Antibody data
- Antibody Data
- Antigen structure
- References [0]
- Comments [0]
- Validations
- Western blot [1]
- Immunocytochemistry [1]
- Immunohistochemistry [3]
- Other assay [66]
Submit
Validation data
Reference
Comment
Report error
- Product number
- 39-8600 - Provider product page

- Provider
- Invitrogen Antibodies
- Product name
- GAPDH Monoclonal Antibody (ZG003)
- Antibody type
- Monoclonal
- Antigen
- Recombinant full-length protein
- Reactivity
- Human, Mouse
- Host
- Mouse
- Isotype
- IgG
- Antibody clone number
- ZG003
- Vial size
- 100 µg
- Concentration
- 0.5 mg/mL
- Storage
- -20°C
No comments: Submit comment
Supportive validation
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
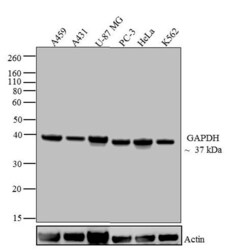
- Experimental details
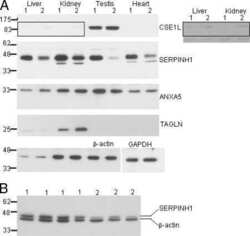
- Western blot analysis of GAPDH was performed by loading 20 µg of A549 (lane1), A431 (lane2), U-87 MG (lane3), Mouse Lung (lane4), PC-3 (lane5), HeLa (Lane6) and K562 (lane7) lysate using Novex®NuPAGE®4-12 % Bis-Tris gel (Product # NP0321BOX), XCell SureLock Electrophoresis System (Product # EI0002), Novex® Sharp Pre-Stained Protein Standard (LC5800), and iBlot® Dry Blotting System (IB21001). Proteins were transferred to a nitrocellulose membrane and blocked with 5 % skim milk for 1 hour at room temperature. GAPDH was detected at ~37 kDa using GAPDH Mouse Monoclonal Antibody (Product # 39-8600) at 0.5-1 µg/mL in 2.5 % skim milk at 4°C overnight on a rocking platform. Goat Anti-Mouse IgG - HRP Secondary Antibody (Product # 62-6520) at 1:4000 dilution was used and chemiluminescent detection was performed using Pierce™ ECL Western Blotting Substrate (Product # 32106).
Supportive validation
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
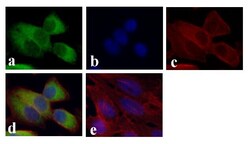
- Experimental details
- Immunofluorescent analysis of GAPDH was done on 70% confluent log phase HeLa cells. The cells were fixed with 4% paraformaldehyde for 15 minutes, permeabilized with 0.25% Triton™ X-100 for 10 minutes, and blocked with 5% BSA for 1 hour at room temperature. The cells were labeled with GAPDH Mouse Monoclonal Antibody (Product # 39-8600) at 1 µg/mL and incubated for 3 hours at room temperature and then labeled with Alexa Fluor 488 Rabbit Anti-Mouse IgG Secondary Antibody (Product # A-11059) at a dilution of 1:400 for 30 minutes at room temperature (Panel a: green). Nuclei (Panel b: blue) were stained with SlowFade® Gold Antifade Mountant with DAPI (Product # S36938). F-actin (Panel c: red) was stained with Alexa Fluor 594 Phalloidin (Product # A12381). Panel d is a merged image showing cytoplasmic localization. Panel e shows no primary antibody control. The images were captured at 20X magnification.
Supportive validation
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
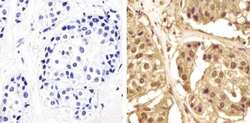
- Experimental details
- Immunohistochemistry analysis of GAPDH showing staining in the cytoplasm and nucleus of paraffin-embedded human breast carcinoma (right) compared to a negative control without primary antibody (left). To expose target proteins, antigen retrieval was performed using 10mM sodium citrate (pH 6.0), microwaved for 8-15 min. Following antigen retrieval, tissues were blocked in 3% H2O2-methanol for 15 min at room temperature, washed with ddH2O and PBS, and then probed with a GAPDH Mouse Monoclonal Antibody (ZG003) (Product # 39-8600) diluted in 3% BSA-PBS at a dilution of 1:100 overnight at 4°C in a humidified chamber. Tissues were washed extensively in PBST and detection was performed using an HRP-conjugated secondary antibody followed by colorimetric detection using a DAB kit. Tissues were counterstained with hematoxylin and dehydrated with ethanol and xylene to prep for mounting.
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
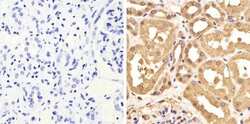
- Experimental details
- Immunohistochemistry analysis of GAPDH showing staining in the cytoplasm and nucleus of paraffin-embedded human kidney tissue (right) compared to a negative control without primary antibody (left). To expose target proteins, antigen retrieval was performed using 10mM sodium citrate (pH 6.0), microwaved for 8-15 min. Following antigen retrieval, tissues were blocked in 3% H2O2-methanol for 15 min at room temperature, washed with ddH2O and PBS, and then probed with a GAPDH Mouse Monoclonal Antibody (ZG003) (Product # 39-8600) diluted in 3% BSA-PBS at a dilution of 1:100 overnight at 4°C in a humidified chamber. Tissues were washed extensively in PBST and detection was performed using an HRP-conjugated secondary antibody followed by colorimetric detection using a DAB kit. Tissues were counterstained with hematoxylin and dehydrated with ethanol and xylene to prep for mounting.
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
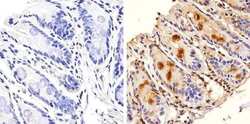
- Immunohistochemistry analysis of GAPDH showing nuclear and weak cytoplasm staining of paraffin-embedded mouse colon tissue (right) compared to a negative control without primary antibody (left). To expose target proteins, antigen retrieval was performed using 10mM sodium citrate (pH 6.0), microwaved for 8-15 min. Following antigen retrieval, tissues were blocked in 3% H2O2-methanol for 15 min at room temperature, washed with ddH2O and PBS, and then probed with a GAPDH Mouse Monoclonal Antibody (ZG003) (Product # 39-8600) diluted in 3% BSA-PBS at a dilution of 1:20 overnight at 4°C in a humidified chamber. Tissues were washed extensively in PBST and detection was performed using an HRP-conjugated secondary antibody followed by colorimetric detection using a DAB kit. Tissues were counterstained with hematoxylin and dehydrated with ethanol and xylene to prep for mounting.
