Antibody data
- Antibody Data
- Antigen structure
- References [0]
- Comments [0]
- Validations
- Western blot [2]
- Immunocytochemistry [1]
- Flow cytometry [1]
- Other assay [52]
Submit
Validation data
Reference
Comment
Report error
- Product number
- 32-2500 - Provider product page

- Provider
- Invitrogen Antibodies
- Product name
- alpha Tubulin Monoclonal Antibody (B-5-1-2)
- Antibody type
- Monoclonal
- Antigen
- Other
- Reactivity
- Human, Mouse, Rat, Canine
- Host
- Mouse
- Isotype
- IgG
- Antibody clone number
- B-5-1-2
- Vial size
- 100 µg
- Concentration
- 0.5 mg/mL
- Storage
- -20°C
No comments: Submit comment
Supportive validation
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- Western blot analysis of Alpha-tubulin was performed by loading 20 µg of U2OS (lane1), HEK-293 (lane2), HeLa (lane3), MCF7 (lane4) and Jurkat (lane5) cell lysate using Novex® NuPAGE® 4-12 % Bis-Tris gel (Product # NP0321BOX), XCell SureLock Electrophoresis System (Product # EI0002), Novex® Sharp Pre-Stained Protein Standard (LC5800), and iBlot® Dry Blotting System (IB21001). Proteins were transferred to a nitrocellulose membrane and blocked with 5 % skim milk for 1 hour at room temperature. Alpha-Tubulin was detected at ~55 kDa using Alpha-Tubulin Mouse Monoclonal Antibody (Product # 32-2500) at 0.5-1 µg/mL in 2.5 % skim milk at 4°C overnight on a rocking platform. Goat Anti-Mouse IgG - HRP Secondary Antibody (Product # 62-6520) at 1:4000 dilution was used and chemiluminescent detection was performed using Pierce™ ECL Western Blotting Substrate (Product # 32106).
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- Western blot analysis was performed on whole cell extracts (30µg lysate) of COS-7 (Lane 1), MDCK (Lane 2), NIH/3T3 (Lane 3), MDA-MB-231 (Lane 4), L6 (Lane 5), RSC96 (Lane 6), tissue extracts of Mouse Lung (Lane 7), Rat Liver (Lane 8) and Mouse Pancreas (Lane 9). The blot was probed with Anti- alpha Tubulin Mouse Monoclonal Antibody (Product # 32-2500, 1µg/mL) and detected by chemiluminescence using Goat anti-Mouse IgG (H+L) Superclonal™ Secondary Antibody, HRP conjugate (Product # A28177, 0.25µg/mL, 1:4000 dilution). A 52 kDa band corresponding to alpha Tubulin was observed across the cell lines and tissues tested. Known quantity of protein samples were electrophoresed using Novex® NuPAGE® 4-12 % Bis-Tris gel (Product # NP0322BOX), XCell SureLock™ Electrophoresis System (Product # EI0002) and Novex® Sharp Pre-Stained Protein Standard (Product # LC5800). Resolved proteins were then transferred onto a nitrocellulose membrane with iBlot® 2 Dry Blotting System (Product # IB21001). The membrane was probed with the relevant primary and secondary Antibody following blocking with 5 % skimmed milk. Chemiluminescent detection was performed using Pierce™ ECL Western Blotting Substrate (Product # 32106).
Supportive validation
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- Immunofluorescent analysis of Alpha - Tubulin was done on 70% confluent log phase U-2 OS cells. The cells were fixed with 4% paraformaldehyde for 15 minutes, permeabilized with 0.25% Triton X-100 for 10 minutes, and blocked with 5% BSA for 1 hour at room temperature. The cells were labeled with Alpha - Tubulin Mouse Monoclonal Antibody (Product # 32-2500) at 0.5 µg/mL in 1% BSA and incubated for 3 hours at room temperature and then labeled with Alexa Fluor 488 Rabbit Anti-Mouse IgG Secondary Antibody (Product # A-11059) at a dilution of 1:400 for 30 minutes at room temperature (Panel a: green). Nuclei (Panel b: blue) were stained with SlowFade Gold Antifade Mountant with DAPI (Product # S36938). F-actin (Panel c: red) was stained with Alexa Fluor 594 Phalloidin (Product # A12381). Panel d is a merged image showing cytoplasmic localization. Panel e shows no primary antibody control. The images were captured at 20X magnification.
