Antibody data
- Antibody Data
- Antigen structure
- References [26]
- Comments [0]
- Validations
- Flow cytometry [1]
- Other assay [10]
Submit
Validation data
Reference
Comment
Report error
- Product number
- MHCD0418 - Provider product page

- Provider
- Invitrogen Antibodies
- Product name
- CD4 Monoclonal Antibody (S3.5), PE-Cyanine5.5
- Antibody type
- Monoclonal
- Antigen
- Other
- Description
- The R-phycoerythrin (PE)-Cy5.5® tandem conjugate permits simultaneous multicolor labeling and detection of multiple targets with excitation by a single excitation source-the 488 nm line of the argon-ion laser.
- Reactivity
- Human
- Host
- Mouse
- Isotype
- IgG
- Antibody clone number
- S3.5
- Vial size
- 500 µL
- Storage
- 4° C, store in dark
Submitted references A SARS-CoV-2 ferritin nanoparticle vaccine elicits protective immune responses in nonhuman primates.
Efficacy and breadth of adjuvanted SARS-CoV-2 receptor-binding domain nanoparticle vaccine in macaques.
Cross-HLA targeting of intracellular oncoproteins with peptide-centric CARs.
Vaccine-induced ICOS(+)CD38(+) circulating Tfh are sensitive biosensors of age-related changes in inflammatory pathways.
Recombinant MVA-prime elicits neutralizing antibody responses by inducing antigen-specific B cells in the germinal center.
Efficacy and breadth of adjuvanted SARS-CoV-2 receptor-binding domain nanoparticle vaccine in macaques.
Evaluation of heterologous prime-boost vaccination strategies using chimpanzee adenovirus and modified vaccinia virus for TB subunit vaccination in rhesus macaques.
Interferon-α alters host glycosylation machinery during treated HIV infection.
Robust T Cell Immunity in Convalescent Individuals with Asymptomatic or Mild COVID-19.
Route of Vaccine Administration Alters Antigen Trafficking but Not Innate or Adaptive Immunity.
CD57(+) Memory T Cells Proliferate In Vivo.
Longitudinal variation in human immunodeficiency virus long terminal repeat methylation in individuals on suppressive antiretroviral therapy.
Principles Governing Establishment versus Collapse of HIV-1 Cellular Spread.
Autoreactive T effector memory differentiation mirrors β cell function in type 1 diabetes.
Fecal Bacterial Communities in treated HIV infected individuals on two antiretroviral regimens.
Perturbed CD8(+) T cell TIGIT/CD226/PVR axis despite early initiation of antiretroviral treatment in HIV infected individuals.
Gut Homing CD4+ and CD8+ T-Cell Frequencies in HIV Infected Individuals on Antiretroviral Treatment.
Adjuvant-dependent innate and adaptive immune signatures of risk of SIVmac251 acquisition.
Human Immunodeficiency Virus Type 1 Monoclonal Antibodies Suppress Acute Simian-Human Immunodeficiency Virus Viremia and Limit Seeding of Cell-Associated Viral Reservoirs.
Collapse of Cytolytic Potential in SIV-Specific CD8+ T Cells Following Acute SIV Infection in Rhesus Macaques.
Selective reduction of IFN-γ single positive mycobacteria-specific CD4+ T cells in HIV-1 infected individuals with latent tuberculosis infection.
Temporal Dynamics of CD8+ T Cell Effector Responses during Primary HIV Infection.
A subset of circulating blood mycobacteria-specific CD4 T cells can predict the time to Mycobacterium tuberculosis sputum culture conversion.
Effects of immunosuppression on circulating adeno-associated virus capsid-specific T cells in humans.
Capsid-specific T-cell responses to natural infections with adeno-associated viruses in humans differ from those of nonhuman primates.
Anti-CD8 antibodies can trigger CD8+ T cell effector function in the absence of TCR engagement and improve peptide-MHCI tetramer staining.
