Antibody data
- Antibody Data
- Antigen structure
- References [0]
- Comments [0]
- Validations
- Western blot [4]
- Immunocytochemistry [5]
- Immunohistochemistry [3]
- Other assay [46]
Submit
Validation data
Reference
Comment
Report error
- Product number
- MA5-11869 - Provider product page

- Provider
- Invitrogen Antibodies
- Product name
- Actin Monoclonal Antibody (ACTN05 (C4))
- Antibody type
- Monoclonal
- Antigen
- Other
- Description
- MA5-11869 reacts with all six known vertebrate isoforms of actin (MW ~42kDa), and with two cytoplasmic actins (beta and gamma), which are highly homologous to each other but differ from the muscle actins by ~25 amino acid residues. This product is highly recommended for monitoring total protein load on Western blots. Staining of formalin-fixed, paraffin-embedded tissues requires boiling tissue sections in 10mM citrate buffer, pH 6.0 for 10-20 minutes, followed by cooling at room temperature for 20 minutes. Suggested positive controls are HeLa cells and skeletal muscle. MA5-11869 is a suggested replacement to Product # 82353, clone # C4.
- Reactivity
- Human, Mouse, Rat, Bovine, Canine, Chicken/Avian, Porcine, Rabbit
- Host
- Mouse
- Isotype
- IgG
- Antibody clone number
- ACTN05 (C4)
- Vial size
- 500 µL
- Concentration
- 0.2 mg/mL
- Storage
- 4° C
No comments: Submit comment
Supportive validation
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- Western blot analysis of actin was performed by loading 25 µg of various whole cell lysates and 10 µL of PageRuler Plus Prestained Protein Ladder (Product # 26619) per well onto a 4-20% Tris-HCl polyacrylamide gel. Proteins were transferred to a PVDF membrane and blocked with 5% BSA in TBST for at least 1 hour. The membrane was probed with a pan actin monoclonal antibody (Product # MA5-11869) at a dilution of 1:3200 overnight at 4°C on a rocking platform, washed in TBS-0.1%Tween-20, and probed with HRP-conjugated goat anti-mouse IgG secondary antibody (Product # 31430) at a dilution of 1:40,000 for at least 1 hour. Chemiluminescent detection was performed using SuperSignal West Pico (Product # 34080).
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- Western blot of Actin pan using Actin pan Monoclonal Antibody (Product # MA5-11869) on HeLa Cells.
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- Western blot analysis of actin was performed by loading 25 µg of various whole cell lysates and 10 µL of PageRuler Plus Prestained Protein Ladder (Product # 26619) per well onto a 4-20% Tris-HCl polyacrylamide gel. Proteins were transferred to a PVDF membrane and blocked with 5% BSA in TBST for at least 1 hour. The membrane was probed with a pan actin monoclonal antibody (Product # MA5-11869) at a dilution of 1:3200 overnight at 4°C on a rocking platform, washed in TBS-0.1%Tween-20, and probed with HRP-conjugated goat anti-mouse IgG secondary antibody (Product # 31430) at a dilution of 1:40,000 for at least 1 hour. Chemiluminescent detection was performed using SuperSignal West Pico (Product # 34080).
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- Western blot analysis of Actin in differentiated THP-1 macrophage lysates. The cell lysates were subjected to 12% SDS-PAGE and proteins were then transferred to PDVF membrane. The membrane was incubated with Pan-Actin monoclonal antibody (Product # MA5-11869) at 1:1000 dilution with 5% skim milk in PBST (0.1% Tween) overnight at 4 degree and then 2 h at RT with HRP-conjugated Goat anti-mouse antibody. Signal was detected by incubation of membrane with HRP substrate (Product # 34579). Data courtesy of Antibody Data Exchange Program.
Supportive validation
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
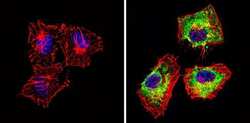
- Experimental details
- Immunofluorescent analysis of Actin pan (green) showing staining in the cytoplasm of Hela cells (right) compared to a negative control without primary antibody (left). Formalin-fixed cells were permeabilized with 0.1% Triton X-100 in TBS for 5-10 minutes and blocked with 3% BSA-PBS for 30 minutes at room temperature. Cells were probed with an Actin pan monoclonal antibody (Product # MA5-11869) in 3% BSA-PBS at a dilution of 1:100 and incubated overnight at 4 ºC in a humidified chamber. Cells were washed with PBST and incubated with a DyLight-conjugated secondary antibody in PBS at room temperature in the dark. F-actin (red) was stained with a fluorescent red phalloidin and nuclei (blue) were stained with Hoechst or DAPI. Images were taken at a magnification of 60x.
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
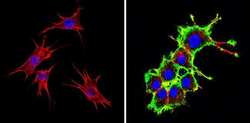
- Experimental details
- Immunofluorescent analysis of Actin pan (green) showing staining in the cytoplasm of NIH-3T3 cells (right) compared to a negative control without primary antibody (left). Formalin-fixed cells were permeabilized with 0.1% Triton X-100 in TBS for 5-10 minutes and blocked with 3% BSA-PBS for 30 minutes at room temperature. Cells were probed with an Actin pan monoclonal antibody (Product # MA5-11869) in 3% BSA-PBS at a dilution of 1:100 and incubated overnight at 4 ºC in a humidified chamber. Cells were washed with PBST and incubated with a DyLight-conjugated secondary antibody in PBS at room temperature in the dark. F-actin (red) was stained with a fluorescent red phalloidin and nuclei (blue) were stained with Hoechst or DAPI. Images were taken at a magnification of 60x.
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
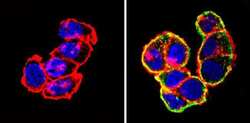
- Experimental details
- Immunofluorescent analysis of Actin pan (green) showing staining in the cytoplasm of T-47D cells (right) compared to a negative control without primary antibody (left). Formalin-fixed cells were permeabilized with 0.1% Triton X-100 in TBS for 5-10 minutes and blocked with 3% BSA-PBS for 30 minutes at room temperature. Cells were probed with an Actin pan monoclonal antibody (Product # MA5-11869) in 3% BSA-PBS at a dilution of 1:100 and incubated overnight at 4 ºC in a humidified chamber. Cells were washed with PBST and incubated with a DyLight-conjugated secondary antibody in PBS at room temperature in the dark. F-actin (red) was stained with a fluorescent red phalloidin and nuclei (blue) were stained with Hoechst or DAPI. Images were taken at a magnification of 60x.