Supportive validation
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- Flow cytometry analysis of Alpha-tubulin was done on HeLa cells. Cells were fixed with 70% ethanol for 10 minutes, permeabilized with 0.25% Tritonª X-100 for 20 minutes, and blocked with 5% BSA for 1 hour at room temperature. Cells were labeled with Alpha-tubulin Mouse Monoclonal Antibody (322500, red histogram) or with mouse isotype control (pink histogram) at 1-3 µg/million cells in 2.5% BSA. After incubation at room temperature for 2-3 hours, the cells were labeled with Alexa Fluor¨ 488 Rabbit Anti-Mouse Secondary Antibody (A11059) at a dilution of 1:400 for 30 minutes at room temperature. The representative 10,000 cells were acquired and analyzed for each sample using an Attune¨ Acoustic Focusing Cytometer. The purple histogram represents unstained control cells and the green histogram represents no-primary-antibody control.
Supportive validation
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- Figure 1. Molecular characterization of mouse intestinal tuft cells. Immunofluorescent stainings for (A) SOX9 and COX1, (B) SOX9 and COX2, (C) HPGDS and COX1, (D) villin and COX1, (E) alpha-tubulin and COX1, and (F) DCLK1 and COX1. Each panel contains a merged image on the left, and gray level pictures of the indicated individual markers corresponding to the yellow inset area on the right. (G) Whole-mount immunofluorescent staining for DCLK1 and F-actin on a dissociated fragment of intestinal epithelium. Panels on the right show higher magnification of the cropped area of the overlay image. Yellow arrowheads point at tuft cells. Nuclei are stained with Hoechst (blue). Bars, 10 um.
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
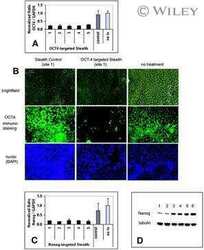
- Figure 6. Atoh1 is required for tuft cell differentiation. Immunofluorescent staining for the SOX9 transcription factor (A and C), the COX1 enzyme (A-D), and for the structural- and morphological-related tuft cells markers DCLK1 and alpha-tubulin (B and D) in intestines from control (A and B) and Atoh1 -deficient mice (C and D), 3 wk after tamoxifen injection. Each panel contains the merged image on the left, and separate pictures of the indicated markers corresponding to the yellow inset on the right. Yellow arrowheads point at tuft cells revealed by SOX9 and COX1 or DCLK1, alpha-tubulin, and COX1 expression. Nuclei are stained with Hoechst (blue). Bars, 10 um.
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- Figure 1 Effect of EGF on the claudins 1 and 3 expression in Caco-2 and HT-29 cells. Cultured Caco-2 and HT-29 cells were treated with EGF (20 ng/mL) for 6 h ( A ), 24 h and 48 h ( B ). Following EGF treatment, total cell lysates were harvested and analyzed by immunoblotting for the expression of claudins 1 and 3. alpha-tubulin was used as a loading control. The numbers represent the ratio of the optical density of EGF-treated to untreated cells normalized by alpha-tubulin. Claudin ; Cld. alpha -tubulin ; alpha-tub.