Joyce MG, King HAD, Elakhal-Naouar I, Ahmed A, Peachman KK, Macedo Cincotta C, Subra C, Chen RE, Thomas PV, Chen WH, Sankhala RS, Hajduczki A, Martinez EJ, Peterson CE, Chang WC, Choe M, Smith C, Lee PJ, Headley JA, Taddese MG, Elyard HA, Cook A, Anderson A, McGuckin Wuertz K, Dong M, Swafford I, Case JB, Currier JR, Lal KG, Molnar S, Nair MS, Dussupt V, Daye SP, Zeng X, Barkei EK, Staples HM, Alfson K, Carrion R, Krebs SJ, Paquin-Proulx D, Karasavva N, Polonis VR, Jagodzinski LL, Amare MF, Vasan S, Scott PT, Huang Y, Ho DD, de Val N, Diamond MS, Lewis MG, Rao M, Matyas GR, Gromowski GD, Peel SA, Michael NL, Bolton DL, Modjarrad K
Science translational medicine 2022 Feb 16;14(632):eabi5735
Science translational medicine 2022 Feb 16;14(632):eabi5735
Efficacy and breadth of adjuvanted SARS-CoV-2 receptor-binding domain nanoparticle vaccine in macaques.
King HAD, Joyce MG, Lakhal-Naouar I, Ahmed A, Cincotta CM, Subra C, Peachman KK, Hack HR, Chen RE, Thomas PV, Chen WH, Sankhala RS, Hajduczki A, Martinez EJ, Peterson CE, Chang WC, Choe M, Smith C, Headley JA, Elyard HA, Cook A, Anderson A, Wuertz KM, Dong M, Swafford I, Case JB, Currier JR, Lal KG, Amare MF, Dussupt V, Molnar S, Daye SP, Zeng X, Barkei EK, Alfson K, Staples HM, Carrion R, Krebs SJ, Paquin-Proulx D, Karasavvas N, Polonis VR, Jagodzinski LL, Vasan S, Scott PT, Huang Y, Nair MS, Ho DD, de Val N, Diamond MS, Lewis MG, Rao M, Matyas GR, Gromowski GD, Peel SA, Michael NL, Modjarrad K, Bolton DL
Proceedings of the National Academy of Sciences of the United States of America 2021 Sep 21;118(38)
Proceedings of the National Academy of Sciences of the United States of America 2021 Sep 21;118(38)
Cross-HLA targeting of intracellular oncoproteins with peptide-centric CARs.
Yarmarkovich M, Marshall QF, Warrington JM, Premaratne R, Farrel A, Groff D, Li W, di Marco M, Runbeck E, Truong H, Toor JS, Tripathi S, Nguyen S, Shen H, Noel T, Church NL, Weiner A, Kendsersky N, Martinez D, Weisberg R, Christie M, Eisenlohr L, Bosse KR, Dimitrov DS, Stevanovic S, Sgourakis NG, Kiefel BR, Maris JM
Nature 2021 Nov;599(7885):477-484
Nature 2021 Nov;599(7885):477-484
Vaccine-induced ICOS(+)CD38(+) circulating Tfh are sensitive biosensors of age-related changes in inflammatory pathways.
Herati RS, Silva LV, Vella LA, Muselman A, Alanio C, Bengsch B, Kurupati RK, Kannan S, Manne S, Kossenkov AV, Canaday DH, Doyle SA, Ertl HCJ, Schmader KE, Wherry EJ
Cell reports. Medicine 2021 May 18;2(5):100262
Cell reports. Medicine 2021 May 18;2(5):100262
Recombinant MVA-prime elicits neutralizing antibody responses by inducing antigen-specific B cells in the germinal center.
Eslamizar L, Petrovas C, Leggat DJ, Furr K, Lifton ML, Levine G, Ma S, Fletez-Brant C, Hoyland W, Prabhakaran M, Narpala S, Boswell K, Yamamoto T, Liao HX, Pickup D, Ramsburg E, Sutherland L, McDermott A, Roederer M, Montefiori D, Koup RA, Haynes BF, Letvin NL, Santra S
NPJ vaccines 2021 Jan 25;6(1):15
NPJ vaccines 2021 Jan 25;6(1):15
Efficacy and breadth of adjuvanted SARS-CoV-2 receptor-binding domain nanoparticle vaccine in macaques.
