Antibody data
- Antibody Data
- Antigen structure
- References [19]
- Comments [0]
- Validations
- Western blot [3]
Submit
Validation data
Reference
Comment
Report error
- Product number
- GTX70143 - Provider product page

- Provider
- GeneTex
- Proper citation
- GeneTex Cat#GTX70143, RRID:AB_372134
- Product name
- DNA ligase III antibody [1F3]
- Antibody type
- Monoclonal
- Reactivity
- Human, Mouse, Chicken/Avian
- Host
- Mouse
Submitted references Inhibition of hydrogen sulfide biosynthesis sensitizes lung adenocarcinoma to chemotherapeutic drugs by inhibiting mitochondrial DNA repair and suppressing cellular bioenergetics.
WRN regulates pathway choice between classical and alternative non-homologous end joining.
Base excision repair defects invoke hypersensitivity to PARP inhibition.
Interaction between DNA Polymerase β and BRCA1.
DNA ligases I and III cooperate in alternative non-homologous end-joining in vertebrates.
Preventing oxidation of cellular XRCC1 affects PARP-mediated DNA damage responses.
Effects of chromatin decondensation on alternative NHEJ.
Functional redundancy between DNA ligases I and III in DNA replication in vertebrate cells.
Pol β associated complex and base excision repair factors in mouse fibroblasts.
Arsenicals affect base excision repair by several mechanisms.
Widespread dependence of backup NHEJ on growth state: ramifications for the use of DNA-PK inhibitors.
DNA polymerase beta-dependent long patch base excision repair in living cells.
Human DNA polymerase beta polymorphism, Arg137Gln, impairs its polymerase activity and interaction with PCNA and the cellular base excision repair capacity.
Rational design of human DNA ligase inhibitors that target cellular DNA replication and repair.
Low levels of DNA ligases III and IV sufficient for effective NHEJ.
ATM mediates oxidative stress-induced dephosphorylation of DNA ligase IIIalpha.
Translocation of XRCC1 and DNA ligase IIIalpha from centrosomes to chromosomes in response to DNA damage in mitotic human cells.
DNA ligase III as a candidate component of backup pathways of nonhomologous end joining.
Physical and functional interaction between DNA ligase IIIalpha and poly(ADP-Ribose) polymerase 1 in DNA single-strand break repair.
Szczesny B, Marcatti M, Zatarain JR, Druzhyna N, Wiktorowicz JE, Nagy P, Hellmich MR, Szabo C
Scientific reports 2016 Nov 3;6:36125
Scientific reports 2016 Nov 3;6:36125
WRN regulates pathway choice between classical and alternative non-homologous end joining.
Shamanna RA, Lu H, de Freitas JK, Tian J, Croteau DL, Bohr VA
Nature communications 2016 Dec 6;7:13785
Nature communications 2016 Dec 6;7:13785
Base excision repair defects invoke hypersensitivity to PARP inhibition.
Horton JK, Stefanick DF, Prasad R, Gassman NR, Kedar PS, Wilson SH
Molecular cancer research : MCR 2014 Aug;12(8):1128-39
Molecular cancer research : MCR 2014 Aug;12(8):1128-39
Interaction between DNA Polymerase β and BRCA1.
Masaoka A, Gassman NR, Horton JK, Kedar PS, Witt KL, Hobbs CA, Kissling GE, Tano K, Asagoshi K, Wilson SH
PloS one 2013;8(6):e66801
PloS one 2013;8(6):e66801
DNA ligases I and III cooperate in alternative non-homologous end-joining in vertebrates.
Paul K, Wang M, Mladenov E, Bencsik-Theilen A, Bednar T, Wu W, Arakawa H, Iliakis G
PloS one 2013;8(3):e59505
PloS one 2013;8(3):e59505
Preventing oxidation of cellular XRCC1 affects PARP-mediated DNA damage responses.
Horton JK, Stefanick DF, Gassman NR, Williams JG, Gabel SA, Cuneo MJ, Prasad R, Kedar PS, Derose EF, Hou EW, London RE, Wilson SH
DNA repair 2013 Sep;12(9):774-85
DNA repair 2013 Sep;12(9):774-85
Effects of chromatin decondensation on alternative NHEJ.
Moscariello M, Iliakis G
DNA repair 2013 Nov;12(11):972-81
DNA repair 2013 Nov;12(11):972-81
Functional redundancy between DNA ligases I and III in DNA replication in vertebrate cells.
Arakawa H, Bednar T, Wang M, Paul K, Mladenov E, Bencsik-Theilen AA, Iliakis G
Nucleic acids research 2012 Mar;40(6):2599-610
Nucleic acids research 2012 Mar;40(6):2599-610
Pol β associated complex and base excision repair factors in mouse fibroblasts.
Prasad R, Williams JG, Hou EW, Wilson SH
Nucleic acids research 2012 Dec;40(22):11571-82
Nucleic acids research 2012 Dec;40(22):11571-82
Arsenicals affect base excision repair by several mechanisms.