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- Figure 1. Generation of stable MDCK tet-off cell clones with inducible expression of PLEKHA7 constructs. ( A ) Schematic diagrams showing the putative minimal regions of PLEKHA7 which can interact with afadin (residues 120-374 in human PLEKHA7), p120ctn (538-696), CGNL1 (620-769), or nezha (680-821), 21 , 23 , 29 and schematic structure of the PLEKHA7 constructs (Full-length FL, N-terminal = N-term., C-terminal = C-term., Central) used to generate stable clones of MDCK tet-off cells. Constructs were tagged N-terminally with YFP, and C-terminally with myc. Putative structural domains are indicated: WW (purple), PH (brown), Pro-rich (P, red), and coiled-coil (CC, blue, CC1 = residues 686-835, and CC2 = residues 1055-1095, based on Phyre2 analysis). Numbers indicate position of PLEKHA7 amino acid residues within constructs. ( B ) Immunoblotting analysis of lysates of MDCK cell lines (two clones for each construct shown) cultured either in the presence (+) or in the absence (-) of doxycycline, using antibodies against the myc tag (to detect exogenous PLEKHA7), E-cadherin (E-cad.), cingulin (CGN) and tubulin (loading control). Numbers on the right indicate the migration of pre-stained markers (kDa). ( C ) Confocal immunofluorescent analysis of induced cultures of stable clones showing the localization of exogenous PLEKHA7 constructs (green, YFP) and alpha-tubulin (red), on the apical plane of focus (ZO-1 labeling is shown for control cells, panel E). Arrows = junctional labeling;
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- Fig 2 Deletion of YIH1 leads to the accumulation of cells in the G2/M phases of the cell cycle. (A) Representative images of cells grown asynchronously to log phase in YPD medium, spheroplasted with zymolyase, permeabilized, fixed and stained with DAPI to visualize the nuclear DNA (left panels) and with anti-tubulin antibodies followed by Alexa-488 anti-mouse IgG to visualize spindles (middle panels). (B) Cells prepared as in A were counted and the percentage of cells with medium sized-buds and short spindles and the localization of the nucleus as indicated (left), or with large buds and elongated spindles (right), were determined. (C) Representative images of live exponentially growing yeast cells stained with DAPI. (D) Quantification of the percentage of cells in G1 (unbudded), S (cells with small buds) and G2/M (cells with medium to large buds) determined from (C). Over 450 cells were analyzed per experiment. Data represent the mean +- S.E. as error bars of three independent experiments for B and D. * p < 0.05; (Student's t test).
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- Figure 1 Characterization of DOX-inducible MIA PaCa-2 cell lines A. qPCR analysis of HuR mRNA expression in Mia.CTRL, Mia.sh290, and Mia.sh700 cells treated with 0 or 2 mug/ml DOX for 5 days ( left ), and Mia.EV and Mia.HuR cells treated with 0 or 2 mug/ml DOX for 2 days ( right ), normalized to 18S rRNA expression. Mia.sh290 and Mia.sh700 are inducible HuR-silencing cell lines, whereas Mia.HuR is an inducible HuR-overexpressing cell line. Mia.CTRL and Mia.EV are the respective control cell lines. B. Western blotting analysis of HuR protein expression in Mia.CTRL, Mia.sh290, and Mia.sh700 cells treated with 0 or 2 mug/ml DOX for 5 days ( left ), and Mia.EV and Mia.HuR cells treated with 0 or 2 mug/ml DOX for 2 days ( right ), normalized to alpha-tubulin protein expression. ns = non-significant; *** = p < 0.001; **** = p < 0.0001.
