Antibody data
- Antibody Data
- Antigen structure
- References [0]
- Comments [0]
- Validations
- Western blot [1]
- Immunocytochemistry [2]
- Other assay [9]
Submit
Validation data
Reference
Comment
Report error
- Product number
- AM4300 - Provider product page

- Provider
- Invitrogen Antibodies
- Product name
- GAPDH Monoclonal Antibody (6C5)
- Antibody type
- Monoclonal
- Antigen
- Purifed from natural sources
- Reactivity
- Human, Mouse, Rat, Canine, Chicken/Avian, Rabbit
- Host
- Mouse
- Isotype
- IgG
- Antibody clone number
- 6C5
- Vial size
- 100 µg
- Storage
- Maintain refrigerated at 2-8°C for up to 1 month. For long term storage store at -20°C
No comments: Submit comment
Supportive validation
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
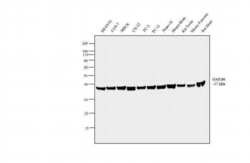
- Western blot analysis was performed on whole cell extracts (30µg lysate) of SH-SY5Y (Lane 1), COS-7 (Lane 2), MDCK (Lane 3), C2C12 (Lane 4), PC-3 (Lane 5), PC-12 (Lane 6), Neuro-2a (Lane 7), tissue extracts of Mouse Brain (Lane 8), Rat Testis (Lane 9), Mouse Pancreas (Lane 10) and Rat Heart (Lane 11). The blot was probed with Anti-GAPDH Mouse Monoclonal Antibody (Product # AM4300, 2µg/mL) and detected by chemiluminescence using Goat anti-Mouse IgG (H+L) Superclonal™ Secondary Antibody, HRP conjugate (Product # A28177, 0.25µg/mL, 1:500 dilution). A 37 kDa band corresponding to GAPDH was observed across the cell lines and tissues tested. Known quantity of protein samples were electrophoresed using Novex® NuPAGE®12 % Bis-Tris gel (Product # NP0342BOX), XCell SureLock™ Electrophoresis System (Product # EI0002) and Novex® Sharp Pre-Stained Protein Standard (Product # LC5800). Resolved proteins were then transferred onto a nitrocellulose membrane with iBlot® 2 Dry Blotting System (Product # IB21001). The membrane was probed with the relevant primary and secondary Antibody following blocking with 5 % skimmed milk. Chemiluminescent detection was performed using Pierce™ ECL Western Blotting Substrate (Product # 32106).
Supportive validation
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
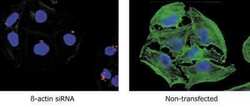
- Experimental details
- An siRNA targeting ß-actin was labeled with Cy™3 using the Silencer™ siRNA Labeling Kit. The labeled siRNA was transfected into HeLa cells and cells were analyzed 96 hours later. Green: ß-actin protein detected with anti-ß-actin (Ambion) and a FITC labeled secondary antibody. Red: Cy3 labeled siRNA. Blue: DAPI stained nuclei. (Cy™3 is a trademark of Amersham Biosciences.)
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- Immunofluorescence analysis of GAPDH was done on 70% confluent log phase HeLa cells. The cells were fixed with 4% paraformaldehyde for 15 minutes, permeabilized with 0.25% Triton™ X-100 for 10 minutes, and blocked with 5% BSA for 1 hour at room temperature. The cells were labeled with GAPDH Mouse Monoclonal Antibody (Product # AM4300) at 1 µg/mL in 1% BSA and incubated for 3 hours at room temperature and then labeled with Goat anti-Mouse IgG (H+L) Superclonal™ Secondary Antibody, Alexa Fluor® 488 conjugate (Product # A28175) at a dilution of 1:2000 for 45 minutes at room temperature (Panel a: green). Nuclei (Panel b: blue) were stained with SlowFade® Gold Antifade Mountant with DAPI (Product # S36938). F-actin (Panel c: red) was stained with Alexa Fluor® 555 Rhodamine Phalloidin (Product # R415, 1:300). Panel d is a merged image showing perinuclear and cytoplasmic localization. Panel e is a no primary antibody control. The images were captured at 60X magnification.
