Antibody data
- Antibody Data
- Antigen structure
- References [7]
- Comments [0]
- Validations
- Immunocytochemistry [2]
- Flow cytometry [2]
- Other assay [9]
Submit
Validation data
Reference
Comment
Report error
- Product number
- PA5-18030 - Provider product page

- Provider
- Invitrogen Antibodies
- Product name
- KLF3 Polyclonal Antibody
- Antibody type
- Polyclonal
- Antigen
- Synthetic peptide
- Description
- This antibody is predicted to react with bovine, canine, mouse, porcine and rat based on sequence homology. This antibody is tested in Peptide ELISA: antibody detection limit dilution 32,000.
- Reactivity
- Human
- Host
- Goat
- Isotype
- IgG
- Vial size
- 100 μg
- Concentration
- 0.5 mg/mL
- Storage
- -20°C, Avoid Freeze/Thaw Cycles
Submitted references Ophiopogonin D promotes bone regeneration by stimulating CD31(hi) EMCN(hi) vessel formation.
Krüppel-like factor 3 inhibition by mutated lncRNA Reg1cp results in human high bone mass syndrome.
Direct competition between DNA binding factors highlights the role of Krüppel-like Factor 1 in the erythroid/megakaryocyte switch.
Krüppel-like Factor 3 (KLF3/BKLF) Is Required for Widespread Repression of the Inflammatory Modulator Galectin-3 (Lgals3).
Directing an artificial zinc finger protein to new targets by fusion to a non-DNA-binding domain.
Loss of Krüppel-like factor 3 (KLF3/BKLF) leads to upregulation of the insulin-sensitizing factor adipolin (FAM132A/CTRP12/C1qdc2).
Generation of mice deficient in both KLF3/BKLF and KLF8 reveals a genetic interaction and a role for these factors in embryonic globin gene silencing.
Yang M, Li CJ, Xiao Y, Guo Q, Huang Y, Su T, Luo XH, Jiang TJ
Cell proliferation 2020 Mar;53(3):e12784
Cell proliferation 2020 Mar;53(3):e12784
Krüppel-like factor 3 inhibition by mutated lncRNA Reg1cp results in human high bone mass syndrome.
Yang M, Guo Q, Peng H, Xiao YZ, Xiao Y, Huang Y, Li CJ, Su T, Zhang YL, Lei MX, Chen HL, Jiang TJ, Luo XH
The Journal of experimental medicine 2019 Aug 5;216(8):1944-1964
The Journal of experimental medicine 2019 Aug 5;216(8):1944-1964
Direct competition between DNA binding factors highlights the role of Krüppel-like Factor 1 in the erythroid/megakaryocyte switch.
Norton LJ, Hallal S, Stout ES, Funnell APW, Pearson RCM, Crossley M, Quinlan KGR
Scientific reports 2017 Jun 9;7(1):3137
Scientific reports 2017 Jun 9;7(1):3137
Krüppel-like Factor 3 (KLF3/BKLF) Is Required for Widespread Repression of the Inflammatory Modulator Galectin-3 (Lgals3).
Knights AJ, Yik JJ, Mat Jusoh H, Norton LJ, Funnell AP, Pearson RC, Bell-Anderson KS, Crossley M, Quinlan KG
The Journal of biological chemistry 2016 Jul 29;291(31):16048-58
The Journal of biological chemistry 2016 Jul 29;291(31):16048-58
Directing an artificial zinc finger protein to new targets by fusion to a non-DNA-binding domain.
Lim WF, Burdach J, Funnell AP, Pearson RC, Quinlan KG, Crossley M
Nucleic acids research 2016 Apr 20;44(7):3118-30
Nucleic acids research 2016 Apr 20;44(7):3118-30
Loss of Krüppel-like factor 3 (KLF3/BKLF) leads to upregulation of the insulin-sensitizing factor adipolin (FAM132A/CTRP12/C1qdc2).
Bell-Anderson KS, Funnell AP, Williams H, Mat Jusoh H, Scully T, Lim WF, Burdach JG, Mak KS, Knights AJ, Hoy AJ, Nicholas HR, Sainsbury A, Turner N, Pearson RC, Crossley M
Diabetes 2013 Aug;62(8):2728-37
Diabetes 2013 Aug;62(8):2728-37
Generation of mice deficient in both KLF3/BKLF and KLF8 reveals a genetic interaction and a role for these factors in embryonic globin gene silencing.