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
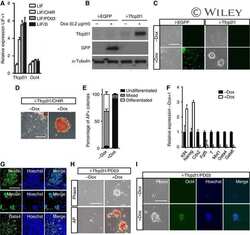
- Fig. 1. Overexpressing Sp5 partially mimics the effect of CHIR in medium with 2i. (A) Flow chart showing the method used to identify candidate genes downstream of CHIR and Stat3. (B) qRT-PCR analysis of Socs3 and Sp5 expression induced by CHIR, PD03 and 2i (left), or LIF (right) for 12 h. Data represent mean+-s.d. of three biological replicates. ** P
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- Fig. 3. Sp5 is a direct target of Stat3. (A) qRT-PCR analysis of Socs3 and Sp5 expression levels in 46C mESCs deprived of LIF overnight and stimulated with LIF for 1 h in the presence or absence of JAK inhibitor I (JAKi) or cycloheximide (CHX, 50 mg/ml). Data represent mean+-s.d. of three biological replicates. ** P
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- Fig. 4. Forced expression of Sp5 promotes EpiSC reprogramming to naive pluripotency. (A) Phase contrast and alkaline phosphatase (AP) staining images of 46C mESCs and 46C mESC-derived EpiSCs. Scale bar: 100 mum. (B) qRT-PCR analysis of expression patterns of Sp5 , Oct4 , Rex1 and Fgf5 in 46C mESCs and EpiSCs. Data represent mean+-s.d. of three biological replicates. ** P
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
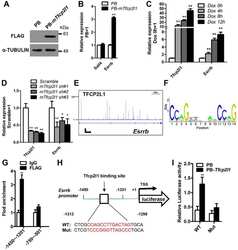
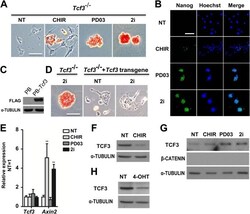
- Fig. 1. CHIR promotes mESC self-renewal via down-regulation of TCF3 protein in a beta-catenin-dependent manner. (A,B) Alkaline phosphatase (AP) staining and immunofluorescence images of Tcf3 -/- mESCs cultured in N2B27 supplemented with the indicated small molecules for two passages. Hoechst, Hoechst 33342. (C) Western blot analysis of the expression of FLAG in Tcf3 -/- mESCs overexpressing FLAG-tagged Tcf3 transgene. alpha-tubulin is a loading control. PB, PiggyBac. (D) AP staining of Tcf3 -/- mESCs and Tcf3 -/- mESCs overexpressing Tcf3 transgene. Cells were cultured in N2B27 medium with or without 2i for two passages. (E,F) qRT-PCR (E) and western blot (F) analysis of Tcf3 and Axin2 expression in 46C ESCs cultured under the indicated conditions for 12 h. (G,H) Western blot analysis of TCF3 expression in Ctnnb1 (beta-catenin) -/- ESCs (G) and Ctnnb1 -/- ESCs overexpressing Ctnnb1 -ERT2 (H). Cells were deprived of 2i/LIF overnight and then treated with the indicated compounds for 12 h in N2B27 medium. NT, no treatment; 4-OHT, 4-hydroxytamoxifen; ERT2, a mutant estrogen ligand-binding domain. Scale bars: 100 mum. Data represent mean+-s.d. of three biological replicates. ** P
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- Figure 2 FN and TNC expression in different endothelial cell models. ( a ) Western analysis (cropped blots) of FN (top) and TNC (bottom) expression in indicated cells. Culture medium (25 ul), left; total cell extracts of equal protein concentration (40 ug), right. Equivalent amounts of FN-depleted serum-containing (10%) culture medium were deposited as control. alpha-tubulin was used as loading control for total cell extracts. Full-length blots are shown in Fig. S5 . ( b ) Detection of cFN isoform transcripts (containing EDB+/EDA+, EDB+/EDA-, EDB-/EDA+, EDB-/EDA-, as schematically presented on the top) by RT-PCR in human endothelial cells and TIFs. 18S rRNA was used as internal control/housekeeping gene. Cropped blots are shown in ( a ) and ( b ). ( c ) QPCR analysis of FN, FN-EDB, FN-EDA and TNC mRNA expression in endothelial cells, normalized with GAPDH and presented as log DeltaCt (+-S.D., n = 5). ( d ) Double immunofluorescence staining (wide-field fluorescence) of total FN (red) and, in green, FN-EDA (left); FN-EDB (middle) or TNC (right) in confluent endothelial cells and TIFs. Cell density was evaluated after Hoechst 33342 staining (30-50 cells in each field). Bar = 100.
