Antibody data
- Antibody Data
- Antigen structure
- References [2]
- Comments [0]
- Validations
- Immunocytochemistry [2]
- Chromatin Immunoprecipitation [5]
- Other assay [2]
Submit
Validation data
Reference
Comment
Report error
- Product number
- 49-1033 - Provider product page

- Provider
- Invitrogen Antibodies
- Product name
- RNA pol II CTD Monoclonal Antibody
- Antibody type
- Monoclonal
- Antigen
- Other
- Reactivity
- Human, Mouse, Xenopus
- Host
- Mouse
- Isotype
- IgG
- Vial size
- 100 µL
- Concentration
- Conc. Not Determined
- Storage
- -20°C, Avoid Freeze/Thaw Cycles
Submitted references Regulatory mechanisms and function of hypoxia-induced long noncoding RNA NDRG1-OT1 in breast cancer cells.
The adenoviral E1A N-terminal domain represses MYC transcription in human cancer cells by targeting both p300 and TRRAP and inhibiting MYC promoter acetylation of H3K18 and H4K16.
Chao HH, Luo JL, Hsu MH, Chen LH, Lu TP, Tsai MH, Chuang EY, Chuang LL, Lai LC
Cell death & disease 2022 Sep 20;13(9):807
Cell death & disease 2022 Sep 20;13(9):807
The adenoviral E1A N-terminal domain represses MYC transcription in human cancer cells by targeting both p300 and TRRAP and inhibiting MYC promoter acetylation of H3K18 and H4K16.
Zhao LJ, Loewenstein PM, Green M
Genes & cancer 2016 Mar;7(3-4):98-109
Genes & cancer 2016 Mar;7(3-4):98-109
No comments: Submit comment
Supportive validation
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
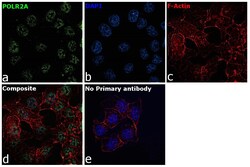
- Experimental details
- Immunofluorescence analysis of RNA pol II CTD Monoclonal Antibody was performed using 70% confluent log phase HCT 116 cells. The cells were fixed with 4% paraformaldehyde for 10 minutes, permeabilized with 0.1% Triton™ X-100 for 15 minutes, and blocked with 2% BSA for 45 minutes at room temperature. The cells were labeled with RNA pol II CTD Monoclonal Antibody (Product # 49-1033) at 1:100 dilution in 0.1% BSA, incubated at 4 degree celsius overnight and then labeled with Donkey anti-Mouse IgG (H+L) Highly Cross-Adsorbed Secondary Antibody, Alexa Fluor Plus 488 (Product # A32766), (1:2000 dilution), for 45 minutes at room temperature (Panel a: Green). Nuclei (Panel b:Blue) were stained with ProLong™ Diamond Antifade Mountant with DAPI (Product # P36962). F-actin (Panel c: Red) was stained with Rhodamine Phalloidin (Product # R415, 1:300 dilution). Panel d represents the merged image showing Nuclear localization. Panel e represents control cells with no primary antibody to assess background. The images were captured at 60X magnification.
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
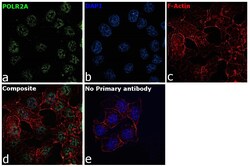
- Experimental details
- Immunofluorescence analysis of RNA pol II CTD Monoclonal Antibody was performed using 70% confluent log phase HCT 116 cells. The cells were fixed with 4% paraformaldehyde for 10 minutes, permeabilized with 0.1% Triton™ X-100 for 15 minutes, and blocked with 2% BSA for 45 minutes at room temperature. The cells were labeled with RNA pol II CTD Monoclonal Antibody (Product # 49-1033) at 1:100 dilution in 0.1% BSA, incubated at 4 degree celsius overnight and then labeled with Donkey anti-Mouse IgG (H+L) Highly Cross-Adsorbed Secondary Antibody, Alexa Fluor Plus 488 (Product # A32766), (1:2000 dilution), for 45 minutes at room temperature (Panel a: Green). Nuclei (Panel b:Blue) were stained with ProLong™ Diamond Antifade Mountant with DAPI (Product # P36962). F-actin (Panel c: Red) was stained with Rhodamine Phalloidin (Product # R415, 1:300 dilution). Panel d represents the merged image showing Nuclear localization. Panel e represents control cells with no primary antibody to assess background. The images were captured at 60X magnification.