King HAD, Gordon Joyce M, Naouar IE, Ahmed A, Cincotta CM, Subra C, Peachman KK, Hack HH, Chen RE, Thomas PV, Chen WH, Sankhala RS, Hajduczki A, Martinez EJ, Peterson CE, Chang WC, Choe M, Smith C, Headley JA, Elyard HA, Cook A, Anderson A, Wuertz KM, Dong M, Swafford I, Case JB, Currier JR, Lal KG, Amare MF, Dussupt V, Molnar S, Daye SP, Zeng X, Barkei EK, Alfson K, Staples HM, Carrion R, Krebs SJ, Paquin-Proulx D, Karasavvas N, Polonis VR, Jagodzinski LL, Vasan S, Scott PT, Huang Y, Nair MS, Ho DD, de Val N, Diamond MS, Lewis MG, Rao M, Matyas GR, Gromowski GD, Peel SA, Michael NL, Modjarrad K, Bolton DL
bioRxiv : the preprint server for biology 2021 Apr 10;
bioRxiv : the preprint server for biology 2021 Apr 10;
Evaluation of heterologous prime-boost vaccination strategies using chimpanzee adenovirus and modified vaccinia virus for TB subunit vaccination in rhesus macaques.
Vierboom MPM, Chenine AL, Darrah PA, Vervenne RAW, Boot C, Hofman SO, Sombroek CC, Dijkman K, Khayum MA, Stammes MA, Haanstra KG, Hoffmann C, Schmitt D, Silvestre N, White AG, Borish HJ, Seder RA, Ouaked N, Leung-Theung-Long S, Inchauspé G, Anantha R, Limbach M, Evans TG, Casimiro D, Lempicki M, Laddy DJ, Bonavia A, Verreck FAW
NPJ vaccines 2020;5(1):39
NPJ vaccines 2020;5(1):39
Interferon-α alters host glycosylation machinery during treated HIV infection.
Giron LB, Colomb F, Papasavvas E, Azzoni L, Yin X, Fair M, Anzurez A, Damra M, Mounzer K, Kostman JR, Tebas P, O'Doherty U, Tateno H, Liu Q, Betts MR, Montaner LJ, Abdel-Mohsen M
EBioMedicine 2020 Sep;59:102945
EBioMedicine 2020 Sep;59:102945
Robust T Cell Immunity in Convalescent Individuals with Asymptomatic or Mild COVID-19.
Sekine T, Perez-Potti A, Rivera-Ballesteros O, Strålin K, Gorin JB, Olsson A, Llewellyn-Lacey S, Kamal H, Bogdanovic G, Muschiol S, Wullimann DJ, Kammann T, Emgård J, Parrot T, Folkesson E, Karolinska COVID-19 Study Group, Rooyackers O, Eriksson LI, Henter JI, Sönnerborg A, Allander T, Albert J, Nielsen M, Klingström J, Gredmark-Russ S, Björkström NK, Sandberg JK, Price DA, Ljunggren HG, Aleman S, Buggert M
Cell 2020 Oct 1;183(1):158-168.e14
Cell 2020 Oct 1;183(1):158-168.e14
Route of Vaccine Administration Alters Antigen Trafficking but Not Innate or Adaptive Immunity.
Ols S, Yang L, Thompson EA, Pushparaj P, Tran K, Liang F, Lin A, Eriksson B, Karlsson Hedestam GB, Wyatt RT, Loré K
Cell reports 2020 Mar 24;30(12):3964-3971.e7
Cell reports 2020 Mar 24;30(12):3964-3971.e7
CD57(+) Memory T Cells Proliferate In Vivo.
Ahmed R, Miners KL, Lahoz-Beneytez J, Jones RE, Roger L, Baboonian C, Zhang Y, Wang ECY, Hellerstein MK, McCune JM, Baird DM, Price DA, Macallan DC, Asquith B, Ladell K
Cell reports 2020 Dec 15;33(11):108501
Cell reports 2020 Dec 15;33(11):108501
Longitudinal variation in human immunodeficiency virus long terminal repeat methylation in individuals on suppressive antiretroviral therapy.
Cortés-Rubio CN, Salgado-Montes de Oca G, Prado-Galbarro FJ, Matías-Florentino M, Murakami-Ogasawara A, Kuri-Cervantes L, Carranco-Arenas AP, Ormsby CE, Cortés-Rubio IK, Reyes-Terán G, Ávila-Ríos S
Clinical epigenetics 2019 Sep 13;11(1):134
Clinical epigenetics 2019 Sep 13;11(1):134
Principles Governing Establishment versus Collapse of HIV-1 Cellular Spread.
Hataye JM, Casazza JP, Best K, Liang CJ, Immonen TT, Ambrozak DR, Darko S, Henry AR, Laboune F, Maldarelli F, Douek DC, Hengartner NW, Yamamoto T, Keele BF, Perelson AS, Koup RA
Cell host & microbe 2019 Dec 11;26(6):748-763.e20
Cell host & microbe 2019 Dec 11;26(6):748-763.e20
Autoreactive T effector memory differentiation mirrors β cell function in type 1 diabetes.
