Antibody data
- Antibody Data
- Antigen structure
- References [3]
- Comments [0]
- Validations
- Western blot [5]
- Immunocytochemistry [1]
- Other assay [2]
Submit
Validation data
Reference
Comment
Report error
- Product number
- MA5-27542 - Provider product page

- Provider
- Invitrogen Antibodies
- Product name
- SCD Monoclonal Antibody (CD.E10)
- Antibody type
- Monoclonal
- Antigen
- Recombinant full-length protein
- Reactivity
- Human, Mouse, Rat
- Host
- Mouse
- Isotype
- IgG
- Antibody clone number
- CD.E10
- Vial size
- 100 µg
- Concentration
- 1 mg/mL
- Storage
- -20° C, Avoid Freeze/Thaw Cycles
Submitted references Lipidomic Analysis of Archival Pathology Specimens Identifies Altered Lipid Signatures in Ovarian Clear Cell Carcinoma.
SCD Inhibition Protects from α-Synuclein-Induced Neurotoxicity But Is Toxic to Early Neuron Cultures.
Expression of SCD and FADS2 Is Lower in the Necrotic Core and Growing Tumor Area than in the Peritumoral Area of Glioblastoma Multiforme.
Mir SA, Wong SBJ, Narasimhan K, Esther CWL, Ji S, Burla B, Wenk MR, Tan DSP, Bendt AK
Metabolites 2021 Sep 3;11(9)
Metabolites 2021 Sep 3;11(9)
SCD Inhibition Protects from α-Synuclein-Induced Neurotoxicity But Is Toxic to Early Neuron Cultures.
Nicholatos JW, Groot J, Dhokai S, Tran D, Hrdlicka L, Carlile TM, Bennion M, Dalkilic-Liddle I, Hirst WD, Weihofen A
eNeuro 2021 Jul-Aug;8(4)
eNeuro 2021 Jul-Aug;8(4)
Expression of SCD and FADS2 Is Lower in the Necrotic Core and Growing Tumor Area than in the Peritumoral Area of Glioblastoma Multiforme.
Korbecki J, Kojder K, Jeżewski D, Simińska D, Tarnowski M, Kopytko P, Safranow K, Gutowska I, Goschorska M, Kolasa-Wołosiuk A, Wiszniewska B, Chlubek D, Baranowska-Bosiacka I
Biomolecules 2020 May 7;10(5)
Biomolecules 2020 May 7;10(5)
No comments: Submit comment
Supportive validation
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
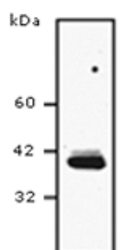
- Experimental details
- Western blot analysis of SCD1 in HEK293T cells. The samples were probed with SCD1 (Product # MA5-27542) monoclonal antibody.
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- Western blot analysis of SCD1 in HEK293T cells. The samples were probed with SCD1 (Product # MA5-27542) monoclonal antibody.
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- CRISPR-Cas9 mediated genome editing ofSCD (as confirmed by next generation sequencing) was achieved by using LentiArray™ Lentiviral sgRNA (Product # A32042, Assay ID CRISPR730414_LV) and LentiArray Cas9 Lentivirus (Product # A32064). Fig (a) Western blot analysis of SCD was performed by loading 30 µg of HeLa Cas9 (Lane 1) and HeLa Cas9 cells transduced with SCD Lentiviral sgRNA (Lane 2) whole cell extracts. The samples were electrophoresed using NuPAGE™ Novex™ 4-12% Bis-Tris Protein Gel (Product # NP0322BOX). Resolved proteins were then transferred onto a nitrocellulose membrane (Product # IB23001) by iBlot® 2 Dry Blotting System (Product # IB21001). The blot was probed with Anti-SCD Monoclonal Antibody (CD.E10) (Product # MA5-27542) using 1:1000 dilution and Goat anti-Mouse IgG (H+L) Superclonal™ Recombinant Secondary Antibody, HRP (Product # A28177 1:10000 dilution).Chemiluminescent detection was performed using SuperSignal™ West Dura Extended Duration Substrate (Product # 34076). A reduced signal in sgRNA transduced cells using the LentiArray™ CRISPR product line confirms that antibody is specific toSCD (Fig (b)). An uncharacterized band was observed in all the samples at ~57 kDa.