Supportive validation
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
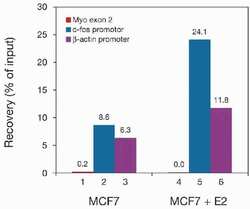
- Experimental details
- ChIP assays were performed using the breast carcinoma cell line MCF7 (treated with estradiol (MCF7 + E2) or not treated (MCF7), the anti–Pol II antibody (Product # 49-1033), and optimized PCR primer sets for myoglobin exon 2, c-fos promoter, β-actin promoter for qPCR. Each ChIP assay used sheared chromatin from 1 million cells and 2.8 µl of anti-Pol II ascites. Pol II binds to active promoters of protein-coding genes. Primer pairs for the following promoter regions were used: c-fos and β-actin. Myoglobin exon 2, which is not a promoter region, was used as negative PCR control. Recoveries (% of input) are shown here above. The percent recovery represents the relative amount of immunoprecipitated DNA compared to input DNA. The recoveries are high using the positive primer pairs c-fos promoter (bars 2 and 5) and β-actin promoter (bars 3 and 6), while there are no recoveries using the negative primer pair myoglobin exon 2 (bars 1 and 4). Moreover, recoveries are higher in cells treated with estradiol (MCF7 + E2) compared to recoveries in untreated MCF7 cells as expected.
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
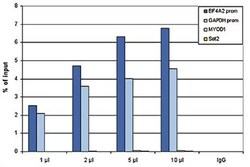
- Experimental details
- ChIP assays were performed using human HeLa cells, the anti-Pol II antibody (Product # 49-1033) and optimized PCR primer pairs for qPCR. ChIP was performed using sheared chromatin from 1 million cells. A titration consisting of 1, 2, 5 and 10 μL of antibody per ChIP experiment was analyzed. IgG (2 μg/ IP) was used as a negative IP control. Quantitative PCR was performed with primers specific for the promoter of the GAPDH and EIF4A2 genes, used as positive controls, and for the MYOD1 gene and the Sat2 satellite repeat, used as negative controls. The figure shows the recovery, expressed as a % of input (the relative amount of immunoprecipitated DNA compared to input DNA after qPCR analysis).
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
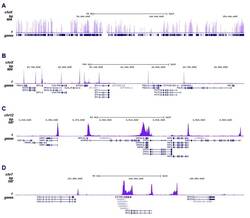
- Experimental details
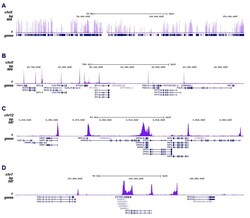
- ChIP was performed on sheared chromatin from 1 million HeLaS3 cells using 2 µL of the anti-Pol II antibody (Product # 49-1033). The IP’d DNA was subsequently analysed on an Illumina HiSeq. Library preparation, cluster generation and sequencing were performed according to the manufacturer’s instructions. The 51 bp tags were aligned to the human genome using the BWA algorithm. The figure shows the enrichment along the complete sequence and a 1 Mb region of the X-chromosome (fig A and B) and in genomic regions of chromosome 12 and 3, surrounding the GAPDH and EIF4A2 positive control genes (fig C and D).