Yeo L, Woodwyk A, Sood S, Lorenc A, Eichmann M, Pujol-Autonell I, Melchiotti R, Skowera A, Fidanis E, Dolton GM, Tungatt K, Sewell AK, Heck S, Saxena A, Beam CA, Peakman M
The Journal of clinical investigation 2018 Aug 1;128(8):3460-3474
The Journal of clinical investigation 2018 Aug 1;128(8):3460-3474
Fecal Bacterial Communities in treated HIV infected individuals on two antiretroviral regimens.
Pinto-Cardoso S, Lozupone C, Briceño O, Alva-Hernández S, Téllez N, Adriana A, Murakami-Ogasawara A, Reyes-Terán G
Scientific reports 2017 Mar 6;7:43741
Scientific reports 2017 Mar 6;7:43741
Perturbed CD8(+) T cell TIGIT/CD226/PVR axis despite early initiation of antiretroviral treatment in HIV infected individuals.
Tauriainen J, Scharf L, Frederiksen J, Naji A, Ljunggren HG, Sönnerborg A, Lund O, Reyes-Terán G, Hecht FM, Deeks SG, Betts MR, Buggert M, Karlsson AC
Scientific reports 2017 Jan 13;7:40354
Scientific reports 2017 Jan 13;7:40354
Gut Homing CD4+ and CD8+ T-Cell Frequencies in HIV Infected Individuals on Antiretroviral Treatment.
Briceño O, Pinto-Cardoso S, Rodríguez-Bernabe N, Murakami-Ogasawara A, Reyes-Terán G
PloS one 2016;11(11):e0166496
PloS one 2016;11(11):e0166496
Adjuvant-dependent innate and adaptive immune signatures of risk of SIVmac251 acquisition.
Vaccari M, Gordon SN, Fourati S, Schifanella L, Liyanage NP, Cameron M, Keele BF, Shen X, Tomaras GD, Billings E, Rao M, Chung AW, Dowell KG, Bailey-Kellogg C, Brown EP, Ackerman ME, Vargas-Inchaustegui DA, Whitney S, Doster MN, Binello N, Pegu P, Montefiori DC, Foulds K, Quinn DS, Donaldson M, Liang F, Loré K, Roederer M, Koup RA, McDermott A, Ma ZM, Miller CJ, Phan TB, Forthal DN, Blackburn M, Caccuri F, Bissa M, Ferrari G, Kalyanaraman V, Ferrari MG, Thompson D, Robert-Guroff M, Ratto-Kim S, Kim JH, Michael NL, Phogat S, Barnett SW, Tartaglia J, Venzon D, Stablein DM, Alter G, Sekaly RP, Franchini G
Nature medicine 2016 Jul;22(7):762-70
Nature medicine 2016 Jul;22(7):762-70
Human Immunodeficiency Virus Type 1 Monoclonal Antibodies Suppress Acute Simian-Human Immunodeficiency Virus Viremia and Limit Seeding of Cell-Associated Viral Reservoirs.
Bolton DL, Pegu A, Wang K, McGinnis K, Nason M, Foulds K, Letukas V, Schmidt SD, Chen X, Todd JP, Lifson JD, Rao S, Michael NL, Robb ML, Mascola JR, Koup RA
Journal of virology 2016 Feb 1;90(3):1321-32
Journal of virology 2016 Feb 1;90(3):1321-32
Collapse of Cytolytic Potential in SIV-Specific CD8+ T Cells Following Acute SIV Infection in Rhesus Macaques.
Roberts ER, Carnathan DG, Li H, Shaw GM, Silvestri G, Betts MR
PLoS pathogens 2016 Dec;12(12):e1006135
PLoS pathogens 2016 Dec;12(12):e1006135
Selective reduction of IFN-γ single positive mycobacteria-specific CD4+ T cells in HIV-1 infected individuals with latent tuberculosis infection.
Riou C, Bunjun R, Müller TL, Kiravu A, Ginbot Z, Oni T, Goliath R, Wilkinson RJ, Burgers WA
Tuberculosis (Edinburgh, Scotland) 2016 Dec;101:25-30
Tuberculosis (Edinburgh, Scotland) 2016 Dec;101:25-30
Temporal Dynamics of CD8+ T Cell Effector Responses during Primary HIV Infection.