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
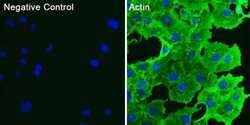
- Experimental details
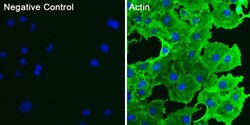
- Immunofluorescent analysis of actin (green) in COS7 cells. Cells were fixed and permeabilized with ice-cold methanol for 10 minutes at room temperature, and blocked with 0.3% BSA in PBS for 1 hour at room temperature. Cells were probed with a pan actin monoclonal antibody (Product # MA5-11869) at a dilution of 1:100 (right panel) or incubated in blocking buffer as a negative control (left panel) overnight at 4°C, washed with PBS, and incubated with DyLight 488 goat anti-mouse IgG secondary antibody (Product # 35502) at a dilution of 1:250 for 1 hour at room temperature. Nuclei (blue) were stained with DAPI (Product # 46190). Images were taken on a Thermo Scientific ToxInsight Instrument at 20X magnification.
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
- Immunofluorescent analysis of actin (green) in COS7 cells. Cells were fixed and permeabilized with ice-cold methanol for 10 minutes at room temperature, and blocked with 0.3% BSA in PBS for 1 hour at room temperature. Cells were probed with a pan actin monoclonal antibody (Product # MA5-11869) at a dilution of 1:100 (right panel) or incubated in blocking buffer as a negative control (left panel) overnight at 4°C, washed with PBS, and incubated with DyLight 488 goat anti-mouse IgG secondary antibody (Product # 35502) at a dilution of 1:250 for 1 hour at room temperature. Nuclei (blue) were stained with DAPI (Product # 46190). Images were taken on a Thermo Scientific ToxInsight Instrument at 20X magnification.
Supportive validation
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- Immunohistochemistry analysis of Actin pan showing positive staining in the cytoplasm of paraffin-treated Human prostate carcinoma (right) compared with a negative control in the absence of primary antibody (left). To expose target proteins, antigen retrieval method was performed using 10mM sodium citrate (pH 6.0), microwaved for 8-15 min. Following antigen retrieval, tissues were blocked in 3% H2O2-methanol for 15 min at room temperature, washed with ddH2O and PBS, and then probed with a Actin pan monoclonal antibody (Product # MA5-11869) diluted by 3% BSA-PBS at a dilution of 1:200 overnight at 4°C in a humidified chamber. Tissues were washed extensively PBST and detection was performed using an HRP-conjugated secondary antibody followed by colorimetric detection using a DAB kit. Tissues were counterstained with hematoxylin and dehydrated with ethanol and xylene to prep for mounting.
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- Immunohistochemistry analysis of Actin pan showing positive staining in the cytoplasm of paraffin-treated Human skeletal muscle (right) compared with a negative control in the absence of primary antibody (left). To expose target proteins, antigen retrieval method was performed using 10mM sodium citrate (pH 6.0), microwaved for 8-15 min. Following antigen retrieval, tissues were blocked in 3% H2O2-methanol for 15 min at room temperature, washed with ddH2O and PBS, and then probed with a Actin pan monoclonal antibody (Product # MA5-11869) diluted by 3% BSA-PBS at a dilution of 1:200 overnight at 4°C in a humidified chamber. Tissues were washed extensively PBST and detection was performed using an HRP-conjugated secondary antibody followed by colorimetric detection using a DAB kit. Tissues were counterstained with hematoxylin and dehydrated with ethanol and xylene to prep for mounting.
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- Immunohistochemistry analysis of Actin pan showing positive staining in the cytoplasm of paraffin-treated Mouse skeletal muscle (right) compared with a negative control in the absence of primary antibody (left). To expose target proteins, antigen retrieval method was performed using 10mM sodium citrate (pH 6.0), microwaved for 8-15 min. Following antigen retrieval, tissues were blocked in 3% H2O2-methanol for 15 min at room temperature, washed with ddH2O and PBS, and then probed with a Actin pan monoclonal antibody (Product # MA5-11869) diluted by 3% BSA-PBS at a dilution of 1:200 overnight at 4°C in a humidified chamber. Tissues were washed extensively PBST and detection was performed using an HRP-conjugated secondary antibody followed by colorimetric detection using a DAB kit. Tissues were counterstained with hematoxylin and dehydrated with ethanol and xylene to prep for mounting.
Supportive validation
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
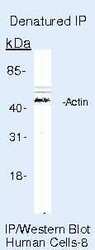
- Immunoprecipitation of Actin pan using Actin pan Monoclonal Antibody (Product # MA5-11869) on denatured Human HeLa Cells.
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
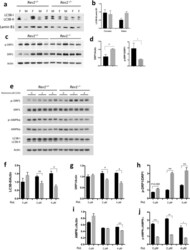
- Figure 4 Analysis of autophagy and mitochondrial dynamics in MEFs and hepatocytes in WT and Rev1 -/- cells. ( a) Analysis of autophagy by immunoblot and ( b) quantification of LC3B-II/Actin ratio in the liver of WT and Rev1 -/- 5 months old mice (females, n = 3; males n = 2). ( c) Immunoblot analysis of total and phosphorylated DRP1 (Ser616) in the liver of WT and Rev1 -/- 5 months old mice (male and females together, n = 5). ( d ) Quantification of total and phosphorylated DRP1 (Ser616) from panel ( c ). ( e ) Immunoblot with WT and Rev1 -/- MEF cells, treated with rotenone (0 muM, 2 uM and 4 uM) for 12 hrs (n = 3; df = 2). ( f) The LC3B-II/Actin ratio in untreated and treated MEF cells after 12 hrs (0 uM: t = 1, 2 uM: t = 2.2, 4 uM: t = 14). ( g , h) Analysis of total DRP1 (0 uM: t = 0.56; 2 uM: t = 5.1; 4 uM: t = 7.36) and phosphorylated DRP1 (Ser616) (0 uM: t = 3.96; 2 uM: t = 21.93; 4 uM: t = 14.14) in untreated and treated MEFs. ( i , j) Analysis of total AMPKalpha (0 uM: t = 2.09; 2 uM: t = 0.56; 4 uM: 81.52) and phospho- AMPKalpha (Thr172) (0 uM: t = 64.37; 2 uM: t = 21.2; 4 uM: t = 4.67) in untreated and treated MEFs. n = sample number; t = t value; df = degree of freedom; *p < 0.05. **p < 0.01. ***p < 0.001. ****p < 0.0001. Data presented are mean +- M.S.E. Dark bars: WT and gray bars: Rev1 -/- .
