PA5-17505
antibody from Invitrogen Antibodies
Targeting: SPI1
OF, PU.1, SFPI1, SPI-1, SPI-A
 Western blot
Western blot Immunoprecipitation
Immunoprecipitation Immunohistochemistry
Immunohistochemistry Flow cytometry
Flow cytometry Chromatin Immunoprecipitation
Chromatin Immunoprecipitation Other assay
Other assayAntibody data
- Antibody Data
- Antigen structure
- References [9]
- Comments [0]
- Validations
- Western blot [2]
- Immunohistochemistry [1]
- Other assay [12]
Submit
Validation data
Reference
Comment
Report error
- Product number
- PA5-17505 - Provider product page

- Provider
- Invitrogen Antibodies
- Product name
- PU.1 Polyclonal Antibody
- Antibody type
- Polyclonal
- Antigen
- Synthetic peptide
- Description
- It is not recommended to aliquot this antibody.
- Reactivity
- Human
- Host
- Rabbit
- Isotype
- IgG
- Vial size
- 100 µL
- Concentration
- 20 µg/mL
- Storage
- -20°C
Submitted references LINC01094/SPI1/CCL7 Axis Promotes Macrophage Accumulation in Lung Adenocarcinoma and Tumor Cell Dissemination.
Chromatin Remodeling of Colorectal Cancer Liver Metastasis is Mediated by an HGF-PU.1-DPP4 Axis.
SIRT1/2 orchestrate acquisition of DNA methylation and loss of histone H3 activating marks to prevent premature activation of inflammatory genes in macrophages.
Differential regulation of TREM2 and CSF1R in CNS macrophages in an SIV/macaque model of HIV CNS disease.
The 10q26 Risk Haplotype of Age-Related Macular Degeneration Aggravates Subretinal Inflammation by Impairing Monocyte Elimination.
Cytotoxicity and Differentiating Effect of the Poly(ADP-Ribose) Polymerase Inhibitor Olaparib in Myelodysplastic Syndromes.
RAG-Mediated DNA Breaks Attenuate PU.1 Activity in Early B Cells through Activation of a SPIC-BCLAF1 Complex.
Suppression of SRCAP chromatin remodelling complex and restriction of lymphoid lineage commitment by Pcid2.
TAM receptor-dependent regulation of SOCS3 and MAPKs contributes to proinflammatory cytokine downregulation following chronic NOD2 stimulation of human macrophages.
Wu Z, Bai X, Lu Z, Liu S, Jiang H
Journal of immunology research 2022;2022:6450721
Journal of immunology research 2022;2022:6450721
Chromatin Remodeling of Colorectal Cancer Liver Metastasis is Mediated by an HGF-PU.1-DPP4 Axis.
Wang L, Wang E, Prado Balcazar J, Wu Z, Xiang K, Wang Y, Huang Q, Negrete M, Chen KY, Li W, Fu Y, Dohlman A, Mines R, Zhang L, Kobayashi Y, Chen T, Shi G, Shen JP, Kopetz S, Tata PR, Moreno V, Gersbach C, Crawford G, Hsu D, Huang E, Bu P, Shen X
Advanced science (Weinheim, Baden-Wurttemberg, Germany) 2021 Oct;8(19):e2004673
Advanced science (Weinheim, Baden-Wurttemberg, Germany) 2021 Oct;8(19):e2004673
SIRT1/2 orchestrate acquisition of DNA methylation and loss of histone H3 activating marks to prevent premature activation of inflammatory genes in macrophages.
Li T, Garcia-Gomez A, Morante-Palacios O, Ciudad L, Özkaramehmet S, Van Dijck E, Rodríguez-Ubreva J, Vaquero A, Ballestar E
Nucleic acids research 2020 Jan 24;48(2):665-681
Nucleic acids research 2020 Jan 24;48(2):665-681
Differential regulation of TREM2 and CSF1R in CNS macrophages in an SIV/macaque model of HIV CNS disease.
Knight AC, Brill SA, Solis CV, Richardson MR, McCarron ME, Queen SE, Bailey CC, Mankowski JL
Journal of neurovirology 2020 Aug;26(4):511-519
Journal of neurovirology 2020 Aug;26(4):511-519
The 10q26 Risk Haplotype of Age-Related Macular Degeneration Aggravates Subretinal Inflammation by Impairing Monocyte Elimination.