Demers KR, Makedonas G, Buggert M, Eller MA, Ratcliffe SJ, Goonetilleke N, Li CK, Eller LA, Rono K, Maganga L, Nitayaphan S, Kibuuka H, Routy JP, Slifka MK, Haynes BF, McMichael AJ, Bernard NF, Robb ML, Betts MR
PLoS pathogens 2016 Aug;12(8):e1005805
PLoS pathogens 2016 Aug;12(8):e1005805
A subset of circulating blood mycobacteria-specific CD4 T cells can predict the time to Mycobacterium tuberculosis sputum culture conversion.
Riou C, Gray CM, Lugongolo M, Gwala T, Kiravu A, Deniso P, Stewart-Isherwood L, Omar SV, Grobusch MP, Coetzee G, Conradie F, Ismail N, Kaplan G, Fallows D
PloS one 2014;9(7):e102178
PloS one 2014;9(7):e102178
Effects of immunosuppression on circulating adeno-associated virus capsid-specific T cells in humans.
Parzych EM, Li H, Yin X, Liu Q, Wu TL, Podsakoff GM, High KA, Levine MH, Ertl HC
Human gene therapy 2013 Apr;24(4):431-42
Human gene therapy 2013 Apr;24(4):431-42
Capsid-specific T-cell responses to natural infections with adeno-associated viruses in humans differ from those of nonhuman primates.
Li H, Lasaro MO, Jia B, Lin SW, Haut LH, High KA, Ertl HC
Molecular therapy : the journal of the American Society of Gene Therapy 2011 Nov;19(11):2021-30
Molecular therapy : the journal of the American Society of Gene Therapy 2011 Nov;19(11):2021-30
Anti-CD8 antibodies can trigger CD8+ T cell effector function in the absence of TCR engagement and improve peptide-MHCI tetramer staining.
Clement M, Ladell K, Ekeruche-Makinde J, Miles JJ, Edwards ES, Dolton G, Williams T, Schauenburg AJ, Cole DK, Lauder SN, Gallimore AM, Godkin AJ, Burrows SR, Price DA, Sewell AK, Wooldridge L
Journal of immunology (Baltimore, Md. : 1950) 2011 Jul 15;187(2):654-63
Journal of immunology (Baltimore, Md. : 1950) 2011 Jul 15;187(2):654-63
No comments: Submit comment
Supportive validation
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
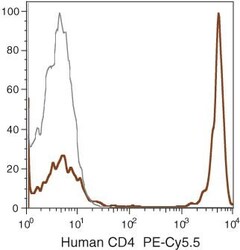
- Experimental details
- Human peripheral blood lymphocytes were stained using PE-Cy®5.5 of anti-human CD4 monoclonal antibody (clone S3.5). The negative control profiles represent unstained cells.
Supportive validation
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
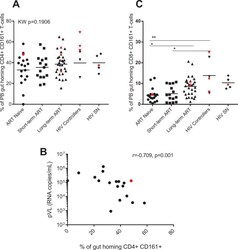
- Fig 3 ART-naive and short-term ART experienced individuals have decreased frequency of CD161-expressing gut homing CD8+ T-cells. Frequencies (expressed as percentages) of peripheral blood (PB) memory gut homing (CD45RO+ CCR9+ B7+) CD4+ (A) and CD8+ (C) T-cells expressing CD161 were determined by flow cytometry on PBMCs of HIV-infected individuals (HIV+), antiretroviral (ART) naive (n = 18), ART experienced individuals on short-term (n = 15) and long-term treatment (n = 32), HIV controllers (n = 6) and HIV seronegative individuals (SN, n = 5). Scatter plots were used to represent the data. Horizontal lines indicate median values. Each symbol represents one individual. The red symbols represent the females in each group. Groups were compared using Kruskal-Wallis test correcting for multiple comparisons using the Dunnett's post-test. Only significant corrected p values are shown ***p
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
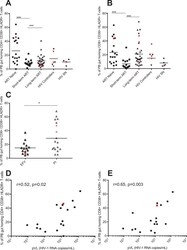
- Fig 4 PB gut-homing CD8+ T activation remains elevated in long-term ART experienced individuals on protease inhibitors. Frequencies (expressed as percentages) of peripheral blood (PB) memory gut homing (CD45RO+ CCR9+ B7+) CD4+ (A) and CD8+ (B) T-cells expressing both CD38 and HLADR was determined by flow cytometry on PBMCs of HIV-infected individuals (HIV+), antiretroviral (ART) naive (n = 18), ART experienced individuals on short-term (n = 15) and long-term treatment (n = 32), HIV controllers (n = 6) and HIV seronegative individuals (SN, n = 5). Frequencies of PB memory gut homing CD8+ T-cells expressing the activation markers CD38 and HLADR in efavirenz (EFV) or protease inhibitors (PI) based therapy is shown in (C). Scatter plots were used to represent the data. Horizontal lines indicate median values. Each symbol represents one individual. The red symbols represent the females in each group. Groups were compared using Kruskal-Wallis test correcting for multiple comparisons using the Dunnett's post-test. Only significant corrected p values are shown ***p