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- Figure 4 Effect of TNC on FN expression and assembly. ( a ) Representative western analysis of FN expression in total cell extracts of HUVECs cultured for 48 h on TNC-coated, BSA-pretreated or non-coated dishes. Equal amounts of protein (40 ug) were loaded in each lane and alpha-tubulin was used as loading control. Fold change over NC (normalized to alpha-tubulin) is shown below cropped blot. Full-length blots are shown in Fig. S7 . ( b ) QPCR analysis of FN mRNA expression levels in different endothelial cells (left, HUVEC; HMEC, middle and HMVEC-d, right) cultured on indicated substrates in comparison to a non-coated (NC) surface ( c ). Time course (12-72 h) expression of FN (left), Axin2 (middle) and DKK1 mRNA (right) in HUVECs on TNC coating, non-coated or BSA-pretreated dish was monitored by QPCR. Fold change over NC (equal to 1, not shown on the graphs) was calculated using the DeltaDeltaCt method (+-S.D., n = 3). ( d ) Immunofluorescence staining of cortactin (white or green), filamentous actin (red) and nuclei (blue) of cells plated for 48 h on non-coated or TNC-coated coverslips. The zoomed area is indicated in white dotted square (right). Bar = 50 um, and 10 um in zoomed area. ( e ) Representative western blot analysis of cofilin phosphorylation at Ser 3 in total cell extracts of HUVECs cultured for 48 h on TNC-coated or non-coated dishes. Equal amounts of protein (40 ug) were loaded in each lane and ERK1/2 was used as loading control. Fold change over NC (normalize
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- Figure 7 Induced HuR silencing suppresses PDA xenograft growth A. Representative images of subcutaneous Mia.sh290 tumors on the flanks of nude female mice, at the termination of the experiment (day 39). B. Tumor growth curves of Mia.sh290 xenografts. Mice were fed normal diet or 200 mg/kg DOX diet starting on day 0 (date of xenograft injection). Each data point represents the mean +- SEM ( n = 10 for -DOX group, and n = 8 for +DOX group). C. Plot of all tumor volumes on the final day of the Mia.sh290 xenograft experiment (day 39). Horizontal bars represent the median tumor volumes. D. Left , qPCR analysis of HuR mRNA expression in all Mia.sh290 xenografts harvested on day 39. 18S rRNA was used as normalization control. Data were plotted relative to the mean HuR mRNA expression in the -DOX group. Horizontal bars represent the medians. Right , plot of HuR mRNA expression vs. tumor volume for all Mia.sh290 tumors. Data were plotted relative to the mean of the -DOX group. E. Left , representative western blot of HuR protein expression in Mia.sh290 tumors harvested on day 39. Alpha-tubulin was used as normalization control. Right , quantification of the western blot. Each bar represents the mean +- SEM ( n = 3 per group). * = p < 0.05; ** = p < 0.01; **** = p < 0.0001.
