeated three times
 Explore
Explore Validate
Validate Learn
Learn Western blot
Western blot Immunoprecipitation
Immunoprecipitation