Supportive validation
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
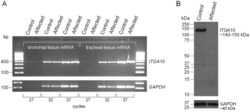
- Figure 7 ITGA10 expression on the RNA and protein level. (A) Semi-quantitative analysis of ITGA10 mRNA expression in bronchial and tracheal tissue samples of an affected NE and an unaffected Australian Kelpie dog. PCR reactions were performed using three cycle numbers, 27, 32 and 37. Amplification of mRNA fragments was roughly equal in both dogs, which indicated that the mutated transcript is stable and not targeted for nonsense mediated decay. (B) A western blot analysis of ITGA10 protein expression. A polyclonal anti-ITGA10 antibody was probed against the total protein lysates from tracheal tissue samples of the affected NE and the unaffected Australian Kelpie. The full-length ITGA10 protein was detected in the unaffected control dog but not in the affected dog. GAPDH was used as a loading control.
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
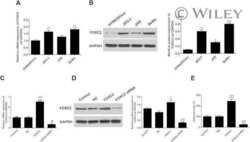
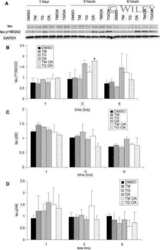
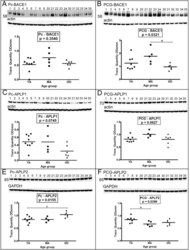
- Figure 2 Western blot analyses of BACE1 and APLP1 and APLP2. Sample numbers correspond to young adult (1-9), middle-aged (20-24) and oldest-old (30-35). Western blotting was used to detect BACE1 in the precuneus (A) and posterior cingulate gyrus (B), APLP1 in the precuneus (C) and posterior cingulate gyrus (D) and APLP2 in the precuneus (E) and posterior cingulate gyrus (F). All Western blots were loaded with a total of 40 ug of protein per lane. APLP2 blots were performed under non-reducing conditions. Data are reported in optical density units and were adjusted for actin (BACE1, APLP1) or GAPDH (APLP2). Actin and GAPDH loading probes are shown below each primary antibody blot. Abbreviations: Pc, precuneus; PCG, posterior cingulate gyrus; YA, young adult; MA, middle-aged; OO, oldest-old; BACE1, beta-site amyloid precursor protein-cleaving enzyme-1; APLP, amyloid precursor-like protein. For statistical treatment see legend to Figure 1 .
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
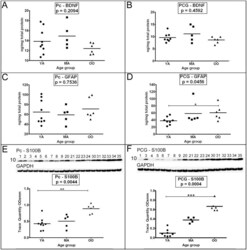
- Figure 6 ELISA quantitative analyses of BDNF and GFAP and Western blot analyses of S100B. As indicated in the figure both the precuneus (A, C and E) and posterior cingulate gyrus (B, D and F) were investigated. Sample numbers, shown above each Western blot correspond to young adult (1-9), middle-aged (20-24) and oldest-old (30-35). ELISA concentrations are reported in ng per mg of total protein. For the S100B Western blot, a total of 25 ug of total protein was loaded per lane. Data are reported in optical density units and were adjusted for GAPDH. The GAPDH loading probe is shown below the primary antibody blot. The molecular weights are shown on the left side of each blot. Abbreviations: Pc, precuneus; PCG, posterior cingulate gyrus; YA, young adult; MA, middle-aged; OO, oldest-old; BDNF, brain-derived neurotrophic factor; GFAP, glial fibrillary acidic protein; S100B, S100 calcium binding protein-B. For statistical analyses see legend to Figure 1 .
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
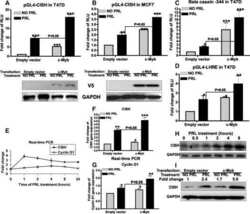
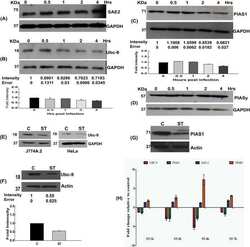
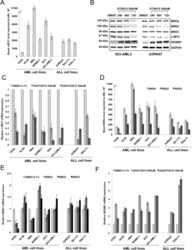
- Figure 3 c-MYC, BRD2/3/4 and HEXIM1 expression in AML and ALL cell lines after OTX015 treatment A. c-MYC basal gene expression in AML and ALL cell lines determined by RT-qPCR, relative to ABL 10 2 . Results are shown as mean +/- SEM from duplicates of three independent experiments. B. Western blot showing c-MYC, BRD2/3/4, and HEXIM1 protein changes in OCI-AML3 and JURKAT cells treated with 500nM OTX015 for 24, 48 or 72h or 0.1% DMSO. GAPDH was used as a loading control. One representative experiment out of three is shown. C. RT-qPCR showing c-MYC decrease in AML and ALL cell lines after 4 and 24h exposure with 500nM OTX015, relative to GAPDH and normalized to 0.1% DMSO. Results are shown as mean +/- SEM from duplicates of three independent experiments. D. RT-qPCR showing BRD4 , BRD2 , and BRD3 basal gene expression in leukemia cell lines, relative to ABL 10 2 . Results are shown as mean +/- SEM from duplicates of three independent experiments. E. RT-qPCR showing BRD4 , BRD2 , and BRD3 mRNA expression levels after 48h exposure to 500nM OTX015 in leukemia cell lines, relative to ABL and normalized to 0.1% DMSO. Results are shown as mean +/- SEM from duplicates of three independent experiments. F. RT-qPCR showing HEXIM1 mRNA increase in AML and ALL cell lines after 4h and 24h exposure with 500nM OTX015, relative to GAPDH and normalized to 0.1% DMSO. Results are shown as mean +/- SEM from duplicates of three independent experiments.