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
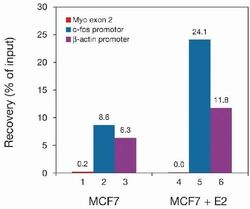
- Experimental details
- ChIP assays were performed using the breast carcinoma cell line MCF7 (treated with estradiol (MCF7 + E2) or not treated (MCF7), the anti–Pol II antibody (Product # 49-1033), and optimized PCR primer sets for myoglobin exon 2, c-fos promoter, β-actin promoter for qPCR. Each ChIP assay used sheared chromatin from 1 million cells and 2.8 µl of anti-Pol II ascites. Pol II binds to active promoters of protein-coding genes. Primer pairs for the following promoter regions were used: c-fos and β-actin. Myoglobin exon 2, which is not a promoter region, was used as negative PCR control. Recoveries (% of input) are shown here above. The percent recovery represents the relative amount of immunoprecipitated DNA compared to input DNA. The recoveries are high using the positive primer pairs c-fos promoter (bars 2 and 5) and β-actin promoter (bars 3 and 6), while there are no recoveries using the negative primer pair myoglobin exon 2 (bars 1 and 4). Moreover, recoveries are higher in cells treated with estradiol (MCF7 + E2) compared to recoveries in untreated MCF7 cells as expected.
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
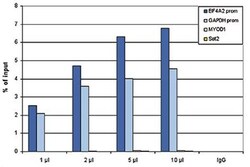
- Experimental details
- ChIP assays were performed using human HeLa cells, the anti-Pol II antibody (Product # 49-1033) and optimized PCR primer pairs for qPCR. ChIP was performed using sheared chromatin from 1 million cells. A titration consisting of 1, 2, 5 and 10 μL of antibody per ChIP experiment was analyzed. IgG (2 μg/ IP) was used as a negative IP control. Quantitative PCR was performed with primers specific for the promoter of the GAPDH and EIF4A2 genes, used as positive controls, and for the MYOD1 gene and the Sat2 satellite repeat, used as negative controls. The figure shows the recovery, expressed as a % of input (the relative amount of immunoprecipitated DNA compared to input DNA after qPCR analysis).
Supportive validation
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
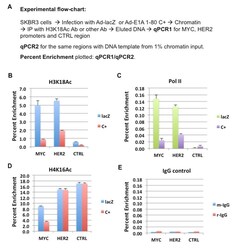
- Experimental details
- Figure 3 E1A 1-80 inhibition of RNA Pol II binding to and H3K18 acetylation on MYC and HER2 promoters in SKBR3 cells A. ChIP-qPCR experimental flow-chart. Percent enrichment of each promoter region is plotted. Percent enrichment is the ratio of qPCR1 from the ChIP DNA divided by qPCR2 from 1% input DNA. B. ChIP with H3K18Ac antibody. SKBR3 cells were infected with 20 PFU/cell of Ad-E1A 1-80 C+ or Ad-lacZ for 22 h, and fixed with formaldehyde before preparation of chromatin. CTRL: a control genomic region, ~ 4 kb upstream of HER2 transcription start site. MYC, HER2, and CTRL qPCR amounts were compared to the corresponding amounts from the ""input"" DNA. Each ChIP assay was repeated with the same chromatin preparation, and the averages of the two ChIP-qPCR assays are plotted, with error bars representing average deviations. C. , D. ChIP with RNA Pol II, and H4K16Ac antibodies, respectively. ChIP-qPCR was performed as in B. The different amounts of enrichment between the panels are likely due to different efficiencies of the antibodies used. E. ChIP control with non-specific antibodies, rabbit IgG (r-IgG) and mouse IgG (m-IgG).
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
- ChIP was performed on sheared chromatin from 1 million HeLaS3 cells using 2 µL of the anti-Pol II antibody (Product # 49-1033). The IP’d DNA was subsequently analysed on an Illumina HiSeq. Library preparation, cluster generation and sequencing were performed according to the manufacturer’s instructions. The 51 bp tags were aligned to the human genome using the BWA algorithm. The figure shows the enrichment along the complete sequence and a 1 Mb region of the X-chromosome (fig A and B) and in genomic regions of chromosome 12 and 3, surrounding the GAPDH and EIF4A2 positive control genes (fig C and D).
 Explore
Explore Validate
Validate Learn
Learn Western blot
Western blot Immunocytochemistry
Immunocytochemistry