Supportive validation
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- Fig. 1 TGFbeta regulates LXRalpha expression in the mesenchymal cell line Snu449 a Hep3B (left) and Snu449 (right) cells were treated with the indicated compounds (10 muM) and TGFbeta1 (5 ng/ml) for 24 h. The TGFbeta type I receptor kinase inhibitor LY2157299 was added at a final concentration of 2 muM for 24 h as a potent positive control. The expression of the indicated proteins was analyzed by immunoblotting. Experiments performed in biological duplicate. Densitometric quantification is provided below each lane. Results indicated as
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- Figure 4 GFP-LIMK1 fusions increase phosphorylation status of FAK signaling components . (A) Western blot analysis of whole cell lysates from MDA-MB-231 stable transductants probed with antibodies against pFAK, total FAK, pPaxillin, total Paxillin, pSrc, total Src, pAKT, total AKT, pErk1/2 and total Erk 1/2. (B) Western blot analysis for tubulin or GAPDH for each extract used in panel A, as loading controls.
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- Fig 4 Increased apoptosis underlies the nol11 cartilage defects. A) nol11 knockdown results in a progressive increase in apoptosis. At stage 14 no significant difference was observed in rates of TUNEL staining between knockdown and control halves of the embryo. At stages 18 and 28 increased apoptosis was evident on the treated side of whole mount and sectioned paraffin embedded embryos. Note that this increased apoptosis occurs primarily within the craniofacial ectomesenchyme. The graph represents the relative quantification of apoptosis rates at stages 14, 18 and 28. This stage specific increase in apoptosis was confirmed by a similar increase in p53 protein levels in 1 cell injected embryos as assayed by western blot (lower right panel). Dotted red lines mark the embryonic midline. B) No significant change in proliferation rates was noted following nol11 knockdown. C) Inhibition of apoptosis by p53 MO results in a partial rescue of cartilage size and morphology. Each pair of columns in the graph compares cartilage size measured in bilateral halves of embryos. The blue pair reveals no significant difference in cartilage measurements in the left vs right side of the UC embryonic head. In the second pair (red), cartilage size is seen to be comparable on either side of the nol11 morphant head. The final pair illustrates that cartilage size is significantly improved on the side of nol11 morphants rescued with p53 MO (green) relative to the side that received nol11 MO only (red).
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
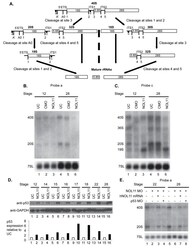
- Experimental details
- Fig 5 Nol11 depletion impairs rDNA transcription and pre-rRNA processing in X . tropicalis . A) Scheme of pre-rRNA processing pathways in X tropicalis . The pre-rRNA is transcribed by RNAPI as a 40S polycistronic precursor. Several cleavages are required to separate the mature rRNAs. The locations of oligonucleotide probes used for northern blots are indicated by lettered lines (a, c) and the cleavage sites indicated. This scheme was adapted from [ 71 - 75 ]. B) Morpholino (MO) depletion of Nol11 impairs pre-rRNA transcription at stage 28. The northern blot was hybridized with probe a (Fig. 5A) and with a probe to the 7SL RNA as a loading control (lower panel). Bands were quantified and analysed by RAMP ([ 60 ]; S6A,B Fig ) C) Morpholino (MO) depletion of Nol11 impairs pre-rRNA transcription and processing. The northern blot was hybridized with probe c (Fig. 5A) and with a probe to the 7SL RNA as a loading control (lower panel). Bands were quantified and analysed by RAMP ([ 60 ]; S6C,D, E, F Fig ). D) Depletion of Nol11 leads to increased p53 levels. The expression of p53 from control and nol11 depleted embryos was analysed by western blot with anti-p53 antibodies. GAPDH levels were used as a loading control. Values for p53 expression normalized to GAPDH are represented in the bar graph. E) MO-resistant human NOL11 (hNOL11) mRNA but not p53 depletion rescues pre-rRNA levels. Embryos injected as shown by + and--in the figure at stages 22 and 28. The pre-rRNAs were visualized w
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
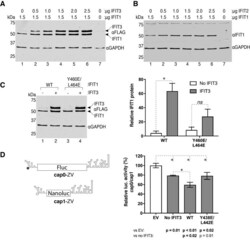
- Experimental details
- Figure 6. IFIT3 stabilizes and promotes IFIT1 activity in cells. ( A and B ) HEK293T cells were transfected with the indicated amounts of plasmid encoding FLAG-tagged versions of IFIT1, IFIT2 and IFIT3. After 24 hours cell lysates were harvested and analyzed by western blotting. (A) anti-FLAG and (B) anti-IFIT1 blots are shown. Empty vector was used to normalize the amount of DNA used in each transfection. The blots shown are representative of three separate experiments. ( C ) HEK293T cells were transfected with 1.5 mug of FLAG-tagged wild type (WT) or mutant IFIT1 and 1.5 mug of FLAG-tagged IFIT3 or empty vector as indicated. After 24 h cell lysates were analyzed as in A. The graph on right shows the quantification of the IFIT1 protein expression relative to GAPDH probed as a loading control. Mean +- the standard deviation of three biological repeats. ( D ) Reporter luciferase was measured as described in the Materials and Methods and expressed as the ratio of Fluc (cap0) over Nanoluc (cap1) signal, normalized to the empty vector control, and shown as the mean +- the standard deviation of three biological repeats. Statistical analysis was performed using an unpaired, two-tailed Students t-test. P -values are indicated, and * denotes statistical significance.