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
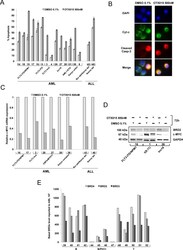
- Figure 4 Induction of apoptosis, expression of c-MYC and BRD2 following OTX015, and basal expression of BRD2/3/4 in AML and ALL patient samples A. Bone marrow mononuclear cells were exposed to 500nM OTX015 for 72h. Apoptotic cells were defined as Annexin V+ with or without PI uptake. Results are shown as mean +/- SEM. B. BM cells from a patient with MLL-rearranged AML ( MLL-AF9 ) (Patient 1, Table 2 ) showing cytochrome c (green), activated caspase-3 (red) and nuclei (blue). In non-apoptotic cells cytochrome c (green) shows dotted staining localized in the mitochondria while no activated caspase-3 could be detected, and in apoptotic cells cytochrome c is released into the cytosol (green) and activated caspase-3 is localized to the cytoplasm (red). Merged images of apoptotic cells appear in yellow. C. RT-qPCR showing c-MYC mRNA expression in nine AML and ALL patient samples after 72h exposure with 500nM OTX015 or 0.1% DMSO, relative to ABL normalized to 0.1% DMSO. D. Western blot showing BRD2, c-MYC and GAPDH expression in three AML patient samples exposed 72h to 500nM OTX015 or 0.1% DMSO ex vivo . E. RT-qPCR showing BRD4 , BRD2 , and BRD3 basal gene expression levels in 13 ALL patient samples, relative to ABL 10 2 .
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
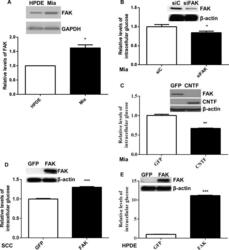
- Fig 2 FAK modulation of intrinsic glucose elevation A. The levels of FAK protein were assessed using Western blot analysis. The band intensity of total FAK (representative images, insets) was determined using Image-J and normalized to that of GAPDH. The relative levels of FAK in Miapaca-2 (Mia) were calculated and statistically analyzed. Data are averages with SEM from 6 biological replicates.*: p
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
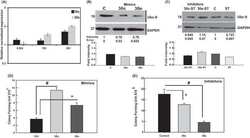
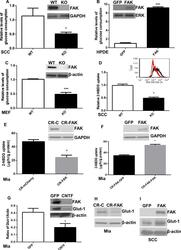
- Fig 3 FAK promotes glucose consumption and uptake A. Wild-type (WT) and FAK (KO) SCC cells were used as to examine the effects of FAK on glucose consumption. The levels of glucose in medium incubated with WT or FAK KO cells were determined. Glucose consumption was calculated (mug glucose/hr/mg protein). Then, the values were normalized to controls and shown as relative levels of glucose consumption. Data are averages with SEM from 6 biological replicates.*: p< 0.05 vs WT. B. HPDE cells were transfected with pGFP or pcFAK constructs, incubated for 56 hr, and subjected to glucose analysis. The amount of glucose used by pcFAK-transfected cells was normalized to cellular protein levels and then to that by pGFP-transfected cells. GFP: Cells expressing the GFP gene, and FAK: Cells expressing the mCherry-tagged FAK gene. Data are averages with SEM from 3 biological replicates. ****: p
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
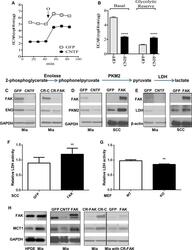
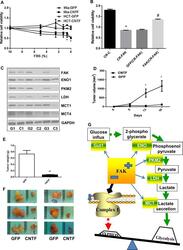
- Fig 5 FAK-promoted glycolysis is associated with increases in key regulatory proteins A. ECAR of GFP- and CNTF-transfected Miapaca-2 cells. Oligomycin A (O; 1mM) was administered to inhibit mitochondrial respiration and induce maximal glycolytic flux. B. Basal glycolysis decreases in cells expressing the CNTF gene. Glycolytic reserve: the difference between Oligomycin A induced ECAR and basal ECAR. Data are averages with SEM from 3 biological replicates. ****: p
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
- Fig 7 FAK-decreased mitochondrial respiration and complex I subunits A. OCR of Miapaca-2 cells expressing the GFP or CNTF gene. B. Basal OCR, ATP dependent OCR (difference between basal OCR and Oligomycin A (O) inhibited OCR), and maximal FCCP (F)-induced OCR. Data are averages with SEM from 3 biological replicates. **p
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
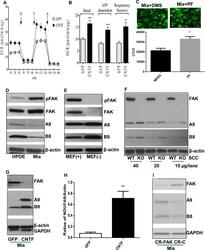
- Fig 8 Attenuation of FAK-promoted oncometabolism restores growth factor-dependency and reduces tumorigenicity A. CNTF inhibition of FAK expression re-sensitizes tumor cells to growth factor withdrawal-decreased cell viability. Miapaca-2 or HCT116 cells expressing the GFP or CNTF gene were cultured in medium containing 0-10% FBS for 24 hr and subjected to WST-1 cell viability assessed. O.D. values were normalized to that of cells in 10% FBS. Mia-GFP: Miapaca-2 cells transfected with the pGFP vector; Mia-CNTF: Miapaca-2 cells transfected with pcFAK constructs; HCT-GFP: HCT116 cells transfected with the pGFP vector; and HCT-CNTF: HCT116 cells transfected with pcFAK constructs. Data are averages with SEM from 3 replicates. *: p< 0.05 vs GFP. B. CRISPR-Cas9 FAK and pcFAK-transfected Miapaca-2 cells were cultured under growth factor and anchorage-limited conditions for 24 hr and subjected to cell viability assay using WST-1. OD values of the cells incubated for 24 hr was normalized to those of respective cells kept for 0 hr. CR-C: Cells expressing the mCherry gene, CR-FAK: Cells transfected with the CRISPR-Cas9 vector targeting the FAK gene, GFP (CR-FAK): FAK deficient cells transfected with vehicles (pGFP), and FAK (CR-FAK): FAK deficient cells transfected with pcFAK vectors. Data are averages with SEM from 3 replicates. *: p< 0.05 vs CR-C, and #: p
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
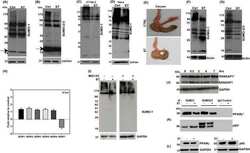
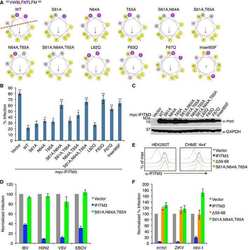
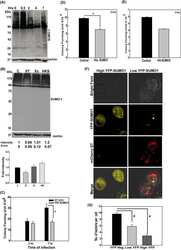
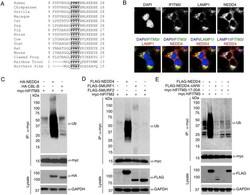
- Fig 1 IFITM3 is ubiquitinated by NEDD4. A) Alignment of IFITM3 N-terminal amino acids from various species. Bold and underlined text highlights the conserved PPxY motif. B) Mouse embryonic fibroblasts (MEFs) were stimulated overnight with IFN-alpha (160 units/mL) to ensure production of IFITM3, and imaged by fluorescent confocal microscopy with staining for endogenous IFITM3, NEDD4, LAMP1, and nuclei (DAPI). Images were taken with a 60x objective and 2.5x zoom. Pseudocolored merged images in different staining combinations are shown. C-E), HEK293T cells were co-transfected with plasmids expressing IFITM3 and epitope tagged ubiquitin ligases, NEDD4, CLB-B, SMURF1 and SMURF2, as indicated. Cell lysates were immunoprecipited with anti-myc resin, and examined by Western blotting with anti-myc and anti-ubiquitin (Ub) antibodies. Western blots of cell lysates with anti-HA (C) or anti-FLAG (D,E) antibodies were performed to confirm expression of the ubiquitin ligases. Anti-GAPDH Western blotting was performed to confirm comparable protein loading.