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
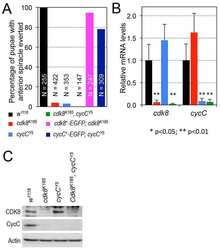
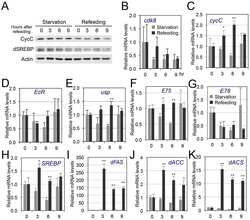
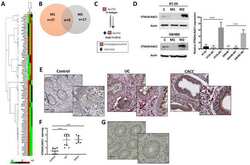
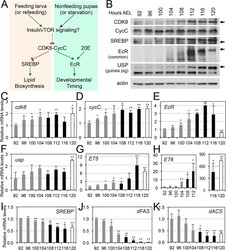
- Fig 7 CDK8-CycC may couple nutrient intake, lipid biosynthesis, and developmental timing. (A) Model for the CDK8-SREBP/EcR regulatory network: In response to nutrient intake, CDK8-CycC may coordinately regulate lipogenesis by directly inhibiting SREBP-activated gene expression and developmental timing by activating EcR-activated gene expression during the larval-pupal transition. Arrows represent activation, and blunt arrows represent inhibition. (B) The protein levels of CDK8, CycC, SREBP, EcR-B1 and USP (upper band is the 54 kDa full-length USP) in wild-type larvae from L3 (92 hr AEL) to the WPP stage (120 hr AEL). For SREBP, the lower band (approximately 49 kDa, arrow) is the mature nuclear form, while the upper band (53-54 kDa) is the N-terminal fragment of SREBP after cleavage by the S1P (see S5B and S5C Fig for detailed analyses of these SREBP isoforms). (C-K) The mRNA levels of cdk8 , cycC , EcR , usp , E75 , E78 , SREBP , dFAS , and dACS from L3 (92 hr AEL) to the WPP stage (120 hr AEL). The black bars represent the wandering stage, while the white bars represent the WPP stage. The x -axis represents the number of hours AEL. * p < 0.05; ** p < 0.01 based on t -tests. Underlying numerical data and statistical analysis for Fig 7C, 7D, 7E, 7F, 7G, 7H, 7I, 7J, and 7K can be found in S1 Data .
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
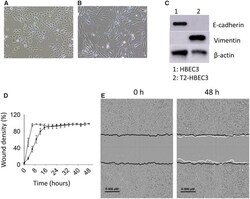
- Figure 3. Morphology, and migratory and invasive potential. Light microscopy images (x10 magnification) of HBEC3 (A) and T2-HBEC3 (B). C, E-cadherin and vimentin protein levels were analyzed in HBEC3 and T2-HBEC3 by immunoblotting. beta-actin was used as loading control. Migration and invasion were measured in a scratch wound closure assay. D, HBEC3 (*) and T2-HBEC3 (*) display significantly different migration curves. Data display wound density (percent) +- SE, n = 3. p < 0.05 (non-linear mixed effects model). E, Invasive potential of T2-HBEC3 analyzed by live cell imaging. Black lines display the original scratch wound made in the cell layer at 0 h and the white lines display the invading front of cells after 48 h.
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
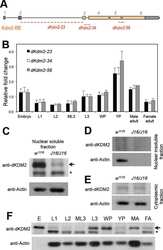
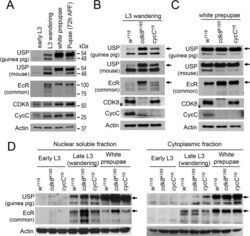
- Fig 4 The levels and subcellular distribution of EcR and USP in cdk8 and cycC mutants in the third instar larvae and pupae. (A) Western blot of EcR and USP in wild-type animals in early L3 (84 hr AEL), L3 wandering (112 hr AEL), white prepupal (120 hr AEL), and pupal stages (72 hr APF). (B and C) Western blot analyses of the protein levels of EcR, USP, CDK8, and CycC in cdk8 and cycC mutants at the L3 wandering stage (B) and the white prepupal stage (C). The arrows in USP blots mark the 54 kDa full-length USP, and the arrows in EcR blots indicate the EcR-B1 isoform. (D) Western blot of EcR and USP in the nuclear and cytoplasmic fractions from early third instar larvae (L3) to white prepupal stages in cdk8 or cycC mutants.
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
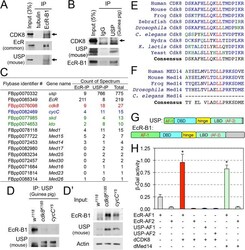
- Fig 6 Biochemical interactions between CDK8-CycC and EcR-USP. (A) EcR-B1 co-immunoprecipitates with CDK8 in white prepupae; USP serves as the positive control. (B) USP co-immunoprecipitates with CDK8 in white prepupae; EcR serves as the positive control. (C) Mediator subunits that can co-immunoprecipitate with EcR or USP are identified by LC-MS/MS analysis. Subunits of the CDK8 submodule are shown in color. The results were combined from two biological replicates. (D) Immunoprecipitation of EcR by using anti-USP (guinea pig) antibody in white prepupae of w 1118 , cdk8 K185 , and cycC Y5 mutants, and the input is shown in (D'). (E) CDK8 has a LXXLL motif that is highly conserved from yeast to human. The LXXLL motif is highlighted in red. (F) The LXXLL motif in Med14 is conserved from Drosophila to humans, but it is not present in Caenorhabditis elegans . (G) Schematic diagram of the EcR-B1 and USP protein depicting the two activating domains (AF1 and AF2), DNA-binding domain (DBD) and the ligand-binding domains (LBD). (H) Yeast two-hybrid analyses show that EcR-AF1, but not EcR-AF2 or USP-AF1/2, can directly bind to CDK8 and Med14. Underlying numerical data and statistical analysis for Fig 6H can be found in S1 Data .
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
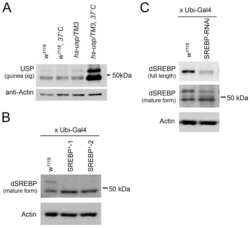
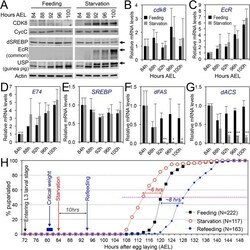
- Fig 8 The effects of starvation on the CDK8-SREBP/EcR regulatory network and timing for the larval-pupal transition. (A) The protein levels of CDK8, CycC, mature SREBP (arrow), EcR-B1 (arrow), and USP (arrow, full-length USP) in feeding versus starved larvae from 84 hr to 100 hr AEL. (B-G) The mRNA levels of cdk8 , EcR , E74 , SREBP , dFAS , and dACS in feeding versus starved larvae from 84 hr to 100 hr AEL. The x -axis represents the number of hours AEL. * p < 0.05; ** p < 0.01 based on t -tests. (H) The effect of starvation and refeeding on the timing of the larval-pupal transition in wild-type ( w 1118 ) larvae. The larvae enter into L3 at 72 hr AEL and reach critical weight between 80 and 82 hr AEL. The larvae were starved starting 84 hr AEL and the timing of pupariation was analyzed once every two hours. For the refeeding experiments, the larvae were put back on normal food after 10 hr of starvation (94 hr AEL). Underlying numerical data and statistical analysis for Fig 8B, 8C, 8D, 8E, 8F, 8G, and 8H can be found in S1 Data .