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- Figure S2 Generation of OTULIN-Targeted Mice, OTULIN Expression, Genotyping, and Reconstitution of Bone Marrow Chimeric Mice, Related to Figure 2 (A) Schematic showing the strategy to generate conditional and cell-type-specific knockouts of Otulin . SA, splice acceptor; neo, neomycin-resistance cassette; pA, polyA signal; PGK, murine PGK-1 promoter; DTA, diphtheria toxin A selection cassette; KanR, Kanamycin-resistance cassette. (B) Genotyping of mouse strains. PCR reactions showing the expected products from each genotype. (C) E13.5 embryos stained with X-gal for beta-galactosidase activity and cleared by methyl salicylate shows Otulin promoter activity in multiple tissues. Pictures are representative of five embryos of each genotype from two independent experiments. (D) Immunoblot analysis showing OTULIN expression in multiple tissues from adult wild-type C57BL/6 mice. Blots are representative of three independent experiments. (E) Ratio of CD45.1 + (wild-type B6.SJL) and CD45.2 + (CreERT2- Otulin +/flox or CreERT2- Otulin Lac Z/flox C57BL/6) expressing splenocytes determined by flow cytometry at the termination of chimera experiments. Data were pooled from two independent experiments. (F) Genotyping of bone marrow cells or blood leukocytes from CreERT2- Otulin flox chimeras treated with tamoxifen or vehicle shows complete or near-complete conversion of flox alleles to del alleles upon tamoxifen treatment. Note that WT(+) products are present in all reactions as BJ6.SJL WT c
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- Figure 2 Deletion of Otulin in Immune Cells Causes Acute Systemic Inflammation in Mice (A) OTULIN immunoblot on immune cells from wild-type mice. NK cell, natural killer cell; DC, dendritic cell; MPhi, macrophage. (B) Schematic representation of mixed bone marrow chimera generation. Wild-type (WT) B6.SJL cells are CD45.1 + , and CreERT2- Otulin flox cells are CD45.2 + . (C) Body weight following i.p. administration of tamoxifen (tx; arrows) to CreERT2- Otulin flox chimeric mice. (D) Neutrophil and lymphocyte counts from blood of CreERT2- Otulin flox chimeras and vehicle-treated controls at day 5 following tamoxifen administration. (E and F) Luminex multiplex analysis of serum cytokines and chemokines from terminal bleeds on day 5 presented as (E) a heatmap of relative changes in concentration of all analytes between CreERT2- Otulin +/flox and CreERT2- Otulin Lac Z/flox chimeras and (F) serum concentrations of cytokines and chemokines increased in CreERT2- Otulin Lac Z/flox chimeras. Data were pooled from two independent experiments. (G and H) Flow cytometry analysis of CD11b + Gr-1 + neutrophils in total cellular infiltrate (CD45.1 + and CD45.2 + ) in peritoneal lavage (PL), spleen, and liver from CreERT2- Otulin flox chimeras presented as (G) representative dot plots with percentage of cells in gate indicated and (H) total cell number. (I) Micrographs of hematoxylin and eosin (H&E) stained sections reveal inflammatory foci (arrowheads) in liver parenchyma. Micrographs are re
 Explore
Explore Validate
Validate Learn
Learn Western blot
Western blot