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
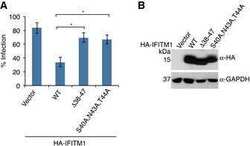
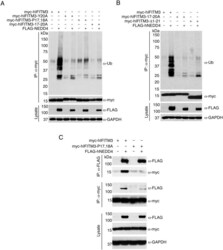
- Fig 2 The IFITM3 PPxY motif is required for ubiquitination by NEDD4. A-C) HEK293T cells were co-transfected with plasmids expressing myc-hIFITM3 or FLAG-NEDD4 as indicated. A-B) Cell lysates were immunoprecipitated with anti-myc resin, and examined by Western blotting with anti-myc and anti-ubiquitin (Ub). Western blotting of cell lysate with anti-FLAG antibodies was performed to confirm expression of NEDD4. Western blotting with anti-GAPDH antibodies was performed to confirm comparable protein loading. C) Cell lysates were immunoprecipitated with anti-myc or anti-FLAG resin, and co-immunoprecipitation was examined by Western blotting with both anti-myc and anti-FLAG antibodies for each immunoprecipitate. Western blots of cell lysates with anti-myc and anti-FLAG antibodies were performed to confirm expression of IFITM3 and NEDD4, respectively. Anti-GAPDH Western blotting was performed to confirm comparable protein loading.
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- Fig 3 NEDD4 catalytic activity is required for IFITM3 ubiquitination. HEK293T cells were transfected with the indicated mouse or human IFITM3 constructs and were co-transfected with plasmids expressing HA-NEDD4 or a catalytically inactive HA-NEDD4-C867A mutant. IFITM3 was immunoprecipitated with anti-myc resin and subjected to anti-myc and anti-ubiquitin (Ub) Western blotting. Cell lysates were probed with anti-NEDD4 antibodies to confirm expression of NEDD4 constructs. Anti-GAPDH staining served as a protein loading control.
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
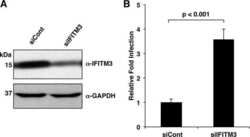
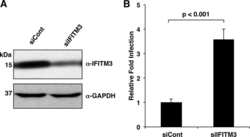
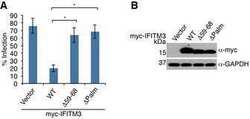
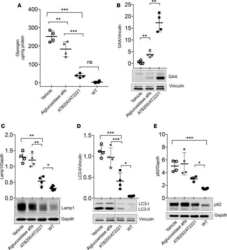
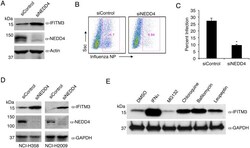
- Fig 7 NEDD4 knockdown in human lung cells increases IFITM3 levels and resistance to influenza virus infection. A-C) A549 cells were transfected for 48 h with control siRNA (siControl) or siRNA targeting human NEDD4 (siNEDD4). A) Cells were collected just prior to infection for confirmation of NEDD4 knockdown by anti-NEDD4 Western blotting, with anti-actin blotting serving as a loading control. Anti-IFITM3 blotting demonstrates an increase in IFITM3 upon NEDD4 knockdown. B,C) Following siRNA treatment, cells were infected with influenza virus strain PR8 at an MOI of 2.5 for 6 h. Cells were then fixed and stained with anti-influenza virus NP to measure the percentage of cells infected using flow cytometry. Results shown are representative of three independent experiments, with samples run in triplicate. Error bars represent standard deviation of triplicate samples. * Indicates a p-value less than 0.0001 calculated with Student's t-test. D) NCI-H358 and NCI-H2009 cells were transfected for 48 h with siControl or siNEDD4. Cell lysates were subjected to immunoblotting with anti-NEDD4 to confirm NEDD4 knockdown, anti-IFITM3 to demonstrate increase in endogenous IFITM3 upon NEDD4 knockdown, and anti-GAPDH as a loading control. E) A549 cells were treated with equal volumes Dimethyl Sulfoxide (DMSO) as a control, MG132 (10 muM), Chloroquine (10 muM), Bafilomycin A1 (1 muM), or Leupeptin (100 muM) for 24 h. Cells were also treated with IFN-alpha (100 units/mL) for comparison. Cell
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
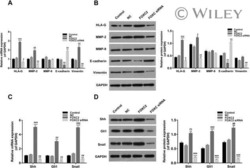
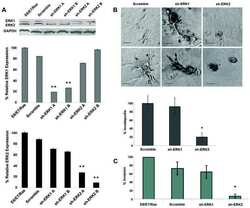
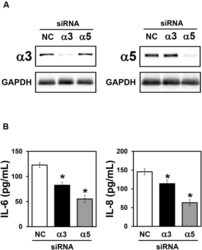
- FIGURE 3 Effect of alpha3 or alpha5 integrin silencing on IL-6 and IL-8 secretion by A549 cells during interaction with H. capsulatum. A549 cells were transfected with alpha3 or alpha5 integrin-directed siRNA or with Negative Control (NC) siRNA for 24 h, and then, incubated with H. capsulatum yeasts for 16 h. After incubation with fungi, culture supernatants were collected for determination of IL-6 and IL-8 levels, and A549 cells were harvested, lysed, and analyzed by Western blot. (A) Silencing of alpha3 and alpha5 integrins was confirmed by Western blot. GAPDH was used as protein loading control. Blots are representative of three independent experiments. (B) IL-6 and IL-8 levels in culture supernatants were determined by ELISA. Values represent the mean of triplicate experiments +- the standard deviation. * p < 0.01 when compared to NC siRNA. Similar results were obtained from three independent experiments.
