Antibody data
- Antibody Data
- Antigen structure
- References [5]
- Comments [0]
- Validations
- Other assay [15]
Submit
Validation data
Reference
Comment
Report error
- Product number
- 34-1400 - Provider product page

- Provider
- Invitrogen Antibodies
- Product name
- NEDD8 Polyclonal Antibody
- Antibody type
- Polyclonal
- Antigen
- Synthetic peptide
- Reactivity
- Human
- Host
- Rabbit
- Isotype
- IgG
- Vial size
- 100 µg
- Concentration
- 0.25 mg/mL
- Storage
- -20°C
Submitted references COP9-Signalosome deneddylase activity is enhanced by simultaneous neddylation: insights into the regulation of an enzymatic protein complex.
The cyclomodulin cycle inhibiting factor (CIF) alters cullin neddylation dynamics.
UBXN7 docks on neddylated cullin complexes using its UIM motif and causes HIF1α accumulation.
A gatekeeper residue for NEDD8-activating enzyme inhibition by MLN4924.
Development and validation of a method for profiling post-translational modification activities using protein microarrays.
Bornstein G, Grossman C
Cell division 2015;10:5
Cell division 2015;10:5
The cyclomodulin cycle inhibiting factor (CIF) alters cullin neddylation dynamics.
Toro TB, Toth JI, Petroski MD
The Journal of biological chemistry 2013 May 24;288(21):14716-26
The Journal of biological chemistry 2013 May 24;288(21):14716-26
UBXN7 docks on neddylated cullin complexes using its UIM motif and causes HIF1α accumulation.
Bandau S, Knebel A, Gage ZO, Wood NT, Alexandru G
BMC biology 2012 Apr 26;10:36
BMC biology 2012 Apr 26;10:36
A gatekeeper residue for NEDD8-activating enzyme inhibition by MLN4924.
Toth JI, Yang L, Dahl R, Petroski MD
Cell reports 2012 Apr 19;1(4):309-16
Cell reports 2012 Apr 19;1(4):309-16
Development and validation of a method for profiling post-translational modification activities using protein microarrays.
Del Rincón SV, Rogers J, Widschwendter M, Sun D, Sieburg HB, Spruck C
PloS one 2010 Jun 28;5(6):e11332
PloS one 2010 Jun 28;5(6):e11332
No comments: Submit comment
Supportive validation
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
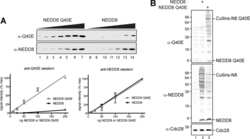
- Figure 4 In vivo validation of substrates ubiquitylated on protein microarrays. (A) Ten putative substrates of ubiquitylation identified on the protein microarrays but not reported in the literature were selected for validation of the modification in vivo . Myc- or GST-substrates were co-expressed with HA-ubiquitin in HEK293T cells. HEK293T cell extracts were prepared using denaturing conditions, substrates immunoprecipitated with anti-Myc or anti-GST antibodies, and ubiquitylation detected by immunoblotting with anti-HA antibodies. Empty vector co-expressed with HA-tagged ubiquitin served as control. Substrates indicated in each lane are: 1- ADRBK2, 2- ACVR1B, 3- PIM2, 4- PRKCgamma, 5- KIF2C, 6- RPS6KA5, 7- ITK, 8- EPHA1, 9- TRIM52, and 10- EPHA5. Of 10 substrates 8 were found to be expressed and immunoprecipitated at detectable levels and of these all demonstrated evidence of ubiquitylation in vivo . To best visualize an ubiquitin smear, substrates 1, 2, 3, 4 were separated by 10% SDS-PAGE gels, while larger molecular weight substrates 5, 6, 7, 8 were separated by 6% SDS-PAGE gels. (B) Ubiquitylation of YY1. HEK293T cells were transfected with plasmids that express HA-ubiquitin, endogenous YY1 protein immunoprecipitated from the denatured extracts, and conjugation to ubiquitin determined by Western blot analysis with anti-HA antibodies (left). Immunoprecipitation efficiency was determined by probing blots with anti-YY1 antibodies (right). Immunoprecipitation with IgG antibo
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- Fig. 2 ATP-depletion resulted in the inhibition of CSN deneddylase activity. a 10 nM His-6-Cul1/ROC1 were pre-neddylated for 30 min at 30degC. Then 10 mug HeLa cells extract were added at 30degC (time plot: lanes 3 - 6 ); or alternatively, ATP-trap mix was added for 45 min at 30degC, followed by the addition of 10 mug HeLa cells extract at 30degC (time plot: lanes 8 - 11 ). b 3 nM His-6-Cul1/ROC1 was pre-neddylated for 30 min at 30degC. Then 100 nM purified CSN was added at 20degC (time plot: lanes 3 - 6 ); or alternatively, ATP-trap mix was added for 45 min at 30degC, followed by the addition of 100 nM purified CSN at 20degC (time plot: lanes 8 - 11 ). c 10 nM His-6-Cul1/ROC1 was incubated as described in b ; time plots were for 1.5-7.5 min, and not for 5-40 min as in b . Blotting was directed against NEDD8, and not against Cul1, as described elsewhere.
