Antibody data
- Antibody Data
- Antigen structure
- References [2]
- Comments [0]
- Validations
- Other assay [10]
Submit
Validation data
Reference
Comment
Report error
- Product number
- MA1-16570 - Provider product page

- Provider
- Invitrogen Antibodies
- Product name
- FANCD2 Monoclonal Antibody (FI-17)
- Antibody type
- Monoclonal
- Antigen
- Other
- Description
- By Western blot, this antibody recognizes a band at ~155 and ~162 kDa (post-translationally modified form). Suggested positive control: Hela whole cell extract.
- Reactivity
- Human, Mouse, Hamster
- Host
- Mouse
- Isotype
- IgG
- Antibody clone number
- FI-17
- Vial size
- 100 μL
- Concentration
- Conc. Not Determined
- Storage
- -20°C, Avoid Freeze/Thaw Cycles
Submitted references FANCD2 Binds Human Papillomavirus Genomes and Associates with a Distinct Set of DNA Repair Proteins to Regulate Viral Replication.
AluY-mediated germline deletion, duplication and somatic stem cell reversion in UBE2T defines a new subtype of Fanconi anemia.
Spriggs CC, Laimins LA
mBio 2017 Feb 14;8(1)
mBio 2017 Feb 14;8(1)
AluY-mediated germline deletion, duplication and somatic stem cell reversion in UBE2T defines a new subtype of Fanconi anemia.
Virts EL, Jankowska A, Mackay C, Glaas MF, Wiek C, Kelich SL, Lottmann N, Kennedy FM, Marchal C, Lehnert E, Scharf RE, Dufour C, Lanciotti M, Farruggia P, Santoro A, Savasan S, Scheckenbach K, Schipper J, Wagenmann M, Lewis T, Leffak M, Farlow JL, Foroud TM, Honisch E, Niederacher D, Chakraborty SC, Vance GH, Pruss D, Timms KM, Lanchbury JS, Alpi AF, Hanenberg H
Human molecular genetics 2015 Sep 15;24(18):5093-108
Human molecular genetics 2015 Sep 15;24(18):5093-108
No comments: Submit comment
Supportive validation
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
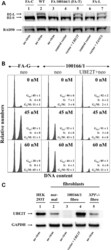
- Figure 1. Identification of UBE2T as new FA gene by retroviral complementation of FA 100166/1 fibroblasts and western blotting. ( A ) Western blot analysis of protein extracts from transduced fibroblast lines immunostained with a FANCD2 antibody to detect non-ubiquitinated (D2-S) and monoubiquitinated (D2-L) FANCD2. Lanes 1-3 depict the FANCD2 ubiquitination status of FANCC -deficient (FA-C), wild-type (WT) and FA100166/1 fibroblasts exposed to 150 n m MMC overnight. Lanes 4-7 show the FANCD2 monoubiquitination status of G418-resistant FA 100166/1 and FANCL-deficient fibroblasts, genetically modified with a control vector or with vectors expressing UBE2T or FANCL cDNAs, respectively. Staining with a RAD50 antibody was used to visually confirm equal loading. ( B ) Flow cytometric cell cycle analysis of transduced primary fibroblasts grown for 3 days in 0, 45 or 60 n m MMC. FA-G: primary FANCG-/- fibroblasts ( 38 ) transduced with the control vector (neo control). FA 100166/1: primary patient fibroblasts transduced with the control vector (neo control) or the UBE2T-coexpressing vector (UBE2T + neo). The distribution of cells in G 0 /G 1 , S and G 2 /M arrest from three different experiments is shown. ( C ) Western blot analysis of protein extracts from the indicated cells upon staining with a polyclonal antibody specific for human UBE2T protein. HEK293T cells, immortalized fibroblasts from a healthy donor (normal) and from a FANCQ/ERCC4/XPF deficient ( XPF-/- ) patient were use
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
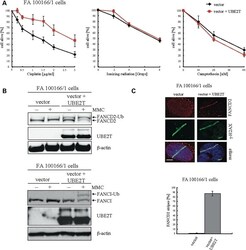
- Experimental details
- Figure 4. The cellular phenotype of human UBE2T-/- fibroblasts is comparable to that of FA cells with defects in FA core complex members. ( A ) Survival of UBE2T-/- fibroblasts after exposure to genotoxins. An isogenic pair of UBE2T-/- fibroblasts transduced with a retroviral vector encoding UBE2T or the corresponding control vector were exposed to increasing doses of the genotoxins cisplatin, ionizing radiation and camptothecin. ( B ) Western blot analysis to visualize intact FANCD2 and FANCI monoubiquitination in immortalized UBE2T-/- fibroblasts after exposure to MMC by UBE2T expression. ( C ) Restoration of FANCD2 recruitment to sites of DNA crosslinks by UBE2T expression. Psoralen interstrand crosslinks were induced along the track of a UVA laser in corrected and non-corrected UBE2T-/- fibroblasts and then visualized by gamma-H2AX staining. FANCD2 accumulation at the sites of psoralen laser tracks only occurred in FA100166/1 cells that had been transduced with the UBE2T expressing retroviral vector.