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
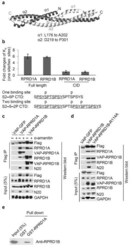
- Figure 4 RPRD1A and RPRD1B dimerization through their coiled-coil domains. ( a ) Crystal structure of RPRD1B C-terminal coiled-coil domain. The two interacting polypeptides are shown in darker and lighter shades. alpha: alpha-helix. ( b ) ITC measurement of the binding affinity (K d ) between full length RPRD1A and RPRD1B or their CIDs with CTD peptides bearing either one or two CID-binding site(s). Ratios of ITC measurements for the interactions of the indicated RPRDs with the two indicated CTD peptides are shown. Underlined are the CID-binding sequences in the CTD peptides. Phosphorylated serines are labeled. Bars indicate ranges of two technical replicates. ( c ) IP-WB analysis using the indicated antibodies showing the interaction between lentiviral transduced VAP-RPRD1A and VAP-RPRD1B in HEK293 cells grown in the presence or absence of 2 mug/ml alpha-amanitin for 72 hr. VAP: versatile affinity purification tag. ( d ) IP-WB analysis showing the effect of the R114 mutation on the interaction between lentiviral transduced VAP-RPRD1B and RPRD1A in HEK293 cells. ( f ) Glutathione-S-transferase (GST) pull-down experiment showing a direct interaction between GST-tagged RPRD1A and His-tagged RPRD1B purified from E. coli . Uncropped images of gels in Fig. 4c-e are shown in Supplementary Data Set 1 .
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
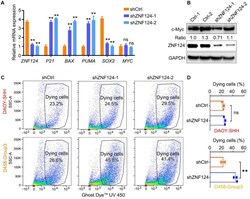
- FIGURE 6 Knockdown ZNF124 promotes apoptosis of G3-MB cells. (A) RT-PCR data showed the efficiency of shZNF124 in MYC amplificated G3-MB D458 cells and the cell cycle inhibitor gene CDKN1A/P21, apoptosis genes BAX and PUMA and tumor stemness gene MYC and SOX2 mRNA expression. Data represent means +- SD, n = 3 independent experiments. ** p < 0.01, two-tailed unpaired t -test. (B) Western blotting analysis of MYC and ZNF124 protein expression in human MB D458 cell after ZNF124 knockdown with Lenti-shRNA for 72 h. Ratio of MYC expression over GAPDH control was shown. (C,D) Dying cells were detected by flow cytometry in G3-D458 and SHH-DAOY cells after knockdown ZNF124 (C) and the comparison of the dead cells in control and shZNF124 treatment cells (D) . Data represent means +- SD, n = 3 independent experiments. ** p < 0.01, two-tailed unpaired t -test.
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
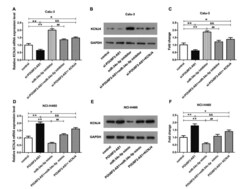
- Figure 3 POU6F2-AS1 served as a ceRNA via sponging miR-34c-5p to elevate KCNJ4. (A) The mRNA and (B) protein expression level of KCNJ4 in Calu-3 cells were demonstrated. (C) The protein expression level of KCNJ4 was quantified. (D) The mRNA and (E) protein expression level of KCNJ4 in NCI-H460 cells were detected. (F) The quantification of (H). *P < 0.05, **P < 0.01, ## P < 0.01 and && P < 0.01.
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
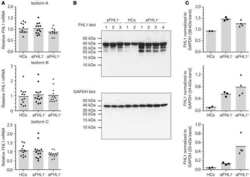
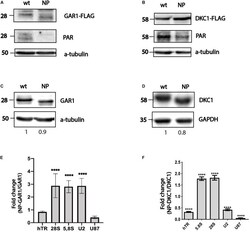
- FIGURE 1 PARylation regulates ribonucleoprotein complex stability. Substitution of amino acid residues at sites proposed for PARylation influences the PAR-attachment to GAR1 (A) and DKC1 (B) mobility of the GAR1 (A,C) and DKC1 (B,D) proteins. Differential binding of RNA to GAR1 and NP-GAR1 (E) and to DKC1 and NP-DKC1 (F) , as revealed by RT-qPCR analysis. The mean values were calculated from triplicate RT-qPCR experiments with three biological replicates, and the bars represent SE. **** P < 0.0001 by Sidak's multiple comparisons test.
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
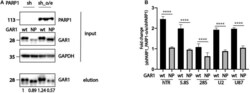
- FIGURE 3 PARylation modulates the ability of RNA-binding proteins to associate with RNA. (A) Western blot analysis of expression of PARP1, GAR1, and NP-GAR1 (input panel) and efficiency of immunoprecipitation (elution panel). (B) RT-qPCR analysis of the levels of RNA co-immunoprecipitated with GAR1. The mean values were calculated from triplicate RT-qPCR experiments with three biological replicates, and the bars represent SE. **** P < 0.0001.
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
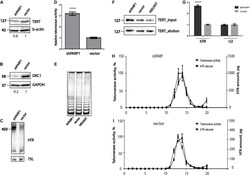
- FIGURE 4 PARP1 is involved in the regulation of telomerase complex composition and stability. (A) Western blot analysis of TERT protein levels in the indicated cell lines. (B) Western blot analysis of DKC1 protein levels in the indicated cell lines. (C) Northern blot analysis of hTR expression in the indicated cell lines. (D) RQ-TRAP telomerase activity analysis in the indicated cell lines. **** indicates unpaired t -test two tailed p value < 0,0001. (E) Analysis of telomerase processivity using TRAP assay followed by PAGE separation of PCR products. (F) Immunoprecipitation of hTERT from the indicated cell lines was followed by immunoblotting with anti-hTERT antibodies. (G) RT-qPCR analysis of the amounts of hTR co-immunoprecipitated with hTERT. U2 RNA was used as a control. **** indicates unpaired t -test two tailed p value < 0,0001. (H) Analysis of hTR distribution and telomerase activity after separation of extracts from HEK293T cells expressing a shRNA targeting PARP1 mRNA, using a sucrose gradient. (I) Analysis of hTR distribution and telomerase activity after separation of extract from HEK293T cells expressing the empty LeGo-Cer vector, using a sucrose gradient.