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
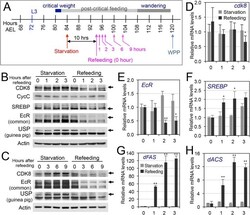
- Fig 9 The effects of refeeding of starved larvae on the CDK8-SREBP/EcR regulatory network. (A) The experimental scheme for the refeeding treatment. Briefly, the wild-type larvae were starved for 10 hr (from 84 hr AEL to 94 hr AEL), and they were then transferred back onto normal food. Samples were collected after 1, 2, 3, 6, 9 hr after refeeding for further analyses. (B) The protein levels of CDK8, CycC, SREBP, EcR-B1 and USP in starved versus refed larvae after 1, 2, 3 hr of refeeding. (C) The protein levels of CDK8, EcR-B1, and USP in starved versus refed larvae after 3, 6, 9 hr of refeeding. The control (anti-actin) is the same as the S12A Fig (D-H) The mRNA levels of cdk8 , EcR , SREBP , dFAS , and dACS in starved versus refed larvae after 1, 2, 3 hr of refeeding. The x -axis represents the number of hours for refeeding. * p < 0.05; ** p < 0.01 based on t -tests. Underlying numerical data and statistical analysis for Fig 9D, 9E, 9F, 9G, and 9H can be found in S1 Data .
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
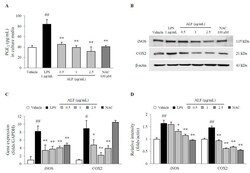
- Figure 3 Prostaglandin E 2 (PGE 2 ) production and inducible nitric oxide synthase (iNOS) and cyclooxygenase-2 (COX-2) expression in BV2 cells. Cell supernatant PGE 2 levels ( A ), protein expression of iNOS and COX-2 ( B ), and gene expression of iNOS and COX-2 relative to that of GAPDH ( C ) were determined using ELISA ( n = 6), Western blot ( n = 3), or real-time PCR ( n = 3) methods. Semiquantification of protein expression was performed with normalization to beta-actin ( D ). The data are expressed as the mean +- SD. # p < 0.05 and ## p < 0.01 compared with the vehicle-treated cells; * p < 0.05 and ** p < 0.01 compared with the LPS-treated BV2 cells.
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
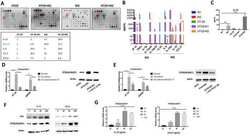
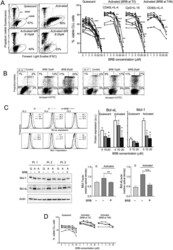
- Figure 1 BRB affects CLL cell viability. ( A ) Left: Flow cytometric dot plots of Propidium Iodide (PI) fluorescence versus Forward Light Scatter (FSC) for the determination of live (intact plasma membrane, PI negative) and dead (disrupted plasma membrane, PI positive) cells, from one CLL patient harboring 17p13 deletion. The cells were either quiescent or stimulated by CpG/ODN2006 + IL-15 and treated with BRB 10 and 20 muM for 48 h. Drug treatment of activated cells started simultaneously with stimulation. Note that the increase of FSC (i.e. cell size) after stimulation was inhibited by the presence of BRB. Right: Cell viability by the PI exclusion test is summarized for leukemic samples from 20 CLL patients, either untreated or treated with BRB at the indicated concentrations, for 48 h. Cell activation was achieved either with CD40L-NIH-3T3 + IL-4 or CpG/ODN2006 + IL-15, as indicated. BRB treatment started at stimulation time (T0), or 48 h after stimulation (T48h). Statistical significance of differences evaluated by a two- sided Wilcoxon signed rank test. ** P
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
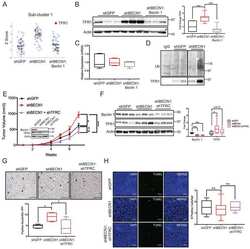
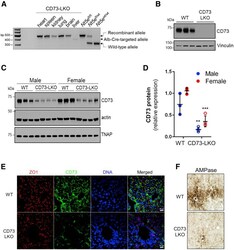
- Figure 2 Generation of liver-specific CD73-LKO mice. ( A ) PCR analysis of Nt5e generated a 349-bp fragment from the Cre-targeted allele in the CD73-LKO mouse liver. Nt5e (wt/wt) , Nt5e (fl/wt) , and Nt5e (fl/fl) mice are controls. Representative immunoblots of CD73 in ( B ) primary hepatocytes, and ( C ) in total liver lysates from male and female WT and CD73-LKO mice. Actin and tissue nonspecific alkaline phosphatase (TNAP) immunoblots serve as controls. ( D ) Semiquantitative analysis of CD73 protein expression based on immunoblot band intensities in panel C . ** P < .01, *** P < .001, 2-way analysis of variance. Error bars represent SD. ( E ) Immunofluorescence staining for CD73 (green), tight junction protein zonula occludens 1 (ZO1) (red), and 4'',6-diamidino-2-phenylindole (DAPI)-stained DNA (blue) on frozen liver tissue sections from WT ( top ) and CD73-LKO ( bottom ) mice. Scale bar : 20 mum. ( F ) AMPase activity in WT and CD73-LKO mice using formalin-fixed liver tissue sections. The brown deposits indicate ecto-AMPase activity in the presence of AMP. Stars indicate the central vein. Scale bar : 400 mum.