Funnell AP, Mak KS, Twine NA, Pelka GJ, Norton LJ, Radziewic T, Power M, Wilkins MR, Bell-Anderson KS, Fraser ST, Perkins AC, Tam PP, Pearson RC, Crossley M
Molecular and cellular biology 2013 Aug;33(15):2976-87
Molecular and cellular biology 2013 Aug;33(15):2976-87
No comments: Submit comment
Supportive validation
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
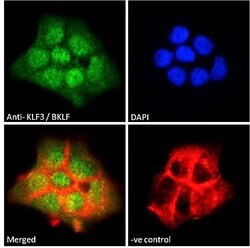
- Experimental details
- Immunocytochemistry analysis of KLF3 using KLF3 Polyclonal Antibody (Product # PA5-18030) in paraformaldehyde fixed A431 cells, permeabilized with 0.15% Triton. Primary incubation 1hr (10 µg/mL) followed by Alexa Fluor 488 secondary antibody (2 µg/mL), showing nuclear staining. Actin filaments were stained with phalloidin (red) and the nuclear stain is DAPI (blue). Negative control: Unimmunized goat IgG (10 µg/mL) followed by Alexa Fluor 488 secondary antibody (2 µg/mL).
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
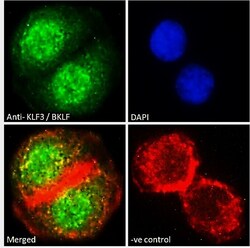
- Experimental details
- Immunocytochemistry analysis of KLF3 using KLF3 Polyclonal Antibody (Product # PA5-18030) in paraformaldehyde fixed A549 cells, permeabilized with 0.15% Triton. Primary incubation 1hr (10 µg/mL) followed by Alexa Fluor 488 secondary antibody (2 µg/mL), showing nuclear staining. Actin filaments were stained with phalloidin (red) and the nuclear stain is DAPI (blue). Negative control: Unimmunized goat IgG (10 µg/mL) followed by Alexa Fluor 488 secondary antibody (2 µg/mL).
Supportive validation
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
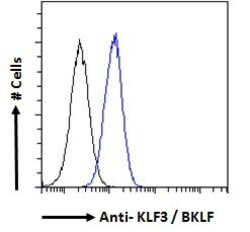
- Experimental details
- Flow cytometric analysis of KLF3 in A431 cells using a polyclonal antibody (Product #PA5-18030). A431 cells (blue line) were paraformaldehyde fixed and permeabilized with 0.5% Triton. The primary antibody was incubated for one hour (10 µg/mL) followed by an Alexa Fluor 488 secondary antibody (1 µg/mL). IgG control: Unimmunized goat IgG (black line) followed by an Alexa Fluor 488 secondary antibody.
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
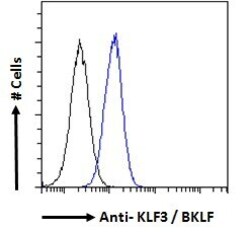
- Experimental details
- Flow cytometric analysis of KLF3 in A431 cells using a polyclonal antibody (Product #PA5-18030). A431 cells (blue line) were paraformaldehyde fixed and permeabilized with 0.5% Triton. The primary antibody was incubated for one hour (10 µg/mL) followed by an Alexa Fluor 488 secondary antibody (1 µg/mL). IgG control: Unimmunized goat IgG (black line) followed by an Alexa Fluor 488 secondary antibody.
Supportive validation
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
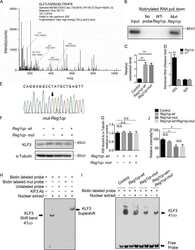
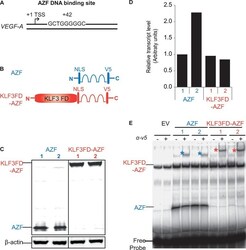
- Figure 1. Construction and verification of AZF and KLF3FD-AZF protein. ( A ) Human VEGF-A locus showing the location of the 9 nt target sequence GCTGGGGGC 42 bases downstream of the +1 TSS that the AZF was designed to recognize and bind ( 15 ). ( B ) Schematic representation of AZF and KLF3FD-AZF with an extra KLF3 functional domain (KLF3 FD) fused to the N-terminus of the artificial zinc fingers (AZFs). Western blot ( C ) and quantitative real time PCR ( D ) showing equivalent protein and mRNA transcript expression, respectively, for the four selected HEK293 clones stably expressing AZF or KLF3FD-AZF. For western blot, beta-actin was included as the loading control and for real-time PCR, transcript expression was normalized to 18S rRNA level and is shown relative to the expression for AZF clone 1 (first bar), which was set to an arbitrary value of 1. ( E ) Electrophoretic mobility shift assay showing equivalent in vitro AZF and KLF3FD-AZF binding to a 32 P radiolabeled EMSA probe containing the 9 nt target sequence GCTGGGGGC. Asterisks indicate the supershift of the protein-DNA probe complex by an anti-V5 antibody confirming the identity of the V5-tagged AZF proteins.
