Antibody data
- Antibody Data
- Antigen structure
- References [6]
- Comments [0]
- Validations
- Flow cytometry [1]
- Other assay [4]
Submit
Validation data
Reference
Comment
Report error
- Product number
- 25-0577-42 - Provider product page

- Provider
- Invitrogen Antibodies
- Product name
- CD57 Monoclonal Antibody (TB01 (TBO1)), PE-Cyanine7, eBioscience™
- Antibody type
- Monoclonal
- Antigen
- Other
- Description
- Description: This TB01 monoclonal antibody reacts with human CD57 (also known as HNK-1 and Leu-7), a 110-kDa cell surface glycoprotein expressed on a subset of natural killer (NK) cells and NK T cells. Applications Reported: This TB01 (TBO1) antibody has been reported for use in flow cytometric analysis. Applications Tested: This TB01 (TBO1) antibody has been pre-titrated and tested by flow cytometric analysis of normal human peripheral blood cells. This can be used at 5 µL (1.0 µg) per test. A test is defined as the amount (µg) of antibody that will stain a cell sample in a final volume of 100 µL. Cell number should be determined empirically but can range from 10^5 to 10^8 cells/test. Light sensitivity: This tandem dye is sensitive to photo-induced oxidation. Please protect this vial and stained samples from light. Fixation: Samples can be stored in IC Fixation Buffer (Product # 00-822-49) (100 µL of cell sample + 100 µL of IC Fixation Buffer) or 1-step Fix/Lyse Solution (Product # 00-5333-54) for up to 3 days in the dark at 4°C with minimal impact on brightness and FRET efficiency/compensation. Some generalizations regarding fluorophore performance after fixation can be made, but clone specific performance should be determined empirically. Excitation: 488-561 nm; Emission: 775 nm; Laser: Blue Laser, Green Laser, Yellow-Green Laser. Filtration: 0.2 µm post-manufacturing filtered.
- Reactivity
- Human
- Host
- Mouse
- Isotype
- IgM
- Antibody clone number
- TB01 (TBO1)
- Vial size
- 100 Tests
- Concentration
- 5 µL/Test
- Storage
- 4° C, store in dark, DO NOT FREEZE!
Submitted references Aberrant newborn T cell and microbiota developmental trajectories predict respiratory compromise during infancy.
Characterization of HIV-induced remodeling reveals differences in infection susceptibility of memory CD4(+) T cell subsets in vivo.
Phenotypic analysis of the unstimulated in vivo HIV CD4 T cell reservoir.
Function of multiple sclerosis-protective HLA class I alleles revealed by genome-wide protein-quantitative trait loci mapping of interferon signalling.
Mass Cytometric Analysis of HIV Entry, Replication, and Remodeling in Tissue CD4+ T Cells.
A novel method for autophagy detection in primary cells: impaired levels of macroautophagy in immunosenescent T cells.
McDavid A, Laniewski N, Grier A, Gill AL, Kessler HA, Huyck H, Carbonell E, Holden-Wiltse J, Bandyopadhyay S, Carnahan J, Dylag AM, Topham DJ, Falsey AR, Caserta MT, Pryhuber GS, Gill SR, Scheible KM
iScience 2022 Apr 15;25(4):104007
iScience 2022 Apr 15;25(4):104007
Characterization of HIV-induced remodeling reveals differences in infection susceptibility of memory CD4(+) T cell subsets in vivo.
Xie G, Luo X, Ma T, Frouard J, Neidleman J, Hoh R, Deeks SG, Greene WC, Roan NR
Cell reports 2021 Apr 27;35(4):109038
Cell reports 2021 Apr 27;35(4):109038
Phenotypic analysis of the unstimulated in vivo HIV CD4 T cell reservoir.
Neidleman J, Luo X, Frouard J, Xie G, Hsiao F, Ma T, Morcilla V, Lee A, Telwatte S, Thomas R, Tamaki W, Wheeler B, Hoh R, Somsouk M, Vohra P, Milush J, James KS, Archin NM, Hunt PW, Deeks SG, Yukl SA, Palmer S, Greene WC, Roan NR
eLife 2020 Sep 29;9
eLife 2020 Sep 29;9
Function of multiple sclerosis-protective HLA class I alleles revealed by genome-wide protein-quantitative trait loci mapping of interferon signalling.
Lundtoft C, Pucholt P, Imgenberg-Kreuz J, Carlsson-Almlöf J, Eloranta ML, Syvänen AC, Nordmark G, Sandling JK, Kockum I, Olsson T, Rönnblom L, Hagberg N
PLoS genetics 2020 Oct;16(10):e1009199
PLoS genetics 2020 Oct;16(10):e1009199
Mass Cytometric Analysis of HIV Entry, Replication, and Remodeling in Tissue CD4+ T Cells.