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
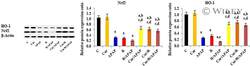
- 8 FIGURE Curcumin up-regulates the relative protein expression ratio of Nuclear factor erythroid-related factor-2 (Nrf2), and Hemeoxygenase-1 (HO-1) to beta-actin, determined by Western blot analysis in liver tissues of intoxicated rats. The results are demonstrated as mean +- SE, n = 10 rats/group. C, control untreated group; CUR, curcumin-pretreated rats; APAP, acetaminophen treated rats; R, gamma-irradiated rats; R/APAP, animals exposed to gamma-irradiation then intoxicated with APAP; CUR/APAP, Curcumin pretreated rats intoxicated with APAP; CUR/R, Curcumin pretreated rats then exposed to gamma-irradiation; CUR/R/APAP, Curcumin pretreated rats, exposed to gamma-irradiation and intoxicated with APAP; a: significant to the untreated control, b: significant to APAP, c: significant to R, and d: significant to R/APAP, ( p
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
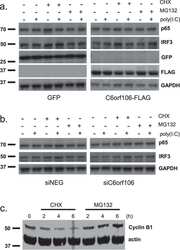
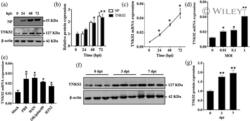
- 1 FIGURE Influenza virus induces TNKS2 expression in vitro and in vivo. (a, b) A549 cells were infected with A/PR/8/34 at an MOI of 0.01 and samples were collected at 0, 24, 48 and 72 hpi. Viral NP and TNKS2 protein levels were determined by western blot and normalised to beta-actin ( N = 6). The results for TNKS2 and NP were expressed as a ratio to 0 and 24 hpi, respectively. (c, d) A549 cells were infected with A/PR/8/34 at an MOI of 0.01 for 0, 24, 48 and 72 hr and different MOI for 24 hr. Relative mRNA expression levels of TNKS2 were measured by real-time PCR and normalised to beta-actin ( N = 3). (e) A549 cells infected with different strains of IAV, including A/PR/8/34, A/WSN/33, pdm/Ok/09, and H3N2 A/OK/309/06, at MOI 0.01 for 48 hpi. Relative mRNA expression levels of TNKS2 were measured by real-time PCR and normalised to beta-actin ( N = 3). (f, g) TNKS2 protein expression in the lungs of mice infected with A/PR/8/34 (250 Pfu) at 3 and 7 days post-infection (dpi) was measured by western blot and expressed as a ratio to 0 dpi ( N = 6). All results are displayed as the mean +- SE . * p < .05, ** p < .01 vs 0 hpi, MOI 0 or Mock. One-way ANOVA followed by dunnett ' s pairwise comparison
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
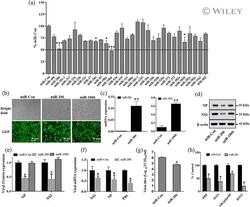
- 3 FIGURE miR-206 inhibits IAV replication. (a) A549 cells were co-transfected with pmirGLO-firefly-TNKS2-3'-UTR reporter plasmid and a miRNA expression plasmid or a miRNA control vector (miR-Con) for 24 hr and luciferase activities were measured. Relative firefly luciferase activities were normalised to Renilla luciferase activities and expressed as a percent of miR-Con. (b-g) HEK293 cells were transfected with a miRNA expression vector (miR-206, miR-106b) or a miRNA control vector (miR-Con) for 24 hr. The cells were then infected with A/PR/8/34 virus (MOI = 0.01) for 48 hr. (b) GPF images. (c) Real-time PCR analysis of miRNA expression normalised to U6. (d, e) Viral NP and NS1 protein expression was determined by western blot and normalised to beta-actin. (f) mRNA expression of viral genes (NP, NS1, and PB1) was determined by real-time PCR, normalised to beta-actin and expressed as a ratio to miR-Con. (g) Virus titers in the culture media were determined by TCID 50 assay. (h) HEK293 cells were transfected with a miR-206 or miR-Con plasmid (100 ng), the IAV firefly luciferase reporter plasmid vNP-luc/pHH21 (20 ng) and pRL-TK Renilla plasmid (5 ng) for 24 hr. The cells were then infected with different strains of IAV for 48 hr, including A/PR/8/34, A/WSN/33, pdm/Ok/09, and H3N2 A/OK/309/06, at MOI 0.01, 0.05, 0.005 and 0.01, respectively. Firefly luciferase activities were normalised to Renilla luciferase activity. The results of three independent experiments are displayed as
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
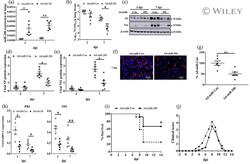
- 6 FIGURE Adenovirus expressing miR-206 suppresses IAV infection in vivo and increases survival of IAV-infected mice. (a-h) Two days before IAV infection (-2 dpi), adenovirus expressing miR-206 (Ad-miR-206) or miR-Con (Ad-miR-Con) was delivered intratracheally into the lungs of mice. The mice were then infected intranasally with a sub-lethal dose (250 PFU/mouse) of A/PR/8/34 on day 0. Lung tissue samples were collected at 3, 7 dpi. (a) miR-206 levels were measured using real-time PCR and normalised to U6 snRNA. (b) The virus titre was determined by TCID 50 assay. (c-e) Viral NP, NS1 protein levels were determined by western blot, normalised to beta-actin. (f, g) Viral replication at 7 dpi in the alveolar regions was determined by immunofluorescent staining for NP. For each section, five pictures were taken randomly from each region with x20 magnification. Total cells and NP positive cells were counted to determine the percentage of NP positive cells. (h) The mRNA levels of viral NS1 and PB1 mRNA were determined by real-time PCR and normalised to beta-actin. The results of six animals are displayed as the mean +- SE . * p < .05, ** p < .01, (a, b, d, e, h) Two-way ANOVA followed by sidak ' s multiple comparisons , (g) Student t test . (i, j) Two days after Ad-miR-206 or Ad-miR-con delivery, mice ( N = 12 per group, female) were infected with 1x MLD50 of A/PR/8/34 in 50 mul intranasally. (i) Kaplan-Meier survival analysis of mice. (j) Clinical scores was measured daily. The resu
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
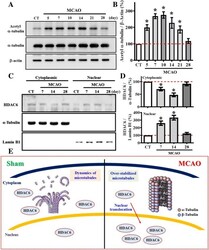
- Fig. 6 Aberrant nuclear translocation of HDAC6 after stroke causes hyper-acetylation of alpha-tubulin. ( a ) Representative western blot images of the acetylated-alpha-tubulin at Lys40, alpha-tubulin and beta-actin levels at different time points after MCAO surgery. ( b ) Quantification of the acetylated-alpha-tubulin that normalized to beta-actin after MCAO surgery. n = 4 biological replicates, * p < 0.05 compared to sham operated control. ( c ) Representative western blot images of the HDAC6 distributed between cytosolic or nuclear fractions at different time points after MCAO surgery. alpha-tubulin and Lamin B1 act as the internal control for either cytosolic or nuclear fraction. ( d ) Quantification of the cytoplasmic (top) or nuclear (bottom) protein levels of HDAC6 that normalized to its internal controls at different time points after MCAO. Data are presented as mean +- SEM. n = 5, * p < 0.05 compared to sham operated control. ( e ) A cartoon model illustrates the aberrant nuclear translocation of HDAC6 after stroke that causes hyper-acetylation alpha-tubulin
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
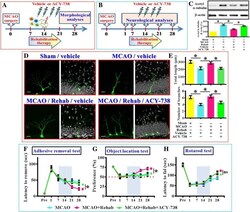
- Fig. 8 Requirement of HDAC6 in rehabilitation therapy induced functional recovery. ( a ) Schematic illustration the procedure for studying HDAC6 in rehabilitation therapy induced reversal of dendritic phenotypes. ( b ) Schematic illustration the procedure for studying HDAC6 in rehabilitation therapy induced long term functional recovery after MCAO. ( c ) Representative western blot images and quantification of the acetylated-alpha-tubulin that normalized to beta-actin from different experimental groups. n = 5 biological replicates, * p < 0.05. ( d ) Representative images of the newly generated neurons for their dendritic elaboration at 14 dpi. ""Rehab"" indicates the rehabilitation therapy for the mice. The scale bar is 25 mum. ( e ) Quantification of total dendritic length and the number of dendritic branches of individual newborn neurons at 14 dpi from different groups. Data are presented as mean +- SEM. n = 20 to 40 cells from four mice, one-way ANOVA with Bonferroni's post hoc analysis. * p < 0.05. ( f ) Behavioral performance of adhesive removal task. Two-way ANOVA analysis showed significant difference between the treatments ( p < 0.001) and Bonferroni's post hoc: p < 0.05 for the time points of 28 days for the MCAO+Rehab vs. MCAO+Rehab+ACY-738. ( g ) Object location memory test before or days after MCAO. Two-way ANOVA analysis showed significance between the treatments ( p < 0.01) and Bonferroni's post hoc: p < 0.05 for the time points of 28 days for the MCAO+Rehab vs.