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
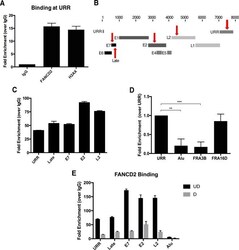
- Experimental details
- FIG 6 FANCD2 is preferentially recruited to HPV DNA. (A) Chromatin immunoprecipitation (ChIP) analysis of FANCD2 and gammaH2AX binding to the URR in CIN612 cells. Quantitative real-time PCR (qRT-PCR) was performed using a LightCycler 480 (Roche), and fold enrichment was quantitated relative to an IgG control. Similar results were seen in three independent experiments. Error bars represent the standard deviations between experiments. (B) Schematic of the HPV31 linearized genome, with primer regions indicated with arrows. (C) ChIP analysis for FANCD2 binding at indicated sites in the viral genome. Fold enrichment was normalized to an IgG control. Similar results were seen in three independent experiments. Error bars represent the standard deviations between experiments. (D) ChIP analysis of FANCD2 binding at the URR compared to Alu repeat and fragile site regions (FRA3B and FRA16D) in the host genome. Enrichment was normalized to an IgG control and is represented as fold change over URR across three independent experiments. The graph represented as percentage of input shows a similar trend ( Fig. S1 ). Error bars represent the standard deviations between experiments. A standard Student's t test was used to determine statistical significance. **, P < 0.005; ***, P < 0.0005. (E) CIN612 cells were differentiated for 72 h in 1.5 mM calcium medium, and ChIP analysis was performed for binding across the HPV genome. Fold enrichment was normalized to an IgG control. Similar result
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
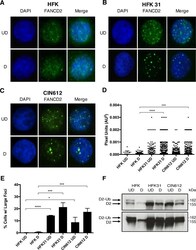
- Experimental details
- FIG 2 HPV infection leads to FA pathway activation. (A to C) Immunofluorescence analysis of FANCD2 localization in HFK, HFK31, and CIN612 cells that were differentiated for 72 h in 1.5 mM calcium medium. Cells were stained with anti-FANCD2 (green) and counterstained with DAPI (blue). UD, undifferentiated; D, differentiated. (D) ImageJ software was used to quantitate focus size by automated particle analysis. The graph represents the size of individual foci represented in pixel units (AU 2 ). Error bars represent the standard errors of the means within the sample. A standard Student's t test was used to determine statistical significance. ***, P
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
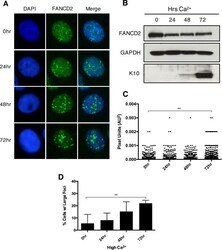
- Experimental details
- FIG 3 FA pathway activation further increases as differentiation progresses in HPV-positive cells. (A) Immunofluorescence analysis of FANCD2 localization in CIN612 cells that were differentiated in 1.5 mM calcium for 24, 48, or 72 h. Cells were stained with anti-FANCD2 (green) and counterstained with DAPI (blue). (B) Western blot analysis of FANCD2 levels in CIN612 cells that were differentiated in high-calcium medium for 24, 48, or 72 h. GAPDH was used as a loading control. Epithelial differentiation was confirmed by levels of cytokeratin 10. (C) ImageJ software was used to quantitate focus size by an automated particle analysis program. The graph represents individual focus size represented in pixel units (AU 2 ). Error bars represent the standard error mean within the sample. A standard Student's t test was used to determine statistical significance. **, P
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
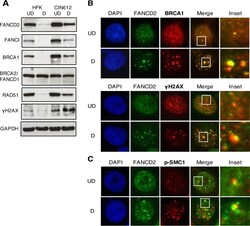
- FIG 4 FANCD2 colocalizes with components of the ATM pathway in discrete nuclear foci. (A) HFKs and CIN612 cells were differentiated for 72 h in 1.5 mM calcium medium. Western blot analysis was performed using antibodies to FANCD2, FANCI, BRCA1, BRCA2, RAD51, and gammaH2AX. GAPDH was used as a loading control. (B and C) CIN612 cells were differentiated for 72 h in 1.5 mM calcium medium and stained with anti-FANCD2 (green) and either anti-BRCA1, anti-gammaH2AX, or anti-p-SMC1 (red). Cells were counterstained with DAPI (blue). UD, undifferentiated; D, differentiated.