Beguier F, Housset M, Roubeix C, Augustin S, Zagar Y, Nous C, Mathis T, Eandi C, Benchaboune M, Drame-Maigné A, Carpentier W, Chardonnet S, Touhami S, Blot G, Conart JB, Charles-Messance H, Potey A, Girmens JF, Paques M, Blond F, Leveillard T, Koertvely E, Roger JE, Sahel JA, Sapieha P, Delarasse C, Guillonneau X, Sennlaub F
Immunity 2020 Aug 18;53(2):429-441.e8
Immunity 2020 Aug 18;53(2):429-441.e8
Cytotoxicity and Differentiating Effect of the Poly(ADP-Ribose) Polymerase Inhibitor Olaparib in Myelodysplastic Syndromes.
Faraoni I, Consalvo MI, Aloisio F, Fabiani E, Giansanti M, Di Cristino F, Falconi G, Tentori L, Di Veroli A, Curzi P, Maurillo L, Niscola P, Lo-Coco F, Graziani G, Voso MT
Cancers 2019 Sep 16;11(9)
Cancers 2019 Sep 16;11(9)
RAG-Mediated DNA Breaks Attenuate PU.1 Activity in Early B Cells through Activation of a SPIC-BCLAF1 Complex.
Soodgupta D, White LS, Yang W, Johnston R, Andrews JM, Kohyama M, Murphy KM, Mosammaparast N, Payton JE, Bednarski JJ
Cell reports 2019 Oct 22;29(4):829-843.e5
Cell reports 2019 Oct 22;29(4):829-843.e5
Suppression of SRCAP chromatin remodelling complex and restriction of lymphoid lineage commitment by Pcid2.
Ye B, Liu B, Yang L, Huang G, Hao L, Xia P, Wang S, Du Y, Qin X, Zhu P, Wu J, Sakaguchi N, Zhang J, Fan Z
Nature communications 2017 Nov 15;8(1):1518
Nature communications 2017 Nov 15;8(1):1518
TAM receptor-dependent regulation of SOCS3 and MAPKs contributes to proinflammatory cytokine downregulation following chronic NOD2 stimulation of human macrophages.
Zheng S, Hedl M, Abraham C
Journal of immunology (Baltimore, Md. : 1950) 2015 Feb 15;194(4):1928-37
Journal of immunology (Baltimore, Md. : 1950) 2015 Feb 15;194(4):1928-37
No comments: Submit comment
Supportive validation
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
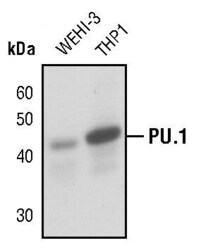
- Experimental details
- Western blot analysis of PU.1 in extracts from WEHI-3 and THP1 cells using PU.1 polyclonal antibody (Product # PA5-17505).
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
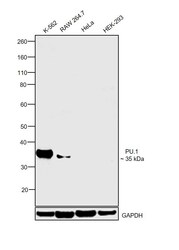

- Experimental details
- Western blot was performed using Anti-PU.1 Polyclonal Antibody (Product # PA5-17505) and a 35 kDa band corresponding to PU.1 was observed across cell lines tested except HeLa and HEK-293 which are reported to be negative. Modified whole cell extracts (1% SDS) (30 µg lysate) of K-562 (Lane 1), RAW 264.7 (Lane 2), HeLa (Lane 3) and HEK-293 (Lane 4) were electrophoresed using NuPAGE™ 4-12% Bis-Tris Protein Gel (Product # NP0322BOX). Resolved proteins were then transferred onto a nitrocellulose membrane (Product # IB23001) by iBlot® 2 Dry Blotting System (Product # IB21001). The blot was probed with the primary antibody (1:1000 dilution) and detected by chemiluminescence with Goat anti-Rabbit IgG (H+L) Superclonal™ Recombinant Secondary Antibody, HRP (Product # A27036, 1:4000 dilution) using the iBright FL 1000 (Product # A32752). Chemiluminescent detection was performed using SuperSignal™ West Dura Extended Duration Substrate (Product # 34076).
Supportive validation
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- Immunohistochemical analysis of PU.1 in paraffin-embedded human tonsil using a PU.1 polyclonal antibody (Product # PA5-17505) showing nuclear localization.