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
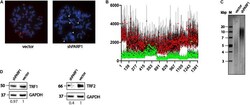
- FIGURE 5 PARP1 is involved in the regulation of telomere length and telomerase activity. (A) Immunofluorescence-FISH analysis of metaphase spreads from cells expressing the LeGo-Cer vector and a shRNA specific to PARP1 mRNA. White arrows point defects in telomeric structures. (B) Telomere length analysis performed by Telometer software by the Johns Hopkins University. Green dots correspond control cells (vector) and red dots correspond spreads from cells with decreased level of PARP1 (shPARP1). (C) Telomere restriction fragment length analysis of PARP1 knockdown cells. (D) Western blot analysis of TRF1 protein levels in the indicated cell lines. (E) Western blot analysis of TRF2 protein levels in the indicated cell lines.
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
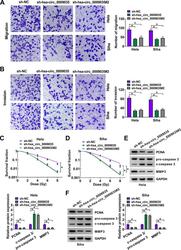
- FIG 4 hsa_circ_0009035 silencing regulated CC cell migration, invasion, and radiosensitivity in vitro . HeLa and Siha cells were stably transduced with sh-NC, sh-hsa_circ_0009035, or sh-hsa_circ_0009035#2. (A and B) Cell migration and invasion by transwell assay. (C and D) Survival analysis by colony formation assay in transduced cells upon radiation (0, 2, 4, 6, and 8 Gy) exposure. (E and F) Protein levels of PCNA, c-caspase 3, procaspase 3, and MMP3 determined by Western blotting in transduced cells. * , P < 0.05.
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
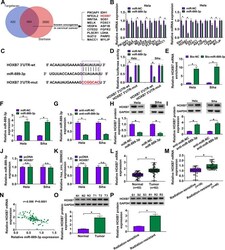
- FIG 7 HOXB7 was a direct target of miR-889-3p in CC cells. (A) Venn diagram showing the putative targets of miR-889-3p predicted by Targetscan and Starbase softwares. (B) qRT-PCR analysis of genes in HeLa cells transfected with the miR-NC mimic or miR-889-3p mimic. (C) Schematic of the target sequence for miR-889-3p identified by Starbase software and the mutant of the seed region. (D) Dual-luciferase reporter assays in both HeLa and Siha cells. (E) RNA pulldown assays in HeLa and Siha cells using Bio-NC or Bio-miR-889-3p. Relative miR-889-3p expression by qRT-PCR in cells transfected with an miR-NC mimic, an miR-889-3p mimic (F), anti-miR-NC, or anti-miR-889-3p (G). HOXB7 protein level determined by Western blotting in cells transfected with an miR-NC mimic, an miR-889-3p mimic (H), anti-miR-NC, or anti-miR-889-3p (I). (J and K) qRT-PCR analysis of miR-889-3p and hsa_circ_0009035 expression levels in cells transfected with a negative-control plasmid (pcDNA) or a HOXB7-overexpressing plasmid (HOXB7). Relative HOXB7 expression determined by qRT-PCR in 82 pairs of CC tissues and adjacent normal tissues (L), CC tissues from 36 primary patients (defined as radiation-sensitive CC) and 46 recurrent patients after radiation treatment (defined as radiation-resistant CC) (M). (N) Correlation between HOXB7 mRNA and miR-889-3p expression levels in CC tissues using the Spearman test. HOXB7 protein expression in 3 pairs of CC tissues and adjacent normal tissues (O), CC tissues from 3 prim
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
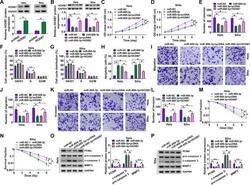
- FIG 8 The effects of miR-889-3p overexpression on CC progression and radiosensitivity in vitro were mediated by HOXB7. (A) Relative HOXB7 protein level determined by Western blotting in cells transfected with a negative-control plasmid (pcDNA) or a HOXB7-overexpressing plasmid (HOXB7). HeLa and Siha cells were transfected with an miR-NC mimic, an miR-899-3p mimic, an miR-899-3p mimic plus pcDNA, or an miR-899-3p mimic plus HOXB7, followed by the determination of HOXB7 protein level by Western blotting (B), cell proliferation by CCK-8 assay (C and D), analysis of cell colony formation by colony formation assay (E), analysis of cell cycle progression and apoptosis by flow cytometry (F to H), analysis of cell migration and invasion by transwell assay (I to L), determination of cell survival fraction by colony formation upon radiation exposure (M and N), and measurement of the levels of PCNA, c-caspase 3, procaspase 3, and MMP3 by Western blotting (O and P). * , P < 0.05.
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
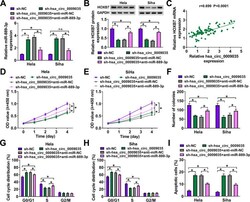
- FIG 9 hsa_circ_0009035 modulated HOXB7 expression and CC progression in vitro by miR-889-3p. Expression levels of miR-889-3p (A) and HOXB7 protein (B) in sh-NC-infected or sh-hsa_circ_0009035-transduced cells transfected with or without anti-miR-NC or anti-miR-889-3p. (C) Correlation between HOXB7 mRNA and hsa_circ_0009035 expression levels in CC tissues using the Spearman test. sh-NC-infected or sh-hsa_circ_0009035-transduced HeLa and Siha cells were transfected with or without anti-miR-NC or anti-miR-889-3p, followed by the assessment of cell proliferation by CCK-8 assay (D and E), cell colony formation by colony formation assay (F), and cell cycle progression and apoptosis by flow cytometry (G to I). * , P < 0.05.
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
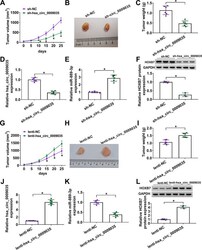
- FIG 11 hsa_circ_0009035 regulated tumor growth in vivo . (A) Growth curves of the xenograft tumors formed by sh-NC-infected or sh-hsa_circ_0009035-transduced HeLa cells ( n = 6 per group). Representative images (B), tumor average weight (C), hsa_circ_0009035 (D) and miR-889-3p (E) levels determined by qRT-PCR, and HOXB7 protein expression determined by Western blotting (F) of the xenograft tumors formed by HeLa cells infected with sh-NC or sh-hsa_circ_0009035, on day 25 after subcutaneous injection ( n = 6 per group). (G) Growth curves of the xenograft tumors formed by lenti-NC-infected or lenti-hsa_circ_0009035-transduced HeLa cells ( n = 6 per group). Representative images (H), tumor average weight (I), hsa_circ_0009035 (J), and miR-889-3p (K) levels determined by qRT-PCR, and HOXB7 protein expression determined by Western blotting (L) of the xenograft tumors formed by HeLa cells infected with lenti-NC or lenti-hsa_circ_0009035, on day 25 after subcutaneous injection ( n = 6 per group). * , P < 0.05.