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- Knockdown of SCD was achieved by transfecting MCF-7 with SCD specific siRNAs (Silencer® select Product # s12504). Western blot analysis (Fig. a) was performed using whole cell extracts from the SCD knockdown cells (lane 3), non-specific scrambled siRNA transfected cells (lane 2) and untransfected cells (lane 1). The blot was probed with SCD Monoclonal Antibody (CD.E10) (Product # MA5-27542, 1:1000 dilution) and Goat anti-Rabbit IgG (H+L) Superclonal™ Recombinant Secondary Antibody, HRP (Product # A27036, 0.25 µg/mL, 1:4000 dilution). Densitometric analysis of this western blot is shown in histogram (Fig. b). Decrease in signal upon siRNA mediated knock down confirms that antibody is specific to SCD.
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- Western blot was performed using anti-SCD Monoclonal Antibody (CD.E10) (Product # MA5-27542) and 28, 37 kDa band corresponding to SCD was observed across cell lines tested. Whole cell extracts (30 µg lysate) of A-431 (Lane 1), MCF-7 (Lane 2), HepG2 (Lane 3) and HeLa (Lane 4) were electrophoresed using NuPAGE® 4-12 % Bis-Tris gel (Product # NP0322BOX). Resolved proteins were then transferred onto a nitrocellulose membrane (Product # IB23001) by iBlot® 2 Dry Blotting System (Product # IB21001). The blot was probed with the primary antibody (1:1000 dilution) and detected by chemiluminescence with Goat anti-Mouse IgG (H+L) Superclonal™ Recombinant Secondary Antibody, HRP (Product # A28177, 1:4000 dilution) using the iBright FL 1000 (Product # A32752). Chemiluminescent detection was performed using Novex® ECL Chemiluminescent Substrate Reagent Kit (Product # WP20005).
Supportive validation
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
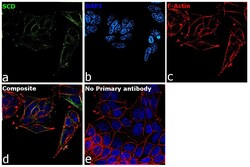
- Experimental details
- Immunofluorescence analysis of SCD was performed using 70% confluent log phase MCF-7 cells. The cells were fixed with 4% Paraformaldehyde for 10 minutes, permeabilized with 0.1% Triton™ X-100 for 10 minutes, and blocked with 2% BSA for 10 minutes at room temperature. The cells were labeled with SCD Monoclonal Antibody (CD.E10) (Product # MA5-27542) at 1:100 dilution in 0.1% BSA, incubated at 4 degree Celsius overnight and then labeled with Goat anti-Mouse IgG (H+L) Superclonal™ Recombinant Secondary Antibody, Alexa Fluor® 488 (Product # A28175, 1:2000 dilution) for 45 minutes at room temperature (Panel a: Green). Nuclei (Panel b: Blue) were stained with SlowFade® Gold Antifade Mountant with DAPI (Product # S36938). F-actin (Panel c: Red) was stained with Rhodamine Phalloidin (Product # R415, 1:300). Panel d represents the merged image showing ER like cytoplasmic localization. Panel e represents control cells with no primary antibody to assess background. The images were captured at 60X magnification.
Supportive validation
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- Figure 4 Representative microphotographs showing the protein expression of desaturase SCD in neurons and glia cells in the peritumoral area ( A ; red and white arrows) and in cells in the growing tumor area ( B ; blue arrows) necrotic core ( C , D ; blue arrows) of brain tumors from patients diagnosed with glioblastoma. Please note that on microphotography of GTA/NC, there are many pathologically changed regions of the brain tissue that indicate brain cell necrosis envelope by hypercellular zones (HcZ), with additional microvascular hyperplasia (green arrows). There are microphotographs from different objective (obj. magnification: A , C , D x20, B x40); scale bar are 200 um or 100 um.
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- Figure 3. High SCD inhibitor concentrations suppress alphaSyn 3K-GFP mRNA and protein in neuroblastoma model. A , Representative Licor western blot depicting alphaSyn 3K-GFP, betaactin, and SCD1 protein levels from samples treated with SCD inhibitors (0.01 to 10 muM), alphaSyn 3K-GFP induction and SCD inhibitor treatment was 48 hours before protein isolation. B , Quantification of alphaSyn 3K protein levels as in A , CAY10566 and MF-438 data was equivalent and merged for this plot. C , mRNA levels of alphaSyn 3K-GFP at equivalent time points and treatments as in C . Note the drop in alphaSyn 3K protein and mRNA under SCD inhibition >= 1 muM. D , Quantification of SCD1 protein levels under increasing concentrations of SCD inhibitor, note the dose dependent increase. Please see Extended Data Figure 3-1 for further SCD mRNA and protein characterization under inhibitor treatment. One-way ANOVA run with Dunnett's multiple test correction, all data displayed as boxplots, *** p < 0.001. All plots n >= 3 independent experiments.
 Explore
Explore Validate
Validate Learn
Learn Western blot
Western blot ELISA
ELISA