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
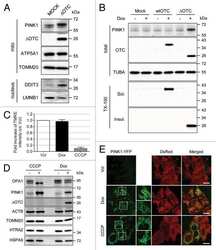
- Figure 3 MS-444 inhibits HuR cytoplasmic localization A. HCT116 cells treated with 10muM MS-444 for 6 hr were subjected to HuR immunofluorescence analysis (shown in green). DAPI (shown in blue) was used to visualize nuclei. B. HCT116 cells were treated with 10muM MS-444 for the indicated times. Cytoplasmic and whole cell lysates were probed for HuR, along with cytoplasmic alpha-tubulin and nuclear Lamin A/C markers. C. RIE-iRas cells were untreated or with 5 mM IPTG for 24 hr to induce oncogenic Ras expression, followed by 10 muM MS-444 for 8 hr. HuR was detected by immunofluorescence (green) along with DAPI was used to visualize nuclei (merged images shown). D. RIE-iRas were grown in the presence or absence of 5 mM IPTG for 24 hr and then treated with indicated amounts of MS-444 for 48 hr. Relative cell survival was performed by MTT assay and is represented as average of 4 independent experiments +- SEM. E. YAMC and YAMC-Ras cells were grown under non-permissive conditions at 37degC and treated with 10 muM MS-444 for 8 hours. HuR localization was assayed by immunofluorescence. F. MTT assay of YAMC and YAMC-Ras treated with MS-444 for 48 hr under non-permissive conditions. Bars = 10 mum.
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- Figure 4 Klotho and alpha smooth muscle actin (alpha-SMA) expression by Western blot: ( a ) alpha-SMA and Klotho expression in renal tissue, normalized by alpha-tubulin in the CTL, AD, Post AD-CTL, and Post AD-Fish oil groups. Relative quantification of alpha-SMA ( b , c ) and Klotho ( d , e ) in the CTL, AD, Post AD-CTL and Post AD-Fish oil groups (** p < 0.01).
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
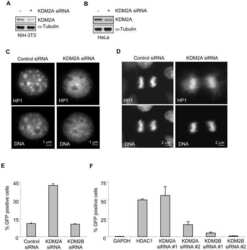
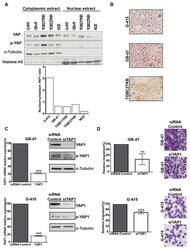
- Figure 3 Knockdown of YAP1 by siRNA affects the migration capacities in GB-d1 and G-415 gallbladder cancer cell lines. ( A ) Cytoplasmic and nuclear protein expression of YAP1 and phospho-YAP1 in GBC cell lines by Western blot and estimation of the YAP1 nuclear/cytoplasmic ratio. Normalized expression of cytoplasmic phospho-YAP1 and nuclear YAP1 was quantified by densitometry. ( B ) Immunocytochemistry of YAP1 in GBC cell lines (Magnification: 20x). GB-d1 and G-415 cancer cells were transfected with 25nM of siRNA against YAP1 or siRNA non-target control; ( C ) the knockdown of YAP1 was validated by assessing mRNA and protein levels in GB-d1 G-415 cell lines at 24 h. YAP1 was quantified by qRT-PCR using QARS and TFCP2 as internal controls, while alpha-tubulin was used as an internal control for loading protein. ( D ) Migration analysis of GB-d1 and G-415 gallbladder cancer cells treated for 24 h with siRNA non-target control and siRNA YAP1, n = 3. ** p < 0.01 and *** p < 0.001 by one-sample t- test. Magnification: 20x.