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
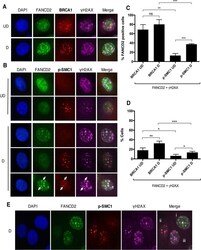
- FIG 5 Distinct populations of foci exist during HPV infection. (A and B) CIN612 cells were differentiated for 72 h in 1.5 mM calcium medium. Immunofluorescence analysis was performed on cells stained with anti-FANCD2 (green) and either anti-BRCA1 or anti-p-SMC1 (red). Cells were then counterstained with anti-gammaH2AX (pink) and DAPI (blue). Arrows indicate foci where FANCD2, gammaH2AX, and p-SMC1 are found together. UD, undifferentiated; D, differentiated. (C) The graph demonstrates the percentage of cells with FANCD2 foci where at least one focus colocalizes with gammaH2AX and either BRCA1 or p-SMC1. (D) The graph represents the percentage of all HPV-positive cells where at least one FANCD2 focus colocalizes with gammaH2AX and either BRCA1 or p-SMC1. Error bars represent the standard deviations between experiments. A standard Student's t test was used to determine statistical significance. *, P
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
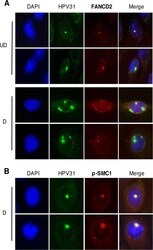
- FIG 7 FANCD2 localizes to HPV replication centers. (A) CIN612 cells were differentiated for 72 h in 1.5 mM calcium medium. Immunofluorescence analysis for FANCD2 (red) was performed followed by fluorescent in situ hybridization (I-FISH) for HPV31 DNA (green). Cells were counterstained with DAPI (blue). In undifferentiated cells, the FANCD2 signal overlapped with 42.47% +- 12.17% of the HPV31 DNA signal and 11.55%+- 1.479% in differentiated cells. UD, undifferentiated; D, differentiated. (B) CIN612 cells were differentiated in 1.5 mM calcium medium, and immunofluorescence analysis for p-SMC1 (red) was performed followed by fluorescent in situ hybridization for HPV31 DNA (green). Cells were counterstained with DAPI (blue). In differentiated cells, the p-SMC1 overlapped with 31.85% +- 8.54% of the HPV31 DNA signal. The percentage of overlap between the HPV31 DNA signal and either FANCD2 or p-SMC1 was measured using ImageJ area analysis and found to be statistically significant where P is
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
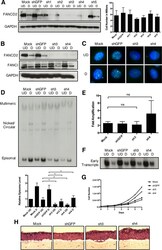
- FIG 8 Knockdown of FANCD2 limits HPV31 replication. (A) CIN612 cells were transiently transduced with lentiviral vectors encoding five individual shRNAs against FANCD2 or GFP as a control. After 48 h, cells were differentiated in 1.5% methylcellulose for an additional 48 h. FANCD2 knockdown was assessed by Western blot analysis using GAPDH as a loading control. At 96 h posttransduction, cells were harvested and stained with trypan blue (Bio-Rad) to assess cell viability. The graph shows the total number of live cells in monolayer culture at the time of collection. Error bars represent the standard deviations between measurements. UD, undifferentiated; D, differentiated. (B) CIN612 cells transduced with shGFP or shFANCD2 were differentiated for 48 h in 1.5% methylcellulose, and Western blot analysis was used to assess FANCD2 and FANCI protein levels. GAPDH was used as a loading control. (C) Immunofluorescence analysis of control and FANCD2 knockdown cells that were differentiated for 72 h in 1.5 mM calcium medium. Cells were stained with anti-FANCD2 (green) and counterstained with DAPI (blue). (D) CIN612 cells were differentiated for 48 h in 1.5% methylcellulose, and total DNA was isolated from control and shFANCD2 cells. Viral replication was assessed by Southern blot analysis. Similar results were seen using high calcium concentrations to induce differentiation ( Fig. S2 ). Quantification of episomal band intensity was determined by densitometry using Image Lab software and
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
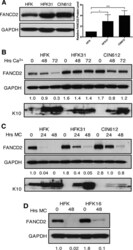
- FIG 1 Levels of FANCD2 are increased in HPV-positive cells and remain elevated through differentiation. (A) Western blot analysis of FANCD2 levels in normal human foreskin keratinocytes (HFKs), HFKs stably transfected with HPV31 whole genomic DNA (HFK31), and CIN612 cells. The graph demonstrates FANCD2 protein levels relative to GAPDH and normalized to FANCD2 levels in HFKs across three independent experiments. Error bars represent standard deviations between experiments. A standard Student's t test was used to determine statistical significance. *, P
 Explore
Explore Validate
Validate Learn
Learn Western blot
Western blot Immunoprecipitation
Immunoprecipitation Other assay
Other assay