Ebert F, Weiss A, Bültemeyer M, Hamann I, Hartwig A, Schwerdtle T
Mutation research 2011 Oct 1;715(1-2):32-41
Mutation research 2011 Oct 1;715(1-2):32-41
Widespread dependence of backup NHEJ on growth state: ramifications for the use of DNA-PK inhibitors.
Singh SK, Wu W, Zhang L, Klammer H, Wang M, Iliakis G
International journal of radiation oncology, biology, physics 2011 Feb 1;79(2):540-8
International journal of radiation oncology, biology, physics 2011 Feb 1;79(2):540-8
DNA polymerase beta-dependent long patch base excision repair in living cells.
Asagoshi K, Liu Y, Masaoka A, Lan L, Prasad R, Horton JK, Brown AR, Wang XH, Bdour HM, Sobol RW, Taylor JS, Yasui A, Wilson SH
DNA repair 2010 Feb 4;9(2):109-19
DNA repair 2010 Feb 4;9(2):109-19
Human DNA polymerase beta polymorphism, Arg137Gln, impairs its polymerase activity and interaction with PCNA and the cellular base excision repair capacity.
Guo Z, Zheng L, Dai H, Zhou M, Xu H, Shen B
Nucleic acids research 2009 Jun;37(10):3431-41
Nucleic acids research 2009 Jun;37(10):3431-41
Rational design of human DNA ligase inhibitors that target cellular DNA replication and repair.
Chen X, Zhong S, Zhu X, Dziegielewska B, Ellenberger T, Wilson GM, MacKerell AD Jr, Tomkinson AE
Cancer research 2008 May 1;68(9):3169-77
Cancer research 2008 May 1;68(9):3169-77
Low levels of DNA ligases III and IV sufficient for effective NHEJ.
Windhofer F, Wu W, Iliakis G
Journal of cellular physiology 2007 Nov;213(2):475-83
Journal of cellular physiology 2007 Nov;213(2):475-83
ATM mediates oxidative stress-induced dephosphorylation of DNA ligase IIIalpha.
Dong Z, Tomkinson AE
Nucleic acids research 2006;34(20):5721-279
Nucleic acids research 2006;34(20):5721-279
Translocation of XRCC1 and DNA ligase IIIalpha from centrosomes to chromosomes in response to DNA damage in mitotic human cells.
Okano S, Lan L, Tomkinson AE, Yasui A
Nucleic acids research 2005;33(1):422-9
Nucleic acids research 2005;33(1):422-9
DNA ligase III as a candidate component of backup pathways of nonhomologous end joining.
Wang H, Rosidi B, Perrault R, Wang M, Zhang L, Windhofer F, Iliakis G
Cancer research 2005 May 15;65(10):4020-30
Cancer research 2005 May 15;65(10):4020-30
Physical and functional interaction between DNA ligase IIIalpha and poly(ADP-Ribose) polymerase 1 in DNA single-strand break repair.
Leppard JB, Dong Z, Mackey ZB, Tomkinson AE
Molecular and cellular biology 2003 Aug;23(16):5919-27
Molecular and cellular biology 2003 Aug;23(16):5919-27
No comments: Submit comment
Supportive validation
- Submitted by
- GeneTex (provider)
- Main image
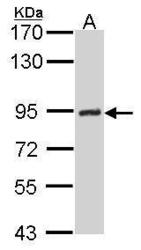
- Experimental details
- Sample (30 ug of whole cell lysate) A: Hela 7.5% SDS PAGE GTX70143 diluted at 1:500
- Validation comment
- WB
- Submitted by
- GeneTex (provider)
- Main image
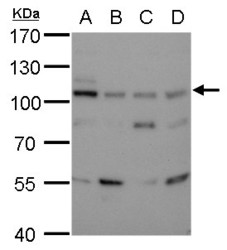
- Experimental details
- DNA ligase III detects DNA ligase III antibody [1F3] protein by western blot analysis.A. 30 ?g 293T whole cell lysate/extractB. 30 ?g A431 whole cell lysate/extractC. 30 ?g HeLa whole cell lysate/extractD. 30 ?g HepG2 whole cell lysate/extract7.5% SDS-PAGEDNA ligase III (GTX70143) dilution: 1:500The HRP-conjugated anti-mouse IgG antibody (GTX213111-01) was used to detect the primary antibody.
- Submitted by
- GeneTex (provider)
- Main image

- Experimental details
- DNA ligase III antibody detects DNA ligase III protein by western blot analysis. Whole cell extracts (30 ?g) was separated by 7.5% SDS-PAGE, and the membrane was blotted with DNA ligase III antibody (GTX70143) diluted by 1:500. The HRP-conjugated anti-mouse IgG antibody (GTX213111-01) was used to detect the primary antibody.
 Explore
Explore Validate
Validate Learn
Learn Western blot
Western blot Immunocytochemistry
Immunocytochemistry