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- Fig. 2 Differential expression of protein phosphatases 1 gamma 2 (PP1gamma2) in whole cell (WC), nuclear (NC) and cytosolic (CYT) fractions of cervical cancer HeLa cell. A - western blot showing PP1gamma2 protein expression in the WC, NC and CYT fractions of HeLa cells, B - densitometry analysis of the western blot confirmed the differential expression of PP1gamma2. beta-actin and Histone-3 were used as a loading control for the cytoplasmic and nuclear fractions, respectively. These results are presented as the mean +- SEM. of 3 independent experiments, ** p < 0.01; as compared to WC, C - sub-cellular localization of PP1gamma2 in HeLa cells showed predominant expression of the PP1gamma2 ( r ) in the nuclear region of the cell, while discrete patches were also observed in the cytoplasm. Blue (b) represents nuclear stain DAPI and green (g) staining shows tubulin expression. Zoom represents 4x magnification of the original. All images were captured using an oil immersion 63x objective lens
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- Fig 4 The ATS1 domains of T . gondii myosin H are necessary for its conoid localization. (A) MyoH-3Ty solubility was evaluated by fractionation after extraction in PBS, PBS/NaCl, PBS/Na2CO3 or PBS/Triton X-100. Their distribution in different fractions was assessed by western blot using anti-Ty antibodies. The profilin (PRF) was used as soluble control. (B) Deoxycholate (DOC) extraction shows that MycMyoH (upper panels) and DD-MyoH-NT-DeltaATS1 (middle panels) remain bound to the conoid (arrowhead). Actin polymerization using jasplakinolide at 1 muM (JAS) showed MycMyoH strongly associated to the conoid (arrowhead) and not to the actin-containing apical extensions (lower panels). Scale bar 2 mum. (C) Schematic representation of the TgMyoH and FKBP destabilization domain (DD) constructs. MyoH contains eight IQ motifs and three ATS1/RCC1 domains. NT: Neck and Tail, T: Tail. (D) Western blot analysis using anti-Myc antibodies shows stabilization of DD-MyoH-NT (100 kDa), DD-MyoH-T (64 kDa), and DD-MyoH-NT-DeltaATS1 (76 kDa) after 24 h of Shield-1 (Shld-1) treatment. Catalase (CAT) serves as loading control. (E) Only DD-MyoH-NT is present at the conoid. (F) Plaque assays performed with DD-MyoH-NT, DD-MyoH-T, and DD-MyoH-NT-DeltaATS1 were fixed after 7 days. A strong impairment in the lytic cycle was observed for DD-MyoH-NT and DD-MyoH-NT-DeltaATS1 in the presence of Shld-1.
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
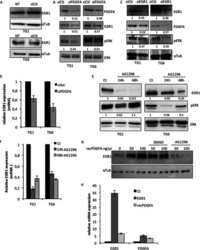
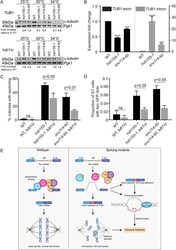
- FIGURE 5: Tubulin stability contributes to genome maintenance in splicing mutants. (A) Western blot of relative alpha-tubulin protein levels. Top, TUB1 ; bottom, intronless TUB1 . (B) Quantification of TUB1 mRNA transcript levels from exon region (left) and TUB1 intron region (right) by RT-qPCR. Asterisks show p values of DeltaDeltaCt: *** p = 0.0002; **** p < 0.0001. (C) CTF phenotypes in TUB1 or tub1 Delta i strains. (D) Endogenous Chr III stability in hsh155-1 and snu114-60 with intronless TUB1 (gray bars). (C, D) Fisher's exact test. (E) Model of defective splicing-induced genome instability in yeast. (B, C, D) Mean values with SEM are shown, n = 3 .
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
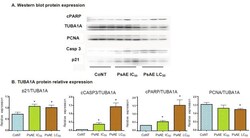
- Figure 6 Western blot analysis of cultured B16-F0 cells. A. After 24 h, protein expression of control, IC 50 and LC 50 PsAE treated cells (0, 2.5 and 9 mug/mL, respectively) are presented. B. Relative expression of p21, caspase 3, PCNA and cPARP with respect to TUBA1A are expressed in terms of mean +- SEM of three assays by triplicated. Results were compared by one-way ANOVA followed by Fisher LSD test. *: indicate significant differences (p
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- Figure 7 Western blot analysis of melanoma tumors. A. After 22 days of tumor induction, control and PsAE treated (150 mg/animal/day) groups are presented. B. Relative expression of p21 and PCNA over TUBA1A are expressed in terms of mean +- SEM. Results were compared by Student T test. *: indicates p = 0.0296. Full, non-adjusted images of p21 blot is provided in Supplementary Figure 5; TUBA1A (upper line) and PCNA (lower line) are provided in Supplementary Figure 6. Figure 7
 Explore
Explore Validate
Validate Learn
Learn Western blot
Western blot