Supportive validation
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- Figure 4 PU.1 upregulates DPP4 expression through remodeling DPP4-associated chromatin. A) Integrated analysis of JASPAR, RNA-seq, and ATAC-seq revealing three potential DPP4 epigenetic regulators. B) Corresponding scores of the potential DPP4 regulators. C) Nucleosome occupancy analysis of enriched or constitutive peaks from liver metastases or primary CRC centered at PU.1 motif with 300 bp flanking window. D) ChIP-qPCR showing PU.1 binding in DPP4 promoter and enhancer regions. E,F) ChIP-qPCR showing PU.1 binding to DPP4 promoter and enhancers in HT29-derived liver metastases versus primary tumors from E) CRC orthotopic-model, and F) patient-derived CRC organoids. G) Western blots showing DPP4 expression levels in HT29 or CRC57 carrying scrambled (control) or PU.1 knockdown (PU.1 KD1 or KD2) shRNAs constructs. H) ChIP-qPCR showing relative H3K27ac, H3K4me1, H3K4me3, and H3K9ac enrichments in HT29 or CRC57 carrying scrambled (control) or PU.1 knockdown (PU.1 KD1) shRNA constructs. LM, liver metastases. CP, CRC primary tumor. E1, E2, and E3, enhancer 1, enhancer 2, and enhancer 3. Data represent the mean +- s.d. p -values were calculated based on Student's t -test. * p < 0.05; ** p < 0.01; *** p < 0.001; **** p < 0.0001.
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- Figure 5 PU.1 is upregulated in CRC liver metastases and promotes CRC progression. A) Analysis of differential PU.1 expression in GEO dataset (GSE41568) between primary CRC and liver metastases. B) Analysis of differential PU.1 expression in the RNA-seq dataset SRR2089755 from five matched patient primary CRC and liver metastases. C) IHC staining of differential PU.1 expression between primary CRC and liver metastases. (Scale bar, 100 um). D) Western blots showing DPP4 expression levels in paired primary CRC and liver metastases collected from four patients. E) Western blots showing DPP4 expression levels in paired primary CRC and liver metastases-derived organoids collected from two patients (CA197 and CA1006). F,G) Images and quantification of F) bioluminescence and G) survival analysis of NSG mice injected with luciferase-labeled HT29 carrying scrambled (control) or PU.1 knockdown (PU.1 KD1 or KD2) shRNA constructs. H,J) Images and quantification of H) bioluminescence and J) survival analysis of NSG mice injected with luciferase-labeled CRC57 carrying scrambled (control) or PU.1 knockdown (PU.1 KD1 or KD2) shRNA constructs. LM, liver metastases. CP, CRC primary tumor. Data represent the mean +- s.e.m. in (A) and (B), and the mean +- s.d. in (F) and (H). p -values were calculated based on Student's t -test (A) and (B), log-rank test in (G) and (J), and ANOVA and Tukey's HSD post hoc test in (F) and (H). * p < 0.05; ** p < 0.01; *** p < 0.001; **** p < 0.0001.
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- Figure 6 HGF upregulates DPP4 through activaton of PU.1. A) Dotplot of pathway analysis on enriched accessible chromatin regions in HT29 liver metatases. B) Western blots showing PU.1 expression level in HT29 or CRC57 cells treated with or without HGF. C) Western blots showing PU.1 phosphorylation in HT29 or CRC57 cells treated with or without HGF or the c-Met inhibitor PHA665752. D,E) Western blots showing that PU.1 complexed with D) C/EBP alpha and - beta and E) SMARCB1. F) ChIP-qPCR showing relative H3K27ac, H3K4me1, H3K4me3, and H3K9ac enrichments in HT29 or CRC57 cultured in RPMI1640 media supplemented with or without 100 ng mL -1 HGF. G) Western blots showing the expressions of DPP4 in HT29 or CRC57 carrying scrambled (control) or PU.1 knockdown shRNA cultured in RPMI media supplemented with or without 100 ng mL -1 HGF. H,I) Images and quantification of H) bioluminescence and I) survival analysis of NSG mice spleen-injected with luciferase-labeled HT29 cells receiving i.p. administered PBS (control) or the HGF inhibitor Norleual. Data represent the mean +- s.d. in (F) and (H). p -values were calculated based on Student's t -test (F) and log-rank test in (I). * p < 0.05; ** p < 0.01; *** p < 0.001; **** p < 0.0001.