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
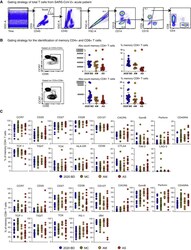
- Figure S1 Quantification and Characterization of CD4 + and CD8 + T Cells in COVID-19, Related to Figure 1 (A) Flow cytometric gating strategy for the identification and quantification of CD4 + and CD8 + T cells. (B) Left: flow cytometric gating strategy for the identification and quantification of memory CD4 + and CD8 + T cells. Right: dot plots summarizing the absolute numbers and relative frequencies of memory CD4 + and CD8 + T cells by group. Each dot represents one donor. Data are shown as median +- IQR. 2020 BD: healthy blood donors from 2020 (n = 18). AM: patients with acute moderate COVID-19 (n = 11). AS: patients with acute severe COVID-19 (n = 17). (C) Dot plots summarizing the expression frequencies of phenotypic markers among memory CD4 + and CD8 + T cells by group. Each dot represents one donor. Bars indicate median values. 2020 BD: healthy blood donors from 2020 (n = 18). MC: individuals in the convalescent phase after mild COVID-19 (n = 31). AM: patients with acute moderate COVID-19 (n = 11). AS: patients with acute severe COVID-19 (n = 17).
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
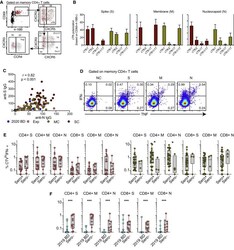
- Figure S5 Functional Polarization of SARS-CoV-2-Specific Memory CD4 + T Cells, Antibody Correlations, and Comparative Analyses of SARS-CoV-2-Specific CD4 + and CD8 + T Cell Responses versus Serostatus in COVID-19, Related to Figures 3 and 4 (A) Representative flow cytometry plots showing the identification of memory CD4 + T cells responding to overlapping peptides spanning the immunogenic domains of the SARS-CoV-2 nucleocapsid protein by subset (AIM assay). Subsets were defined as CXCR5 + (cTfh), CCR4 - CCR6 - CXCR3 + CXCR5 - (Th1), CCR4 + CCR6 - CXCR3 - CXCR5 - (Th2), CCR4 - CCR6 + CXCR3 - CXCR5 - (Th17), CCR4 - CCR6 + CXCR3 + CXCR5 - (Th1/17), and CCR4 - CCR6 - CXCR3 - CXCR5 - (non-Th1/2/17). (B) Bar graphs summarizing the functional polarization of memory CXCR5 + (cTfh) CD4 + T cells responding to overlapping peptides spanning the immunogenic domains of the SARS-CoV-2 spike (S), membrane (M), and nucleocapsid proteins (N). Data are shown as median +- IQR. Key as in C. (C) Correlation between anti-spike (S) and anti-nucleocapsid (N) IgG levels. Each dot represents one donor. 2020 BD: healthy blood donors from 2020 (n = 31). Exp: exposed family members (n = 28). MC: individuals in the convalescent phase after mild COVID-19 (n = 31). SC: individuals in the convalescent phase after severe COVID-19 (n = 23). Spearman rank correlation. (D) Representative flow cytometry plots showing functional SARS-CoV-2-specific memory CD4 + T cell responses in a seronegative convalescent donor
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
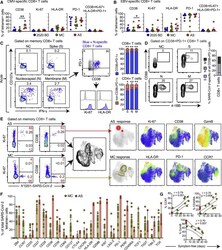
- Figure 2 Phenotypic Characteristics of SARS-CoV-2-Specific T Cells in Acute and Convalescent COVID-19 (A and B) Dot plots summarizing the expression frequencies of activation/cycling markers among tetramer + CMV-specific (A) or EBV-specific CD8 + T cells (B) by group. Each dot represents one specificity in one donor. Data are shown as median +- IQR. * p < 0.05, ** p < 0.01. Kruskal-Wallis rank-sum test with Dunn's post hoc test for multiple comparisons. (C) Representative flow cytometry plots (left) and bar graphs (right) showing the expression of activation/cycling markers