11-3109-41
antibody from Invitrogen Antibodies
Targeting: HAVCR2
CD366, FLJ14428, Tim-3, TIM3, TIMD3
Antibody data
- Antibody Data
- Antigen structure
- References [8]
- Comments [0]
- Validations
- Flow cytometry [1]
- Other assay [3]
Submit
Validation data
Reference
Comment
Report error
- Product number
- 11-3109-41 - Provider product page

- Provider
- Invitrogen Antibodies
- Product name
- CD366 (TIM3) Monoclonal Antibody (F38-2E2), FITC, eBioscience™
- Antibody type
- Monoclonal
- Antigen
- Other
- Description
- Description: This F38-2E2 monoclonal antibody reacts with human CD366, also known as T cell immunoglobulin and mucin domain-containing protein 3 (TIM3) or HAVCR2. This cell surface receptor is expressed on activated CD4+ T cell subsets (e.g. Th1, Th17, and Treg), CD8+ T cells, monocytes, dendritic cells, and mast cells. Due to alternative splicing, CD366 exists as membrane-bound and soluble forms. Galectin-9 has been identified as the ligand for CD366. In humans, this receptor negatively regulates CD4+ T cells, influencing the secretion of some Th1- and Th17-related cytokines. CD366 has also been implicated in tolerance, autoimmune disease (e.g. multiple sclerosis), and HIV infection. Applications Reported: This F38-2E2 antibody has been reported for use in flow cytometric analysis. Applications Tested: This F38-2E2 antibody has been pre-titrated and tested by flow cytometric analysis of stimulated normal human peripheral blood cells. This can be used at 5 µL (0.125 µg) per test. A test is defined as the amount (µg) of antibody that will stain a cell sample in a final volume of 100 µL. Cell number should be determined empirically but can range from 10^5 to 10^8 cells/test. Excitation: 488 nm; Emission: 520 nm; Laser: Blue Laser. Filtration: 0.2 µm post-manufacturing filtered.
- Reactivity
- Human
- Host
- Mouse
- Conjugate
- Green dye
- Isotype
- IgG
- Antibody clone number
- F38-2E2
- Vial size
- 25 Tests
- Concentration
- 5 µL/Test
- Storage
- 4° C, store in dark, DO NOT FREEZE!
Submitted references TIGIT(+) TIM-3(+) NK cells are correlated with NK cell exhaustion and disease progression in patients with hepatitis B virus‑related hepatocellular carcinoma.
Phosphatidylserine binding directly regulates TIM-3 function.
SLC1A1 mediated glutamine addiction and contributed to natural killer T-cell lymphoma progression with immunotherapeutic potential.
Early IFN-α signatures and persistent dysfunction are distinguishing features of NK cells in severe COVID-19.
Combined p14ARF and Interferon-β Gene Transfer to the Human Melanoma Cell Line SK-MEL-147 Promotes Oncolysis and Immune Activation.
Effect of analytical treatment interruption and reinitiation of antiretroviral therapy on HIV reservoirs and immunologic parameters in infected individuals.
Immunological biomarkers predict HIV-1 viral rebound after treatment interruption.
TIM-3 is expressed on activated human CD4+ T cells and regulates Th1 and Th17 cytokines.
Yu L, Liu X, Wang X, Yan F, Wang P, Jiang Y, Du J, Yang Z
Oncoimmunology 2021;10(1):1942673
Oncoimmunology 2021;10(1):1942673
Phosphatidylserine binding directly regulates TIM-3 function.
Smith CM, Li A, Krishnamurthy N, Lemmon MA
The Biochemical journal 2021 Sep 17;478(17):3331-3349
The Biochemical journal 2021 Sep 17;478(17):3331-3349
SLC1A1 mediated glutamine addiction and contributed to natural killer T-cell lymphoma progression with immunotherapeutic potential.
Xiong J, Wang N, Zhong HJ, Cui BW, Cheng S, Sun R, Chen JY, Xu PP, Cai G, Wang L, Sun XJ, Huang JY, Zhao WL
EBioMedicine 2021 Oct;72:103614
EBioMedicine 2021 Oct;72:103614
Early IFN-α signatures and persistent dysfunction are distinguishing features of NK cells in severe COVID-19.