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
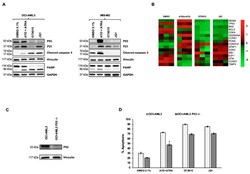
- Figure 2 Treatment with OTX015 (MK-8628) and JQ1 induces P53-independent apoptosis in OCI-AML3 cells. ( A ) Western blot showing P53, P21, caspase 3, and PARP protein changes in OCI-AML3 or IMS-M2 cells treated with similar doses of indicated drugs for 72 h. Vinculin was used as loading control for P53, P21, and caspase 3 and GAPDH for PARP. One representative experiment out of three is shown for WB experiments. Experiments were performed as triplicates. ( B ) Heatmap displaying expression of genes related to the TP53 pathway in OCI-AML3 cells treated for 72 h with 0.1% DMSO, 1000 nM ATO + 1000 nM ATRA, 500 nM OTX015 and 500 nM JQ1. The experiment was conducted on biological triplicates. ( C ) OCI-AML3 cells were transfected by electroporation with Alt-R CRISPR/Cas9 RNP. Knock out of TP53 was controlled with Western blot detecting P53 protein for indicated cells. Vinculin was used as loading control. One representative experiment out of three is shown. ( D ) OCI-AML3 knocked out for TP53 by CRISPR was exposed to 500 nM OTX015 (MK-8628), 500 nM JQ1, 1000 nM ATO + 1000 nM ATRA, or 0.1% DMSO for 72 h. Apoptosis was detected with annexin V and PI by using flow cytometry. Results are shown as mean +- SEM from duplicates of three independent experiments. Statistically significant differences were calculated from medians compared by the Mann-Whitney U test. * represents p
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
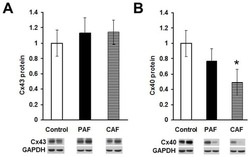
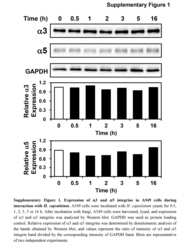
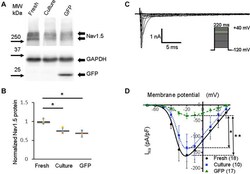
- FIGURE 3 Sodium channel protein (Nav1.5) and sodium current ( I Na ) in 3wRbCMs. (A) Western blot of Nav1.5, tetrodotoxin-resistant voltage-gated sodium channel subunit alpha 5 in freshly isolated (fresh), 48 h cultured, non-transduced (culture), and 48 h cultured, Green Fluorescent Protein (GFP)-transduced cells. For illustrative purposes (same protein samples), the images for GAPDH and GFP are reused in Figure 7A (Kv11.1). (B) Quantification of the western blots ( N = 4) shows significant downregulation of total Nav1.5 protein during 48 h of culturing with and without GFP transduction compared to freshly isolated cells. (C) Representative I Na traces of a freshly isolated 3wRbCM. The voltage clamp protocol is shown in the insert. (D) Cumulative I-V curves of freshly isolated cells (black circles, N = 4, n = 18), 48 h cultured, non-transduced cells (blue squares, N = 4, n = 10), 48 h cultured GFP-transduced cells (green triangles, N = 4, n = 17). The symbols * and ** correspond to p < 0.05 and p < 0.01, respectively.
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
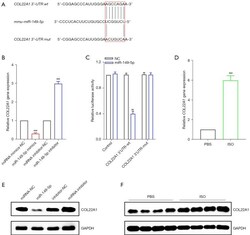
- Figure 5 COL22A1 is the target of miR-149-5p in HL-1 cells. (A) The predictive binding sequences between miR-149-5p and COL22A1. (B,C) The qRT-PCR and western blot assays showed that the overexpression of miR-149-5p significantly suppressed COL22A1 gene and protein expression, while the inhibition of miR-149-5p resulted in the upregulation of COL22A1 both in gene and protein as compared to the miRNA mimics and inhibitor NCs; U6 and GAPDH were used as the respective internal controls for each. (D) Luciferase assays were applied to confirm the binding between miR-149-5p and the 3'-UTR of COL22A1. Luciferase activity was measured. (E,F) qRT-PCR and western blot assays were performed to examine the mRNA and protein expression of COL22A1 in the mouse hearts induced by ISO or control; GAPDH was used as the internal control. ## , P
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
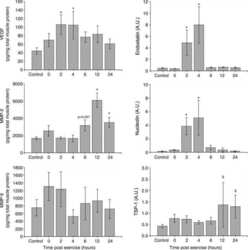
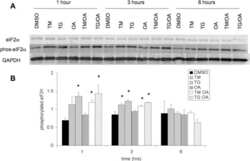
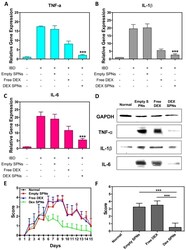
- Figure 6 Scoring of intestinal bleeding/ Histological change and ex vivo gene/ protein expression. A. TNF-alpha, B. IL-1beta, C. IL-6, after three consecutive treatment sessions with Normal, IBD control, Empty SPNs, Free DEX and DEX-SPNs. Results are presented as mean +- SD (n=3/group). *** indicates P < 0.001, DEX SPNs vs IBD control, Empty SPNs, Free DEX, or Normal. D. Protein levels of TNF-alpha, IL-1beta, and IL-6 within the intestinal tissue in DSS-induced mouse models of colitis. TNF-alpha (17 kDa), IL-1beta (17 kDa), and IL-6 (24 kDa) bands are shown. GAPDH is used as the internal control (40 kDa). E. Intestinal bleeding score. F. histology score for the four experimental groups (normal, Empty SPNs, Free DEX and DEX SPNs). Results are presented as mean +- SD (n=3/group). *** indicates P < 0.001.