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- Fig. 1 Cul1 deneddylation is mediated specifically by CSN. a 1 nM His-6-Cul1/ROC1 was pre-neddylated for 30 min at 30degC; and for 20 min more, at 30degC, following the addition of elevated volumes of HeLa cells extract. Lanes 5 - 6 represent equivalent volumes of HeLa cells extract blotted with anti-Cul1 antibody; this compares the resultant fraction of endogenic Cul1 with its neddylated conjugate, to the extract blotted without the addition of His-6-Cul1/ROC1. Same deneddylation activity was achieved while using reticulocyte lysates as a source for COP9/Signalosome under the same conditions; the image was cropped since most of the work was conducted with HeLa cells extract due to technical reasons and the complete blot is provided as an additional data file termed Additional file 1 . b 10 nM His-6-Cul1/ROC1 were pre-neddylated for 30 min at 30degC, followed by the addition of elevated volumes of HeLa cells extract, and by preparations that were either immunodepleted from JAB1 or that had undergone the same procedure without the antibody itself, as discussed in the methods section. The incubation time with the extract and its parallel derivatives was 20 min at 30degC. c 3 nM His-6-Cul1/ROC1 was pre-neddylated for 30 min at 30degC, followed by the addition of an elevated concentration of purified CSN for 10 min at 20degC. The first part of the image was cropped since it tested different conditions for neddylation and is not relevant to the data exemplified regarding the dened
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- Fig. 3 Reconstitution of ATP with AMP-PNP does not inhibit Cul1 deneddylation. a 3 nM His-6-Cul1/ROC1 was pre-neddylated for 30 min at 30degC, with either ATP or AMP-PNP, then followed with an addition of 100 nM purified CSN, on a time plot of 5-10 min at 20degC. Part of the image was cropped since the experiment also tested the use of CSN after the addition of ATP-trap mix with either ATP or AMP-PNP as was exemplified and discussed in Fig. 2 (regarding the use of ATP). Those parts of the experiment are not relevant for the question of CSN hypothetic autophosphorylation. The complete blot is provided as an additional data file termed Additional file 6 . b 10 nM His-6-Cul1/ROC1 was pre-neddylated for 30 min at 30degC with AMP-PNP. Then 100 nM purified CSN was added at 20oC (time plot: lanes 3 - 6 ); or alternatively, ATP-trap mix was added for 45 min at 30degC, followed by the addition of 100 nM purified CSN at 20degC (time plot: lanes 8 - 11 ).
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
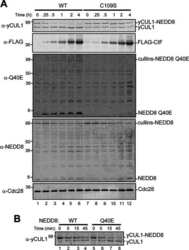
- Fig. 4 Endogenic Cul1 deneddylation is enhanced during simultaneous neddylation. a 20 mug HeLa cells extract were incubated for 30 min at 30degC ( lanes 2 - 7 ) or with the addition of ATP-trap mix prior to the incubation ( lanes 8 - 13 ). Simultaneous ATP-depletion was performed in lanes 3 - 7 after 30 min incubation, on a time plot of 1.5-7.5 min at 30degC. Simultaneous neddylation was performed in lanes 9 - 13 after 30 min, by the addition of AMP-PNP, on a time plot of 1.5-7.5 min at 30degC. b 10 nM His-6-Cul1/ROC1 was neddylated at 30degC on a time plot of 2.5-30 min, without or with the addition of ATP-trap mix ( lanes 2 - 6 and lanes 7 - 11 , respectively). In lane 12 , the neddylation mix was added to the reaction after pre-incubation with ATP-trap mix. c 10 nM His-6-Cul1/ROC1 was neddylated at 30degC using AMP-PNP on a time plot of 2.5-30 min, either directly ( lanes 2 - 6 ) or after pre-incubation with ATP-trap mix ( lanes 7 - 11 ).
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- Fig. 5 Direct inhibition of APP-BP1/UBA3 with MLN-4924 results in inhibition of CSN deneddylase activity. a 10 nM His-6-Cul1/ROC1 was pre-neddylated for 30 min at 30degC. Then 10 mug HeLa cells extract was added, on a time plot at 20degC ( lanes 3 - 6 ); or, alternatively, MLN-4924 was added for 15 min at 30degC, followed by the addition of 10 mug HeLa cells extract, on a time plot at 20degC ( lanes 8 - 11 ). In lane 1 , incubation with MLN-4924 was prior to neddylation. b The same as a using 100 nM purified CSN instead of HeLa cells extract. Both images 5a, b were cropped and the lanes representing the end of the time plot (200 s) were omitted due to technical reasons. The complete blots are provided as additional data files termed Additional file 9 and Additional file 10 , respectively.
 Explore
Explore Validate
Validate Learn
Learn Western blot
Western blot Other assay
Other assay