Krämer B, Knoll R, Bonaguro L, ToVinh M, Raabe J, Astaburuaga-García R, Schulte-Schrepping J, Kaiser KM, Rieke GJ, Bischoff J, Monin MB, Hoffmeister C, Schlabe S, De Domenico E, Reusch N, Händler K, Reynolds G, Blüthgen N, Hack G, Finnemann C, Nischalke HD, Strassburg CP, Stephenson E, Su Y, Gardner L, Yuan D, Chen D, Goldman J, Rosenstiel P, Schmidt SV, Latz E, Hrusovsky K, Ball AJ, Johnson JM, Koenig PA, Schmidt FI, Haniffa M, Heath JR, Kümmerer BM, Keitel V, Jensen B, Stubbemann P, Kurth F, Sander LE, Sawitzki B, Deutsche COVID-19 OMICS Initiative (DeCOI), Aschenbrenner AC, Schultze JL, Nattermann J
Immunity 2021 Nov 9;54(11):2650-2669.e14
Immunity 2021 Nov 9;54(11):2650-2669.e14
Combined p14ARF and Interferon-β Gene Transfer to the Human Melanoma Cell Line SK-MEL-147 Promotes Oncolysis and Immune Activation.
Cerqueira OLD, Clavijo-Salomon MA, Cardoso EC, Citrangulo Tortelli Junior T, Mendonça SA, Barbuto JAM, Strauss BE
Frontiers in immunology 2020;11:576658
Frontiers in immunology 2020;11:576658
Effect of analytical treatment interruption and reinitiation of antiretroviral therapy on HIV reservoirs and immunologic parameters in infected individuals.
Clarridge KE, Blazkova J, Einkauf K, Petrone M, Refsland EW, Justement JS, Shi V, Huiting ED, Seamon CA, Lee GQ, Yu XG, Moir S, Sneller MC, Lichterfeld M, Chun TW
PLoS pathogens 2018 Jan;14(1):e1006792
PLoS pathogens 2018 Jan;14(1):e1006792
Immunological biomarkers predict HIV-1 viral rebound after treatment interruption.
Hurst J, Hoffmann M, Pace M, Williams JP, Thornhill J, Hamlyn E, Meyerowitz J, Willberg C, Koelsch KK, Robinson N, Brown H, Fisher M, Kinloch S, Cooper DA, Schechter M, Tambussi G, Fidler S, Babiker A, Weber J, Kelleher AD, Phillips RE, Frater J
Nature communications 2015 Oct 9;6:8495
Nature communications 2015 Oct 9;6:8495
TIM-3 is expressed on activated human CD4+ T cells and regulates Th1 and Th17 cytokines.
Hastings WD, Anderson DE, Kassam N, Koguchi K, Greenfield EA, Kent SC, Zheng XX, Strom TB, Hafler DA, Kuchroo VK
European journal of immunology 2009 Sep;39(9):2492-501
European journal of immunology 2009 Sep;39(9):2492-501
No comments: Submit comment
Supportive validation
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
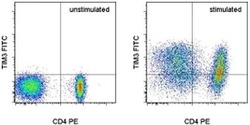
- Experimental details
- Staining of unstimulated (left) or Anti-Human CD3 and CD28 Functional Grade Purified (Product # 16-0037-81 and Product # 16-0289-81)-stimulated (right) normal human peripheral blood cells with Anti-Human CD4 PE (Product # 12-0047-42) and Anti-Human CD366 (TIM3) FITC. Total viable cells, as determined by Fixable Viability Dye eFluor® 660 (Product # 65-0864-14), were used for analysis.