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- Figure 5 SPI1 promotes CCL7 transcription. (a) Transcription factors that can bind to CCL7 promoter with an over 0.5 correlation coefficient in TCGA-LUAD analyzed using the R TFBS package; (b) binding sites between SPI1 and CCL7 promoter; (c) conservative binding sequence of SPI1; (d) a sketch map for the fragmentation of CCL7 promoter sites using the Crispr-Cas9 system; mRNA (e) and protein (f) levels of CCL7 in A549 cells after knockout of A, B, and C fragments alone or simultaneously; (g) CCL7 promoter fragments enriched by anti-SPI1 examined by the ChIP-qPCR assay; (h) construction of the pGL4-Luc luciferase vector containing the CCL7 promoter sequence for the examination of SPI1-CCL7 binding by luciferase assay. Repetition = 3. Data are presented as the mean +- SD. ** p < 0.01.
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- Overexpression of SPI1 in CCL7-low cells promotes macrophage migration and M2 polarization. mRNA (a) and protein (b) levels of SPI1 and CCL7 in H125 CCL7 Low and A549 CCL7 High cells after SPI1 overexpression plasmid or shRNA transfection detected by qPCR and WB analyses; (c) number of migrated THP-1 cells examined by crystal violet staining; mRNA (d) and protein (e) levels of CD86 and CD206 in cocultured THP-1 cells examined by qPCR and WB analyses; (f) expression of M1 cytokines (IL-6, IL-1 beta , and TNF- alpha ) and M2 cytokines (IL-10 and TGF- beta ) in the culture medium determined by ELISA kits. Repetition = 3. Data are presented as the mean +- SD. ** p < 0.01.
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- Figure 1. RAG DSB Signals Induce Genome-wide Changes in PU.1 Binding (A) qPCR analysis of Igk genomic DNA from Rag1 -/- :Bcl2 (red) and Art -/- :Bcl2 (blue) abl pre-B cells treated with imatinib for 48 h. Schematic shows germline(GL) Igk locus and unrepaired Jkappa1 coding end withlocation of PCR primers. PCR is normalizedto Rag1 -/- :Bcl2 abl pre-B cells, which do not generate RAG DSBs and have only intact germline Igk DNA. Data are representative of three independentexperiments. (B) Dot plot and heatmap of fold changes and signal Intensity for PU.1peaks Identified by ChIP-seq In Rag1 -/- :Bcl2 and Art -/- :Bcl2 ablpre-B cells treated with Imatinib for 48 h. Data are from common peaksidentified in two replicates for each cell. (C) Representative tracks at indicated regions for PU.1 ChIP-seq from(B). ChIP-qPCR validation for PU.1 binding at each locus is also shown. Data aremean and SE for three independent experiments. **p
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- Figure 2. Expression of SPIC Alters PU.1 Binding (A) Spic mRNA expression in Rag1 -/- :Bcl2 and Art -/- :Bcl2 ablpre-B cells treated with imatinib for 48 h. Art -/- :Bcl2 ablpre-B cells were also treated with vehicle (-) or 15 muM ATMinhibitor KU55933 (+ iATM). Data are relative to Rag1 -/- :Bcl2 andare mean and SE for three independent experiments. (B) Spic mRNA expression in Rag1 -/- :Bcl2:Spic tet abl pre-B cells treated with imatinib alone (-) or with imatinib and 2muM doxycycline (Dox; +) for 48 h. Data are relative to Rag1 -/- :Bcl2:Spic tet without doxycycline and are mean and SE for three independent experiments. (C) Western blot shows PU.1 and SPIC (determined by anti-FLAG antibody)in Rag1 -/- :Bcl2:Spic tet abl pre-B cells treated as in (B). Data are representative of three independentexperiments. (D) Dot plot and heatmap of fold changes and signal intensity for PU.1peaks identified by ChIP-seq in Rag1 -/- :Bcl2:Spic tet abl pre-B cells treated with imatinib alone (- Dox, no SPIC) or withimatinib and 2 muM doxycycline (+ Dox, + SPIC) for 48 h as in (B). Dataare from common peaks identified in two replicates for each cell line. *p