among CD107a + and/or IFN-gamma + SARS-CoV-2-specific CD4 + and CD8 + T cells (n = 6 donors). Numbers indicate percentages in the drawn gates. Data are shown as median +- IQR. NC, negative control. * p < 0.05, ** p < 0.01. Paired t test or Wilcoxon signed-rank test. (D) Representative flow cytometry plots (left) and bar graph (right) showing the upregulation of CD69 and 4-1BB (AIM assay) among CD38 + PD-1 + SARS-CoV-2-specific CD8 + T cells (n = 6 donors). Numbers indicate percentages in the drawn gates. S, spike; M, membrane; N, nucleocapsid. (E) Left: representative flow cytometry plots showing the expression of activation/cycling markers among tetramer + SARS-CoV-2-specific CD8 + T cells by group (red) and by total frequency (black). Center: UMAP plot showing the clustering of memory CD8 + T cells. Right: UMAP plots showing the clustering of tetramer + SARS-CoV-2-specific CD8 + T cells by group and the
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
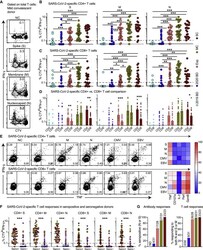
- Figure 4 Antibody Responses and Proliferation Capabilities of SARS-CoV-2-Specific T Cells in Convalescent COVID-19 (A) Representative flow cytometry plots showing the proliferation (CTV - ) and functionality (IFN-gamma + ) of SARS-CoV-2-specific T cells from a convalescent individual (group MC) after stimulation with overlapping peptides spanning the immunogenic domains of the SARS-CoV-2 S, M, and N proteins. Numbers indicate percentages in the drawn gates. (B) and (C) Dot plots summarizing the frequencies of CTV - IFN-gamma + SARS-CoV-2-specific CD4 + (B) and CD8 + T cells (C) by group and specificity. Each dot represents one donor. The dotted line indicates the cutoff for positive responses. * p < 0.05, ** p < 0.01, *** p < 0.001. Kruskal-Wallis rank-sum test with Dunn's post hoc test for multiple comparisons. (D) Dot plots comparing the frequencies of CTV - IFN-gamma + SARS-CoV-2-specific CD4 + versus CD8 + T cells by group and specificity. Each dot represents one donor. Data are shown as median +- IQR. Key as in (B) and (C). * p < 0.05, ** p < 0.01, *** p < 0.001. Paired t test or Wilcoxon signed-rank test. (E) Left: representative flow cytometry plots showing the production of IFN-gamma and TNF among CTV - virus-specific CD4 + (top) and CD8 + T cells (bottom) from a convalescent individual (group MC). Numbers indicate percentages in the drawn gates. Right: heatmaps summarizing the functional profiles of CTV - IFN-gamma + virus-specific CD4 + (top) and CD8 + T cells (bott
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
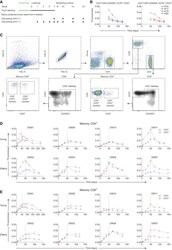
- Figure 1 CD57 - and CD57 + Memory T Cells Exhibit Similar Rates of Deuterium Incorporation (A) Schematic representation of the 2 H 2 O labeling protocol and sampling time points. (B) Experimental labeling data for CD57 - and CD57 + memory CD8 + T cells sampled from the HIV-1-infected volunteers in cohort 1. The corresponding flow cytometric gating strategy is shown in Figure S1 . (C) Successive panels depict the flow cytometric gating strategy used to sort CD57 - and CD57 + memory T cells from the CD4 + and CD8 + lineages (cohort 2). Lymphocytes were identified in a forward scatter-area versus side scatter-area plot, and single cells were identified in a forward scatter-area versus forward scatter-height plot. Boolean gates were drawn for analysis only to exclude fluorochrome aggregates. Viable CD3 + CD14 - CD19 - cells were then identified in the CD4 + and CD8 + lineages, and sort gates were fixed on CD57 - and CD57 + memory cells after exclusion of potentially naive CD27 bright CD45RO - cells. (D) Experimental labeling data for CD57 - and CD57 + memory CD4 + T cells sampled from the healthy volunteers in cohort 2. (E) Experimental labeling data for CD57 - and CD57 + memory CD8 + T cells sampled from the healthy volunteers in cohort 2.