- Conjugate
- Green dye
Supportive validation
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
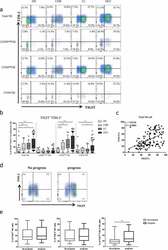

- Experimental details
- Figure2. The co-expression of TIGIT and TIM-3 is elevated on NK cells of progression patients with HBV-HCC. a-b Percentages of TIGIT + TIM-3 + NK cells on total NK cells and NK cell subsets(CD56 bright NK cells,CD56 dim NK cells, and CD56 - NK cells) from HBV-HCC, HDs, CHB and HBV-LC patients by flow cytometry analysis. c Correlation analysis of TIGIT and TIM-3 on NK cells from patients with HBV-HCC. d-e Flow-cytometry analyses (d) of TIGIT and TIM-3 were performed on PBMCs collected from HBV-HCC patients. Representative plots (e) display the expression of TIGIT + NK cells, TIM-3 + NK cells and total TIGIT + TIM-3 + NK cells from patients with progression (n = 61) and no progression (n = 72). P values were calculated by using the Kruskal-Wallis nonparametric H test (a-c). P values were obtained by the unpaired t test (d-e). * P < .05, ** P < .01, *** P < .001, **** P < .0001
- Conjugate
- Green dye
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- Fig. 5 Asparaginase treatment increased NKTCL cell sensitivity to anti-PD-1 antibody. (a) SLC1A1 expression on NK-92 cells transfected with SLC1A1 vector or control vector (upper panel) and SNK-6 cells transfected with SLC1A1 shRNA or scramble (lower panel). (b) Ki-67 and TIM-3 positivity of CD3+/CD8+ T cells in PBMC co-cultured with NK-92 cells (upper panel) or SNK-6 cells (lower panel) transfected with indicated vectors or shRNAs in medium with or without extra glutamine (2mM). (c) PD-L1 mRNA expression in NK-92 cells transfected with SLC1A1 vector or control vector (upper panel) and SNK-6 cells transfected with SLC1A1 shRNA or scramble (lower panel) upon asparaginase (10 IU/mL) treatment. The control vector or scramble values were normalized to 1, respectively. (d and e) Median fluorescence intensity of PD-L1 (d) on NK-92 cells (upper panel) or SNK-6 cells (lower panel), as well as Ki-67 and TIM-3 positivity of CD3+/CD8+ T cells in PBMC co-cultured with NK-92 cells (upper panel) or SNK-6 cells (lower panel) upon indicated treatment. (f) Gene expression correlation of tumor SLC1A1 with PD-L1 in NKTCL patients (n=128). (g) Tumor EAAT3 expression according to the TSIM, HEA, and MB subtypes in NKTCL patients (n=100). (h) PD-L1 mRNA expression of NK-92 cells transfected with TP53 R248Q, TP53 R273H, EP300 vector, or MGA shRNA upon indicated treatment. (i) Ki-67 positivity of CD3+/CD8+ T cells in PBMC co-cultured with NK-92 cells transfected with TP53 R248Q, TP53 R273H, EP300 vec
- Conjugate
- Green dye
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
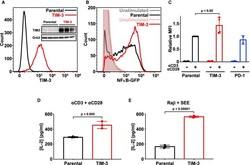
- Experimental details
- Figure 1. Ectopic expression of TIM-3 enhances activation of Jurkat cells. ( A ) Flow cytometry analysis of human TIM-3 expression in parental (black) and TIM-3 lentivirus-transduced NF-kappaB GFP reporter Jurkat cells (red), detected with phycoerythrin-conjugated TIM-3 antibody. The inset shows Western blot analysis of TIM-3 expression in the same cells as marked -- with three biological repeats of each. ( B ) Representative histogram of NF-kappaB-driven GFP expression in stimulated NF-kappaB reporter cells (black), and those expressing TIM-3 (red) -- compared with unstimulated parental (gray) and TIM-3-expressing (pink) cells. Stimulation employed 1 mug/ml alphaCD3 plus 1 mug/ml alphaCD28 for 16 h to mimic antigen binding. T cell activation was quantified by flow cytometry analysis of the GFP reporter of NFkappaB transcriptional activity. Enhanced T cell activation in TIM-3-expressing cells requires cell stimulation, with no change being observed in unstimulated cells. ( C ) Mean GFP fluorescence intensity (MFI) values for NF-kappaB GFP reporter assays in cells stimulated as in ( B ) relative to parental NF-kappaB reporter cells (black). Data represent the mean +- standard deviation (SD) for four biological repeats for parental and TIM-3-expressing cells (red) and two repeats for PD-1-expressing cells (blue). P -value for the TIM-3/parental comparison was determined with a two-tailed, unpaired Student's t -test. ( D ) ELISA analysis of IL-2 secretion into the medium of pare
- Conjugate
- Green dye
 Explore
Explore Validate
Validate Learn
Learn Flow cytometry
Flow cytometry