Antibody data
- Antibody Data
- Antigen structure
- References [7]
- Comments [0]
- Validations
- Flow cytometry [1]
- Other assay [5]
Submit
Validation data
Reference
Comment
Report error
- Product number
- 17-1179-42 - Provider product page

- Provider
- Invitrogen Antibodies
- Product name
- CD117 (c-Kit) Monoclonal Antibody (YB5.B8), APC, eBioscience™
- Antibody type
- Monoclonal
- Antigen
- Other
- Description
- Description: The YB5.B8 monoclonal antibody reacts with human CD117, also known as c-Kit, Steel factor receptor and stem cell factor receptor. A member of the tyrosine kinase receptor family, this 145 kDa molecule is expressed by hematopoietic progenitor cell subsets and mast cells. The interaction of c-Kit and Steel factor promotes proliferation and differentiation of hematopoietic progenitor cells and mast cell differentiation. CD117 is also expressed by melanocytes and plays a role in signaling and activation of these cells. When interested in staining HSCs from peripheral blood, we recommend the use of clone 104D2. Please refer to Product # 17-1178. Applications Reported: This YB5.B8 antibody has been reported for use in flow cytometric analysis. Applications Tested: This YB5.B8 antibody has been pre-titrated and tested by flow cytometric analysis of the TF-1 cell line, a human erythroleukemia cell line. This can be used at 5 µL (0.5 µg) per test. A test is defined as the amount (µg) of antibody that will stain a cell sample in a final volume of 100 µL. Cell number should be determined empirically but can range from 10^5 to 10^8 cells/test. Excitation: 633-647 nm; Emission: 660 nm; Laser: Red Laser. Filtration: 0.2 µm post-manufacturing filtered.
- Reactivity
- Human
- Host
- Mouse
- Isotype
- IgG
- Antibody clone number
- YB5.B8
- Vial size
- 100 Tests
- Concentration
- 5 µL/Test
- Storage
- 4° C, store in dark, DO NOT FREEZE!
Submitted references Human Mast Cell Proteome Reveals Unique Lineage, Putative Functions, and Structural Basis for Cell Ablation.
Stimulatory Effects of Mesenchymal Stem Cells on cKit+ Cardiac Stem Cells Are Mediated by SDF1/CXCR4 and SCF/cKit Signaling Pathways.
Immunophenotypic comparison of heterogenous non-sorted versus sorted mononuclear cells from human umbilical cord blood: a novel cell enrichment approach.
Human primordial germ cell commitment in vitro associates with a unique PRDM14 expression profile.
Lineage depletion of stromal vascular fractions isolated from human adipose tissue: a novel approach towards cell enrichment technology.
Oxidative stress leads to increased mutation frequency in a murine model of myelodysplastic syndrome.
CD45/CD11b positive subsets of adult lung anchorage-independent cells harness epithelial stem cells in culture.
Plum T, Wang X, Rettel M, Krijgsveld J, Feyerabend TB, Rodewald HR
Immunity 2020 Feb 18;52(2):404-416.e5
Immunity 2020 Feb 18;52(2):404-416.e5
Stimulatory Effects of Mesenchymal Stem Cells on cKit+ Cardiac Stem Cells Are Mediated by SDF1/CXCR4 and SCF/cKit Signaling Pathways.
Hatzistergos KE, Saur D, Seidler B, Balkan W, Breton M, Valasaki K, Takeuchi LM, Landin AM, Khan A, Hare JM
Circulation research 2016 Sep 30;119(8):921-30
Circulation research 2016 Sep 30;119(8):921-30
Immunophenotypic comparison of heterogenous non-sorted versus sorted mononuclear cells from human umbilical cord blood: a novel cell enrichment approach.
Indumathi S, Harikrishnan R, Rajkumar JS, Dhanasekaran M
Cytotechnology 2015 Jan;67(1):107-14
Cytotechnology 2015 Jan;67(1):107-14
Human primordial germ cell commitment in vitro associates with a unique PRDM14 expression profile.
Sugawa F, Araúzo-Bravo MJ, Yoon J, Kim KP, Aramaki S, Wu G, Stehling M, Psathaki OE, Hübner K, Schöler HR
The EMBO journal 2015 Apr 15;34(8):1009-24
The EMBO journal 2015 Apr 15;34(8):1009-24
Lineage depletion of stromal vascular fractions isolated from human adipose tissue: a novel approach towards cell enrichment technology.
Indumathi S, Mishra R, Harikrishnan R, Rajkumar JS, Kantawala N, Dhanasekaran M
Cytotechnology 2014 Mar;66(2):219-28
Cytotechnology 2014 Mar;66(2):219-28
Oxidative stress leads to increased mutation frequency in a murine model of myelodysplastic syndrome.