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
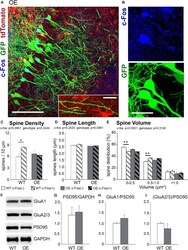
- FIGURE 3 The increase in spine density on c-Fos(+) neurons compared to c-Fos(-) neurons is impaired in OE mice. (A) Confocal images of the CA1 neurons expressing GFP (green) and astrocytes expressing tdTomato (red). Some neurons show c-Fos immunoreactivity (blue), scale bar is 100 mum. High magnification image shows example of dendritic spines located in a close proximity to tdTomato-expressing astrocytes, scale bar is 20 mum (insert). (B) High magnification images of c-Fos(+) (blue) and GFP-expressing (green) CA1 pyramidal neurons. (C-E) Graphs show the average number of dendritic spines per 10 mum dendrite (C) , spine length (D) , and spine volume (E) in c-Fos(+) and c-Fos(-) neurons from WT and OE mice. (C) There was an increased dendritic spine density in WT c-Fos(+) neurons compared with WT c-Fos(-) neurons [two-way ANOVA, c-Fos F (1,41) = 2.920, p = 0.0951; genotype F (1,41) = 0.995, p = 0.3244; interaction F (1,41) = 4.787, p = 0.0344; Bonferroni's post hoc * p < 0.05 c-Fos(+) WT vs. c-Fos(-) WT]. (D) Spine length was no different between c-Fos(-) and c-Fos(+) neurons in both WT and OE mice. (E) A significant decrease in the percentage of smaller spines (
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
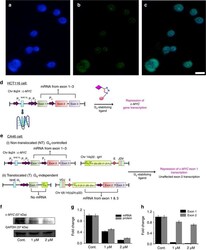
- Figure 7 Biological validation of Tz 1 as c-MYC G 4 stabilizer in cancer cells. Fluorescence microscopic images of HCT116 cells incubated with Tz 1 and DAPI, Scale bar, 10 mum; cells stained with ( a ) DAPI and ( b ) Tz 1 and ( c ) co-localization of Tz 1 and DAPI. ( d ) Schematic representation of G-quadruplex-mediated control over c-MYC expression in HCT116 cancer cell line. ( e ) A reciprocal translocation in the CA46 Burkitt's lymphoma cell line between the IgH heavy chain gene of chromosome 14 and the MYC gene on chromosome 8--NT mRNA expression is under the control of G-quadruplex forming region and T mRNA expression is independent of G-quadruplex. The NHE III 1 region moved along with exon 1 produces no mRNA product; therefore represents expression form NT allele; whereas, exon 2 mRNAs are produced from both T and NT allele. ( f ) Immunoreactive bands of c-MYC and GAPDH protein expression as analysed by Western blot. ( g ) Concentration dependent downregulation of c-MYC mRNA and protein levels in HCT116 cells after treatment with compound Tz 1 for 24 h. ( h ) Exon-specific qRT-PCR assay with the CA46 Burkitt's lymphoma cell line after 24 h treatment with Tz 1 . ( n =3, +-s.e.m.)
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
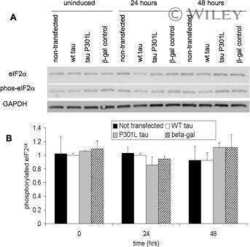
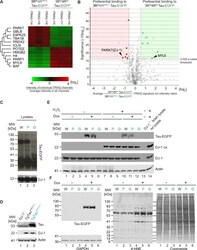
- Figure 5 Preferential binding of DJ-1 to mutant 4R P301L Tau, relative to 4R wt Tau, in IMR cells. ( A ) Hierarchical clustering analysis of proteins that exhibited preferential binding to 4R P301L or 4R wt Tau. ( B ) Orthogonal confirmation of proteins shortlisted in panel 'A' by volcano plot (ANOVA, p = 0.025). Red and green background shading signify preferential binding to 3R wt /4R P301L Tau and 3R wt /4R wt Tau, respectively (fold change greater than 50%). ( C ) Western blot documenting that the 12 hours induction of wild-type or P301L mutant Tau-EGFP fusion proteins did not alter steady-state levels of DJ-1. Note that in this and subsequent western blot panels samples derived from wild-type human Tau-EGFP expressing cells, P301L mutant Tau-EGFP expressing cells, and EGFP expressing cells are labeled as 'W in black font', 'P' in blue font and 'G' in green font, respectively. ( D ) Similarly, transgenic overexpression of P301L mutant Tau had no apparent effect on DJ-1 expression levels in 2-year-old mice. Specificity of the anti-DJ-1 antibody was validated with age-matched brain extracts from a DJ-1 -/- mouse. ( E ) Hydrogen peroxide (H 2 O 2 ) treated IMR cells expressing wild-type or P301L mutant Tau-EGFP fusion proteins exhibited no differences in total DJ-1 levels or of its oxidation at residue C106. As negative and positive controls for oxidized DJ-1 served brain lysates from DJ-1 -/- and wild-type mice, respectively, that were subjected to 1 mM H 2 O 2 for 10 min a
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
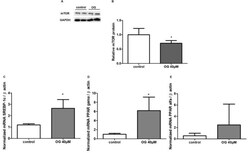
- Figure 3 Expression of mTOR protein in HepG2 cells after 24 h of treatment with OG by Western blot analysis (A). GAPDH was used to normalize protein expression. Results are expressed as protein levels (fold induction) of the mTOR expression. Treatment with OG decreased the expression of mTOR (B) in HepG2 cells (*p
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
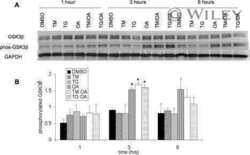
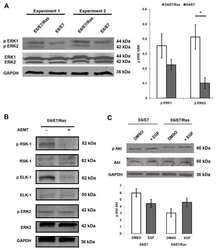
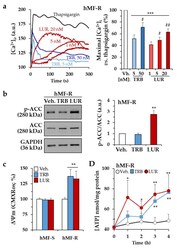
- Figure 3 Time-lapse effects of TRB and LUR in hMF intracellular Ca +2 signaling and mitochondrial membrane potential. ( a ) Intracellular Ca +2 concentration in hMF-R after thapsigargin (200 nM), TRB, or LUR treatment; ( b ) p-ACC/ACC ratio after 30 min of treatment with TRB (50 nM) and LUR (20 nM); ( c ) hMF-R and hMF-S mitochondrial membrane potential after 24 h of treatment with TRB (5 nM) and LUR (1 nM); ( d ) intracellular ATP levels in hMF treated with TRB (5 nM) and LUR (1 nM). Results show a representative blot out of three ( b ) and the means + S.E.M. from 8 different donors, * p < 0.05; ** p < 0.01; *** p < 0.001 vs. the vehicle condition; # p < 0.05; ## p < 0.01; vs. the corresponding condition with TRB (5 nM) or LUR (1 nM), respectively. a.u., arbitrary units.
 Explore
Explore Validate
Validate Learn
Learn Western blot
Western blot ELISA
ELISA