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
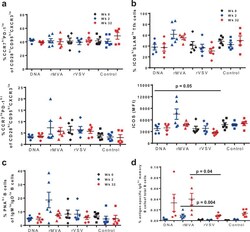
- Fig. 4 Increased expression of ICOS on Tfh CD4+ T cells is associated with induction of Env-specific B-cell responses after priming with rMVA. In each panel, black symbols denote pre-immunization (Week 0), blue symbols post-prime (Week 2) and red symbols show post-boost measurements. Plasmid DNA group is represented by using circles, rMVA by triangles, rVSV by diamonds, and control group by squares. Bars depict SEM. a Both upper and lower panel show the percentages of Tfh CD4+ T cells based on the expression of CCR7 and PD-1 in lymph node-derived lymphocytes. b Upper panel shows the percentages of ICOS hi SLAM lo Tfh CD4+ T cells for individual monkeys from each vaccine group at weeks 0, 2, and 32. Lower panel shows the expression of ICOS per Tfh CD4+ T cell--based on Mean Fluorescence Intensity (MFI). c Dot plots showing accumulated data of relative frequencies (%) of bulk germinal center memory B cells in lymph node (LN) tissues at the indicated time points as judged by PNA staining--memory B cells in LN tissues obtained at the indicated time points. d Immunogen-specific B cells responses were detected by using a specific Env probe in bulk memory B cells from LN tissues. The relative frequencies are shown as a percentage of total B cells. All LN samples were analyzed in one experiment. Non-parametric Mann-Whitney U test was used for the statistical analysis and mean and SEM are shown for each vaccine group.
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
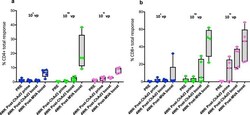
- Fig. 2 Antigen-specific CD4+ and CD8+ T cell in the lung measured for different ChAd 3-5Ag vaccine doses. a Percentage of CD4+ or b CD8+ T cells from lung washes (BAL), producing either IFN-gamma, TNF-alpha, and/or IL-2 (in total) after antigen-specific ex vivo stimulation with peptide pools of individual antigens contained in the respective viral vector constructs. BAL samples were collected before vaccination (PRE), 4 weeks after ChAd3-5Ag priming (study week WK4), ChAd3-5Ag boosting (study week WK14) and MVATG18633 (10 8 PFU) boosting (study week Wk28). vp viral particles. The box extends from the 25th to 75th percentiles. The line in the middle signifies the median and the whiskers include the smallest value up to the largest.
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
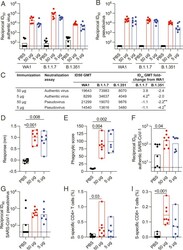
- Fig. 5. Cross-reactive immune responses to SARS-CoV-2 variants and SARS-CoV-1. Serum and PBMC collected 2 wk after the last vaccination was assessed for cross-reactivity to VOCs and SARS-CoV-1. ( A ) Authentic virus and ( B ) pseudovirus neutralizing antibody responses to variants B.1.1.7 and B.1.351. Corresponding responses to SARS-CoV-2 WA1/2020 authentic virus and Wuhan-1 pseudovirus are shown. Bars indicate the GMT. ( C ) Reciprocal ID 50 GMT fold change from wild-type neutralization (WA1 or Wuhan-1) with statistical significance indicated ( P < 0.05, *; P < 0.01, **). ( D ) Serum binding responses to SARS-CoV-1 RBD assessed by biolayer interferometry. ( E ) Antibody-dependent cellular phagocytosis of SARS-CoV-1 S trimer-coated fluorescent beads. ( F ) Authentic SARS-CoV-1 (Urbani) neutralization titers (ID 50 ). ( G ) SARS-CoV-1 (Urbani) pseudovirus neutralization (ID 90 ). ( H ) SARS-CoV-1 (Urbani) S-specific memory CD4+ Th1 and ( I ) CD8+ responses assessed by peptide pool stimulation and ICS (INFgamma, IL-2, and TNF). Significance was assessed with a Kruskal-Wallis test followed by a Dunn's posttest. Bars indicate the GMT.
 Explore
Explore Validate
Validate Learn
Learn Flow cytometry
Flow cytometry