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
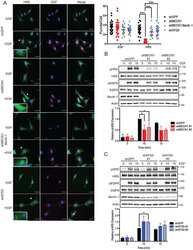
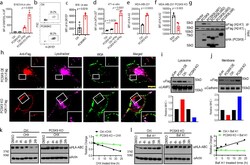
- Fig. 4. PCSK9 promotes lysosome-mediated degradation of MHC I in tumor cells. a. Quantitative estimates of SIINFEKL-H-2Kb levels in B16F10-OVA cells with or without IFNgamma treatment in control and PCSK9KO B16F10 cells. n=4,4,6,5 biologically independent samples, respectively. b-c . Flow cytometry estimates of H-2Kb/Db levels on the surface of subcutaneously grown control and PCSK9-deficient B16F10 cells. n=3 tumors per group. d . Quantitative estimate of MHC I levels on the surface of in tissue cultured control and PCSK9-deficient 4T1 cells treated with or without IFNgamma. n=5 biologically independent samples. e . The effect of PCSK9 deficiency on HLA-A2 expression on the surface of control and PCSK9KO MDA-MB-231 cells. n=3 biologically independent samples. f . The effect of exogenous PCSK9 protein on HLA-A2 degradation on the surface of MDA-MB-231 PCSK9KO cells. n=4 biologically independent samples. g . Interaction of mouse PCSK9 and H2-K1 in 293T cells were transduced with FLAG-tag labeled H2-K1 gene in combination with HA-tag labeled full-length or various deleted PCSK9 genes. h . PCSK9-promoted H2-K1 migration into the lysosome. Representative fluorescence confocal images of MHC I (H2-K1Flag) distribution in PCSK9 (PCSK9-HA) over-expressing (top panels) or PCSK9 KO (lower panels) B16 F10 cells. Scale bar: 10 mum. Insets showed magnified areas with additional details of co-localization. i-j. Western blot quantification of H2-K1-Flag in the lysosome (i) and plasma membra
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
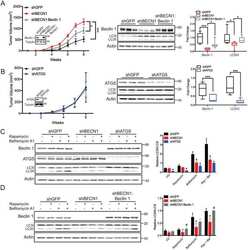
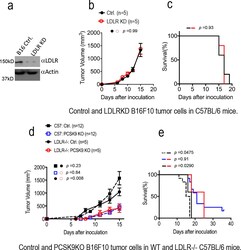
- Extended Data Fig. 3. The influence of tumor or host cell LDLR and host cholesterol levels on tumor growth from control or PCSK9KO tumor cells in immunocompetent hosts. a . Western blot analysis of CRISPR-Cas9 mediated knockdown (KD) of LDLR in B16F10 cells. The analysis was done once. b . Tumor growth from vector control and LDLR KD B16F10 cells in C57BL/6 mice. n=5 tumors per group. Error bars, mean +- S.E.M. P value calculated by two-way ANOVA. c . Kaplan Meier survival curve of mice (from b) bearing control and LDLR KD B16F10 tumors. n=5 mice per group. P value calculated by log-rank test. d . Tumor growth from vector control and PCSK9 knockout B16F10 cells in WT and LDLR-/- mice fed with high-fat diet. n=12, 12, 5, and 5 tumors in C57BL/6 mice inoculated with control and PCSK9KO tumor cells, and LDLR -/- mice inoculated with control and PCSK9 KO tumor cells. Error bars, mean +- S.E.M. P values calculated by two-way ANOVA with multiple comparisons. e. Kapan-Meier survival curve of wild type and LDLR-/- mice (from d) bearing vector control and PCSK9 knockout tumors. P value calculated by log-rank test.
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
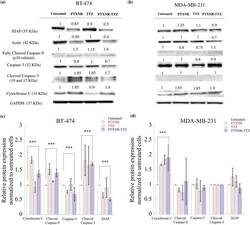
- Figure 7 Western blot analysis: The cell signaling protein expressions of ( a ) BT-474 and ( b ) MDA-MB-231 cells using PBS, PTX alone, TTZ alone, and PTXNR-TTX treatments. The relative intensity of XIAP, actin, cleaved caspase-9, caspase 3, cleaved caspase 3, and cytochrome C is analyzed after 48 h of treatment in ( c ) BT-474 and ( d ) MDA-MB-231 cells using ImageJ/ Fiji. For each protein expression the experiment was replicated at least n = 2-4 times and the average data is presented as mean +- standard deviation. The relative protein expressions are normalized to the untreated control cells and to the housekeeping GAPDH protein expression (hypothesized expression mean, u = 1). The p values for overexpression or downregulation of cytochrome-C, cleaved caspase-3, and XIAP in BT-474 are 0.039, 0.003, and 0.000049, respectively. The p value for the overexpression of cytochrome-C in MDA-MB-231 is 0.047. ***Represents p < 0.05 (for detailed p value calculations, please see SI Table 6 ).