Chung YJ, Robert C, Gough SM, Rassool FV, Aplan PD
Leukemia research 2014 Jan;38(1):95-102
Leukemia research 2014 Jan;38(1):95-102
CD45/CD11b positive subsets of adult lung anchorage-independent cells harness epithelial stem cells in culture.
Peter Y, Sen N, Levantini E, Keller S, Ingenito EP, Ciner A, Sackstein R, Shapiro SD
Journal of tissue engineering and regenerative medicine 2013 Jul;7(7):572-83
Journal of tissue engineering and regenerative medicine 2013 Jul;7(7):572-83
No comments: Submit comment
Supportive validation
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- Staining of the TF-1 cell line with Mouse IgG1 kappa Isotype Control APC (Product # 17-4714-81) (open histogram) or Anti-Human CD117 (c-Kit) APC (filled histogram). Total viable cells were used for analysis.
Supportive validation
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
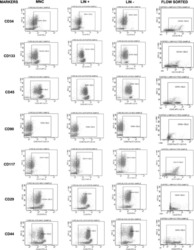

- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
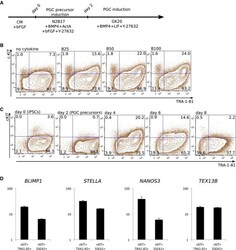
- Figure 2 Induction of PGCLCs from human iPSCs Schematic presentation of PGC-precursor and PGCLC induction. FACS analysis of the concentration-dependent effect of BMP4 on TRA-1-81 + /cKIT + PGCLCs on day 4. B25: 25 ng/ml BMP4; B50: 50 ng/ml BMP4; B100: 100 ng/ml BMP4. FACS analysis of TRA-1-81 and cKIT during PGC induction of up to day 8. Gene expression analysis of selected PGC markers in TRA-1-81 + /cKIT + and SSEA1 + /cKIT + FACS fractions of d4 PGCLCs. Samples were calibrated with iPSC values, and iPSC values depict 1. Data information: Data are presented as means +- SD ( n = 3).
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- Figure 4 Effects of KSR and WNT3A on PGC-precursor and PGCLC induction Effects of KSR and WNT3A during PGC-precursor induction on the expression of selected pluripotency, PGC, and a mesodermal gene of d2 cultures. The value for iPSCs is set as 1, and values are on log 2 scale. Data are presented as means +- SD ( n = 3). Effects of KSR and WNT3A on the induction of d6 PGCLCs as analyzed by FACS gated for TRA-1-81 and cKIT. Gene expression analysis of selected PGC and hematopoietic markers in d6 PGCLCs that were cultured in 0% (control) or 20% KSR (KSR) condition during PGC-precursor induction. Samples were calibrated with iPSC values, and iPSC values depict 1. Data are presented as means +- SD ( n = 3). Immunofluorescence analysis for OCT4 (red), BLIMP1 (green), and T (yellow) in d2 precursors cultured in increasing KSR concentrations. Nuclei were stained with Hoechst (blue). Scale bar: 100 mum.
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- Figure 5 Transcriptomics profiles during PGC-precursor and PGCLC induction Unsupervised hierarchical clustering of HuES6 ESCs, 383.2 iPSCs, d2 PGC-precursor cultures, and FACS-sorted PGCLCs. Principal component analysis of HuES6 ESCs, 393.2 iPSCs, d2 PGC-precursor cultures, and FACS-sorted PGCLCs. Scatter plots of global gene expression microarrays comparing d2 PGC-precursor cultures, or d4 or d6 FACS-sorted PGCLCs with iPSCs. Venn diagrams of the transcriptomics meta-analysis with the intersections of the DEGs of mouse in vitro PGCLCs, human in vivo PGCs, and human in vitro PGC-like cells in relation to their respective pluripotent counterparts in their respective platforms. Heatmap of pluripotent, germ cell, mesodermal, and chromatin-related markers in mouse (left) and human (right) transcriptomics samples ( Tfap2c is not targeted in the Illumina MouseRef-8 v2). The color bars codify the gene expression in log 2 scale. Red corresponds to high gene expression. Blimp1 +/ Stella + PGCLCs are denoted ""B+S+"", and Blimp1 + /Stella - PGCLCs are denoted ""B+S-"". Effect of knockdown of BLIMP1 and PRDM14 on PGCLC induction. Knockdown of BLIMP1 (F) and PRDM14 (G) was done using the lentiviral system with two individual shRNA vectors for each gene. The knockdown efficiencies were assessed by qPCR (left panel, each). The induction of d4 PGCLCs analyzed by FACS, gated for TRA-1-81 and cKIT (right panel, each). Data are presented as means +- SD ( n = 3).
 Explore
Explore Validate
Validate Learn
Learn Flow cytometry
Flow cytometry