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
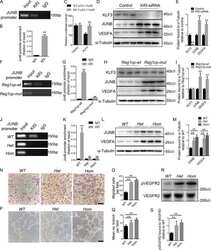
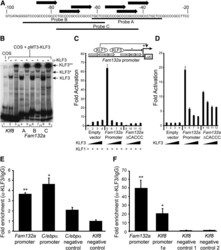
- FIG. 5. KLF3 binds the Fam132a promoter in vitro and in vivo and is able to repress its activity. A : Probe design to assess binding of KLF3 to the Fam132a proximal promoter by EMSA: three probes (probes A, B, and C) were designed to provide coverage of major CACCC boxes identified in the promoter. Black arrows indicate CACCC boxes; numbering is with respect to the transcriptional start site. B : EMSA; KLF3 was expressed in COS-7 cells and binding of nuclear extracts to probe A ( lanes 4 and 5 ), probe B ( lanes 6 and 7 ), and probe C ( lanes 8 and 9 ) assessed. Also shown is binding of KLF3 to a previously validated consensus sequence in the Klf8 promoter ( 29 ) ( lanes 2 and 3 ). Binding of untransfected COS nuclear extract to the Klf8 probe is shown in lane 1 . alpha-KLF3 indicates an anti-KLF3 antibody used in supershift to validate KLF3 specific binding. Arrows show the positions of probe complexes bound by KLF3. *KLF3, higher-order multimeric complexes. **KLF3, supershift complexes. C : SL-2 cells were transiently transfected with either pGL4.10[ luc2 ], pGL4.10[ luc2 ] containing a region of the wild-type Fam132a proximal promoter, or pGL4.10[ luc2 ] containing a mutant version of the promoter lacking major CACCC boxes ( Fam132a DeltaCACCC). pPac-KLF1 (250 ng) was used to drive activation of the reporter constructs. Increasing amounts of pPac-KLF3 were included to assess repression: 0 ng ( lanes 1, 5 , and 9 ), 25 ng ( lanes 2, 6 , and 10 ), 50 ng ( lanes 3, 7 , and 11
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
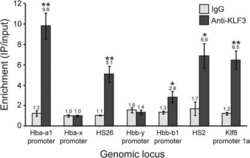
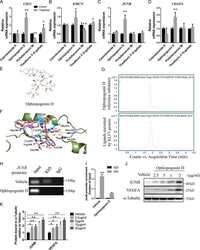
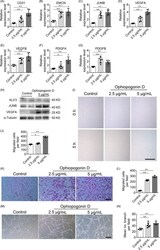
- Figure 3 Ophiopogonin D acts as a KLF3 inhibitor and promotes vessel formation in vitro. A-G, qRT-PCR analysis of the relative levels of CD31 (A), EMCN (B), JUNB (C), VEGFA (D), VEGFB (E), PDGFA (F) and PDGFB (G). H, Western blotting analysis of the KLF3, JUNB and VEGFA expression in HMECs treated with vehicle or different doses of ophiopogonin D. I and J, Representative images (I) and quantification (J) of migration HMECs in wound healing assay. Scale bar, 500 mum. K-L, Representative images (K) and quantification (L) of a transwell migration assay. Scale bar, 200 mum. M and N, Representative images (M) and quantification (N) of tube branch numbers of a matrigel tube formation assay. Scale bar, 200 mum. Data are shown as mean +- SD, (The cell experiments were repeated for three times, n = 6 in A-G, J, L and N). * P < .05; ** P < .01 by one-way ANOVA
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
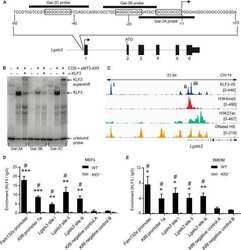
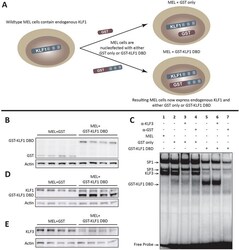
- Figure 1 The GST-KLF1 DBD construct is capable of binding to the canonical KLF1 binding site and has a biological effect in MELs. ( A ) Schematic demonstrating the experimental design of the two different constructs nucleofected into MEL cells. ( B ) Western blots from MEL lines showing protein expression of GST and GST-KLF1 DBD for each clone. Actin is presented as a loading control. ( C ) EMSAs from representative MEL clones indicating GST-KLF1 DBD is capable of binding the canonical KLF1 consensus sequence ( beta - globin CACCC probe) in vitro . ( D ) Western blots from MEL clones with an antibody that recognises both endogenous KLF1 and GST-KLF1 DBD. beta-Actin is presented as a loading control. ( E ) Western blots from MEL clones with an antibody that recognises KLF3, indicating reduced expression of endogenous KLF3 in cells overexpressing GST-KLF1 DBD compared to GST only clones. beta-Actin is presented as a loading control. n = 4 for each construct.
 Explore
Explore Validate
Validate Learn
Learn Western blot
Western blot Immunocytochemistry
Immunocytochemistry