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
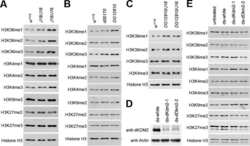
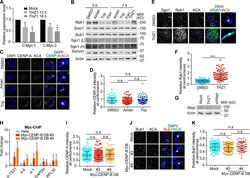
- Figure S2. Treament of transcriptional inhibitors does not change CENP-A levels at centromeres and expression of CENP-B DB does not alter Bub1 levels at centromeres. (A) RNA extracted from HeLa Tet-On cells with mock treatment or treated with THZ1 was subjected to real-time PCR analysis with the indicated primers. The average and standard error were calculated based on three independent experiments. (B) Lysates of HeLa Tet-On cells treated with DMSO, alpha-amanitin (Aman), triptolide (Trip), flavopiridol (Flav), or THZ1 were subjected to Western blots with the indicated antibodies. L, light exposure; H, heavy exposure. (C) Nocodazole-arrested HeLa Tet-On cells treated with DMSO, alpha-amanitin (Aman), or triptolide (Trip) were subjected to chromosome spread and stained with the indicated antibodies. The scale bars in the left and right panels represent 5 um and 1 um, respectively. (D) Quantification for relative CENP-A intensity (CENP-A/DNA) at centromeres in C. Quantification details are recorded in the Materials and methods. The experiment was repeated twice, and the results are highly reproducible. Quantification was performed based on the results from a single experiment. The average and standard deviation are shown here. At least 90 centromeres (six per cell) were scored for each condition. (E) Thymidine-arrested HeLa Tet-On cells were released into fresh medium containing DMSO or THZ1 and incubated for 12 h with the treatment with nocodazole in the last 2 h. Mitotic cel
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
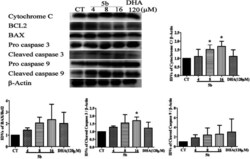
- Fig. 6 5b induces apoptosis by regulating caspase family proteins and Bcl2 family proteins in human GBM cells. Cells were treated with the indicated concentrations of 5b and DHA for 16 h. The proteins were collected for western blot assay and beta-actin was used as protein loading control. The result was obtained by three different independent experiments.
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
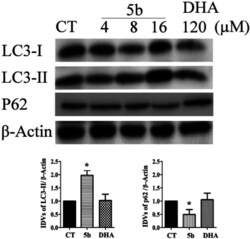
- Fig. 7 5b induces autophagy in human GBM cells. Cells were treated with the indicated concentrations of 5b and DHA for 16 h. The proteins were collected for western blot assay with LC3B, p62 and beta-actin antibodies. The result was obtained by three different independent experiments.
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
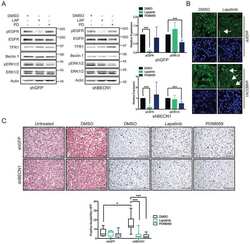
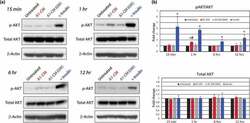
- Figure 4. Protein analyses of AKT signalling. (a) Representative Western blot images for protein phospholylation (p-) states and total AKT expression levels across time points including 15-min, 1-, 6- or 12-h post-treatment are shown for untreated, A1-CM, A1-CM (GW4869) and insulin-treated (1 uM) NRVMs. (b) Graphical representations of the quantitation for p-AKT (S473) to total AKT ratio (top graph) and total AKT levels (lower graph). Experiments were performed in triplicate, total AKT band intensities were calibrated to beta- actin as the loading control and expression values normalized to untreated controls. Statistical significance differences between untreated NRVMs and treatment groups are indicated by * ( p < 0.01), and # indicates statistically significant changes between A1-CM and A1-CM (GW) treatments ( p < 0.01).
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
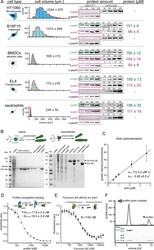
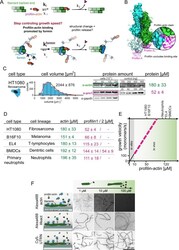
- Figure 1--figure supplement 1. Quantification of profilin-actin levels and purification of mammalian profilin-actin. ( A ) Profilin-actin concentration determination in mammalian cells. Left to right: name of cell type and origin, histograms of the single cell volume from fluorescence eXclusion measurements, quantitative western blot analysis of cellular actin, profilin1 and profilin-2 amount (left: titration of cell number, right: standard curve of recombinant proteins), calculated mean protein concentration [muM] per cell with experimental error (SD, N = 3 independent experiments, see Materials and methods). ( B ) Purification of mammalian cytoplasmic actin from two sources. Top: Schematic workflow of cytoplasmic actin purification. Left: native purification of beta,gamma-actin by gelsolin affinity chromatography. Right: Isolation of beta-actin from recombinant expression of beta-actin-linker-thymosin beta 4 -10xhis and purification via IMAC followed by chymotrypsin cleavage (^). For both strategies, finally released monomers were polymerized and separated from contaminants. Both purification protocols result in very pure and high yields of protein (see last pellet fraction). ( C ) Barbed end polymerization rate of cytoplasmic native mammalian (beta,gamma)-actin as a function of the actin monomer concentration. The mean values +- SD were fitted with a linear function. ( D ) Binding of profilin1 and 2 to ATP-bound cytoplasmic actin monomers measured by fluorescence anisotrop
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
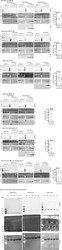
- Figure 1--figure supplement 2. Western blots and graphical summary of profilin-actin levels per cell. Determination of protein-levels in different cell types by quantitative western blot analysis. For each cell type (HT1080, B16F10, BMDC, neutrophils, HT1080/B16F10 overexpressing profilin1 and beta-actin) the amount of total profilin1/2 and actin was detected by monoclonal antibodies and calculated with in vitro protein controls in multiple experiments (see Materials and methods). The amount of profilin2 control is equal to the amount of profilin1 unless otherwise indicated. Right: Calculated profilin1/2 and actin concentrations per cell for individual experiments, errors indicate the SD. Last: Western blots from wt HT1080 cells and HT1080 overexpressing profilin1 and beta-actin from a polyclonal integrated construct (+one to +4) after sorting them by their mScarletI fluorescence intensity (see also Figure 6--figure supplement 2B ). For each sample, 250.000 cells were applied.