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- Figure 3. SPIC and PU.1 Bind to Identical Genomic Regions (A) Dot plot and heatmap of fold changes and signal intensity for PU.1and SPIC (by anti-HA ChIP) peaks identified by ChIP-seq in Rag1 -/- :Bcl2:Spic tet abl pre-B cells treated with imatinib for 48 h in the absence (for PU.1 ChIP) orpresence (for SPIC ChIP) of 2 muM doxycycline (Dox). Data are from commonpeaks identified in two replicates of each cell line. (B) Representative ChIP-seq binding of PU.1 and SPIC at indicatedregions. PU.1 ChIP-seq was performed in Rag1 -/- :Bcl2:Spic tet abl pre-B cells treated with imatinib alone (- Dox, no SPIC) or withimatinib and doxycycline to induce expression of SPIC(+ Dox, + SPIC) for 48 h.ChIP-seq for SPIC was performed as in A in Rag1 -/- :Bd2:Spic tet abl pre-B cells treated with imatinib and doxycycline for 48 h. (C) ChIP-qPCR validation for PU.1 and SPIC binding at each locus shownin (B). Data are mean and SE for three independent experiments. **p
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- Figure 6. BCLAF1 Regulates SYK Expression in Primary Pre-B Cells (A-C) Art -/- :muIgh:Bcl2 pre-B cells were transduced with a retrovirus expressing a scrambled shRNA(-) or shBclaf1 (+) and then subsequently withdrawn from IL-7. (A and B) Spic and Syk mRNA expressionassessed in indicated small pre-B cells 2 days after IL-7 withdrawal. Data aremean and SE for three independent experiments. (C) Western blot of SYK and BCLAF1 in indicated small pre-B cells 2 daysafter IL-7 withdrawal. Data are representative of three independentexperiments. (D) Flow cytometric analysis showing EGFP (y axis) and FSC (x axis) inbone marrow pre-B cells(B220 lo CD43 - IgM - ) fromwild-type and Spic igfp / igfp mice. Data are representative of five independent experiments. (E) Percentage of EGFP-positive small pre-B cells in Spic igfp / igfp (circles) and Atm -/- :Spic igfp / igfp (squares) mice was quantified by flow cytometry as in (D). Data are mean and SEfrom three independent mice of each genotype. (F-H) Syk mRNA expression (F), ChIP-PCR of PU.1at Syk promoter (G), and ChIP-PCR of BCLAF1 at Syk promoter (H) in EGFP-negative (-) andEGFP-expressing (+) small pre-B cells sorted from Spic igfp / igfp mice. Data in (F) are the mean and SE from three independent experiments. Datain (G) and (H) are representative of two independent experiments. *p
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- Figure 6. SIRT1 and 2 interact with DNMTs to drive DNA hypermethylation at inflammatory loci. (A) Co-immunoprecipitation assays were performed in MOs differentiated to MACs in the presence of M-CSF for 5 days. Protein extracts were immunoprecipitated utilising anti-DNMT1, -DNMT3A, -DNMT3B, -SIRT1 and -SIRT2 antibodies, in which IgG was used as a negative control and total protein extract was used as input. (B) Co-immunoprecipitation of MACs differentiated in the presence of DMSO or 50 uM cambinol for 5 days. (C) Chromatin immunoprecipitation of two independent experiments of SIRT1 (pink), SIRT2 (red) and PU.1 (blue) was performed in isolated monocytes and MACs differentiated in the presence of M-CSF for 5 days in the presence and absence of cambinol, before and after LPS stimulation for 18 hrs. (D) MOs were transfected with 100nM of corresponding siRNA and differentiated to MACs in the presence of M-CSF for 5 days, as described in Materials and Methods. Annotated numbers indicate relative band intensities compared to non-targeting control (NT). (E) Pyrosequencing of hypermethylated CpGs in MOs and MACs transfected with NT, siSIRT1 and siSIRT2 prior to LPS activation. (F) Gene expression of SCL1A2 , TNFAIP3 , JAK3 , RUNX3 and ADORA2A in MOs and LPS-activated MACs transfected with NT, siSIRT1 and siSIRT2, as normalised against RPL38 . (D-F) Statistical significance of at least three independent experiments was calculated using paired student t-tests (* p-value < 0.05, ** p-valu
 Explore
Explore Validate
Validate Learn
Learn