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
- Figure 1. Filament assembly at physiological profilin-actin concentrations. ( A ) Scheme of barbed end elongation from profilin-actin alone (top) or with formins (bottom). ( B ) Structural model of profilin at filament barbed ends (Materials and methods). The incoming profilin-actin complex is transparent. Actin is shown as green surface while profilin as magenta ribbons. Inset highlights the clash between the incoming actin monomer and profilin. ( C ) Profilin-actin measurements in HT1080 cells. Left to right: single cell volume histogram, western blots of actin, profilin1 (left: cell titration, right: standard curve of recombinant proteins), values are mean (N = 3) and SD, Materials and methods. ( D ) Table of total concentrations of actin and profilin-1/2 in various mammalian cell types ( Figure 1--figure supplement 1 ). ( E ) Scheme of a linearly substrate-dependent actin elongation rate. Top axis: Profilin-actin amounts for various cell types as indicated. ( F ) Scheme (left) and TIRFM images (right) of elongating filaments at indicated profilin-actin concentrations visualized with top- Alexa488-labeled monomers (20% labeled), middle - 10 nM Alexa488-lifeact, bottom - 10 nM Cy5-UTRN 261 . Figure 1--source data 1. Data Figure 1 . Figure 1--figure supplement 1. Quantification of profilin-actin levels and purification of mammalian profilin-actin. ( A ) Profilin-actin concentration determination in mammalian cells. Left to right: name of cell type and origin, histograms of t
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
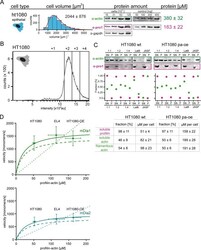
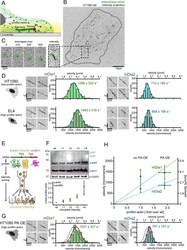
- Figure 6--figure supplement 2. Profilin1-actin overexpression and in vivo formin speeds at different profilin1-actin levels. ( A ) Profilin-actin concentration determination in mammalian HT1080 cells overexpressing profilin1 and beta-actin. Left to right: name of cell type and origin, histograms of the single cell volume from fluorescence eXclusion measurements, quantitative western blot analysis of cellular actin, profilin1 amount (left: titration of cell number, right: standard curve of recombinant proteins), calculated mean protein concentration [muM] per cell with experimental error (SD, N = 3 independent experiments, see Materials and methods). ( B ) Histogram from FACS analysis showing the ScarletI intensity distribution from a polyclonal population of HT1080 cells overexpressing profilin1, mScarletI and beta-actin. This population was sorted into subpopulations as indicated (+one to +4). ( C ) Mean velocities of either mDia1 (top) and mDia2 (bottom) in different mammalian cell types (wt HT080, EL4 and HT1080 overexpressing profilin-actin) plotted as a function of the quantified profilin-actin concentration. Error indicators are SD. Dashed lines are the formin speeds determined in vitro (see Figure 5C ), dotted lines are linear fits through the origin to the in vivo data. Continuous lines fit to the in vivo data by a hyperbolic model with only one free parameter (vmax, the maximal growth velocity). K 0.5 (the profilin-actin concentration at half-maximal elongation speed
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
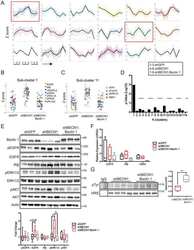
- Figure 6. Formin single molecule imaging reveals buffered elongation rates in mammalian cells. ( A ) Scheme of TIRFM imaging of single formins in the actin cortex of cells. ( B ) Maximum intensity projection of a TIRFM time-lapse shows growth trajectories of single mNeonGreen-mDia1 molecules in the cortex of a HT1080 cell. Inset: Close-up of a single trajectory as in ( C ). ( C ) TIRFM time-lapse images (left) and intensity projection (right) of an individual mNeonGreen-mDia1 molecule. ( D ) Measurements of mDia1/2 elongation velocities in vivo. Left to right: Scheme of HT1080 (top) and EL4 (lower) cells, kymographs of single mNeonGreen- mDia1 (left) or mDia2 (right) molecules followed by velocity distributions. Lines are Gaussian fits. Means and SD are indicated. [N cells >= 10, n molecules/cell >=30, n total >=650 per condition]. ( E ) Workflow to generate profilin1 and beta-actin overexpressing HT1080 cells. Polycistronic constructs for beta-actin, mScarletI and profilin1 were integrated into the genome. Cells were sorted into four sub-populations dependent on mScarletI fluorescence intensity ( Figure 6--figure supplement 2B , Materials and methods). ( F ) Top: Western blot of HT1080 cells (wt or overexpressing sub-populations). No translational read-through is visible (1xRP: actin-mScarletI, 2xRP: actin-mScarletI-profilin1 at expected Mw). Bottom: Relative profilin1 and actin levels (fold over wt) for indicated sub-populations. ( G ) mDia1/2 velocities in profilin-actin o
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
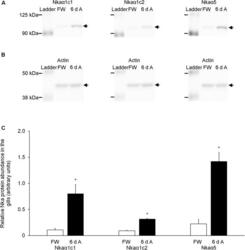
- FIGURE 8 Effects of exposure to 30 mmol l -1 NH 4 Cl on the protein abundance of Na + /K + -ATPase alpha-subunit (Nkaalpha) isoforms in the gills of Heteropneustes fossilis. Protein abundances of Nkaalpha1c1, Nkaalpha1c2, and total Nkaalpha (as reflected by immunostaining with the alpha5 comprehensive anti-Nkaalpha antibody) in the gills of H. fossilis kept in fresh water (control; FW) or exposed to 30 mmol l -1 NH 4 Cl for 6 days (6 d A). (A) An example of the immunoblot of Nkaalpha1c1, Nkaalpha1c2, and total Nkaalpha. (B) An example of the immunoblot of actin. (C) The protein abundance of the various Nka bands normalized with respect to that of actin. Results represent mean + SEM ( N = 4). *Significance difference between the NH 4 Cl-exposed group (6 d A; closed bar) and the corresponding freshwater group (FW; open bar) ( P < 0.05).
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
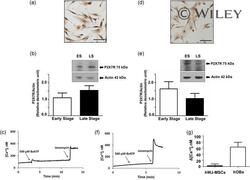
- 2 Figure Expression and function of P2X7R in hOBs and hWJ-MSCs. (a) Representative immunostaining of P2X7R in freshly isolated hOBs. (b) Representative western blot in the early stage (ES) and in the late stage (LS) of hOBs differentiation is reported with densitometric analysis of all samples analyzed. Data are expressed as mean +- standard deviation ( SD ) of P2X7R/Actin ratio. Data were analyzed by Student's t test, p < .05. (c) Representative trace showing the intracellular Ca 2+ rise following hOBs stimulation with 500 uM BzATP. (d) Representative immunostaining of P2X7R in hWJ-MSCs. (e) Representative western blot in the early stage (ES) and in the LS of differentiation of hWJ-MSCs is reported with densitometric analysis of all samples analyzed. Data are expressed as mean +- SD of P2X7R/Actin ratio. (f) Representative trace showing the intracellular Ca 2+ rise following hWJ-MSCs stimulation with 500 uM BzATP. (g) Intracellular Ca 2+ rise following stimulation with 500 uM BzATP. All experiments were carried out at least three times. hOBs, human primary osteoblast; hWJ-MSC, Wharton's jelly mesenchymal stromal cells
 Explore
Explore Validate
Validate Learn
Learn Western blot
Western blot Immunoprecipitation
Immunoprecipitation