Cavrois M, Banerjee T, Mukherjee G, Raman N, Hussien R, Rodriguez BA, Vasquez J, Spitzer MH, Lazarus NH, Jones JJ, Ochsenbauer C, McCune JM, Butcher EC, Arvin AM, Sen N, Greene WC, Roan NR
Cell reports 2017 Jul 25;20(4):984-998
Cell reports 2017 Jul 25;20(4):984-998
A novel method for autophagy detection in primary cells: impaired levels of macroautophagy in immunosenescent T cells.
Phadwal K, Alegre-Abarrategui J, Watson AS, Pike L, Anbalagan S, Hammond EM, Wade-Martins R, McMichael A, Klenerman P, Simon AK
Autophagy 2012 Apr;8(4):677-89
Autophagy 2012 Apr;8(4):677-89
No comments: Submit comment
Supportive validation
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
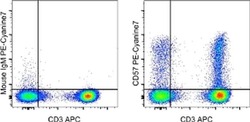
- Experimental details
- Staining of normal human peripheral blood cells with Anti-Human CD3 APC (Product # 17-0036-42) and 1.0 µg of Mouse IgM Isotype Control APC (Product # 17-4752-80) (left) or 1.0 µg of Anti-Human CD57 PE-Cyanine7 (right). Cells in the lymphocyte gate were used for analysis.
Supportive validation
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- Figure 5. Design of tailored sort strategies to isolate three populations memory CD4+ PD1+ T cells harboring different frequencies of kNN latent cells. Sort strategies were designed for each of the four leukapheresis donors analyzed by PP-SLIDE. Shown are the CyTOF datasets, the unstimulated atlas cells are shown as gray contours and kNN latent cells shown as red contours. Gates in black correspond to upstream gates. Each sorting strategy isolates three populations of memory CD4+ PD1+ T cells: two disenriched ( pink , purple ), and one enriched ( red ). Results were pre-gated on live, singlet memory CD4+ T cells (CD3+CD8-CD19-CD45RO+CD45RA-). The three functional assays applied to sorted cells are listed. The gray inset on the right shows frequencies of kNN latent cells in the final enriched populations (per million memory CD4+ T cells) and the fold-enrichment of kNN latent cells in the final sorted enriched population relative to each of the disenriched populations. In instances where a disenriched population did not harbor any kNN cells, the fold-enrichment is listed as NA (not available) because the fold-enrichment is infinity when divided by zero.
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
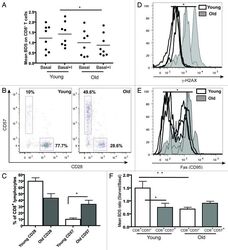
- Figure 5. Aging and replicative senescence markers in human T lymphocytes. PBMCs from healthy young (< 28 y) and old (> 56 y) donors were stained for CD28 and CD57 and run on LSR II flow cytometer. (A) Autophagy levels (Mean BDS) in CD8 + T cells from young (< 28 y, n = 8) and old (> 56 y, n = 8) donors under basal and basal+I (for 2 h) conditions (mean +- SEM, *p < 0.0499 between young and old basal+I). (B) Representative dot plots from a young and an old donor showing percentages of CD28 and CD57 cells gated on CD8 + T cells. (C) Bar graph showing % of CD8 + lymphocytes with CD28 and CD57 markers in four young and old donors (mean +- SEM, p = 0.0571 for CD8 CD28 population and *p = 0.0286 for CD8 CD57 population). (D) Overlaid histogram of gammaH2AX (DNA double-strand break) levels of CD8 + lymphocytes from three young and old donors gated on CD28 + CD57 - population (geometric mean +- SEM, *p = 0.0286). (E) Overlaid histogram of FAS (CD95) levels of CD8 + lymphocytes from four young and old donors gated on CD28 + CD57 - population (geometric mean +- SEM, *p = 0.0286). PBMCs from four healthy young and old donors were cultured under control and starved conditions for 2 h and stained for CD8, CD57, LC3 and Lyso-ID. (F) Colocalization of LC3 and lysosomal marker in CD8 + CD57 +/- cells, expressed as mean BDS ratio between starved and basal treatments (mean +- SEM, n = 5 (young donors), n = 8 (old donors), **p = 0.0049, *p = 0.035).
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- Figure 6. Levels of gammaH2AX foci in senescent T cells with low autophagy. PBMCs from three healthy donors were bead sorted for CD8 + T cells and stained for CD57 gammaH2AX, LC3 and Lyso-ID and run on ImageStream. (A) Representative BDS overlay histogram for CD8 + CD57 + (red) for CD8 + CD57 - (green) depicts the low and high autophagy gates. (B) Bar graph showing proportion of CD8 + CD57 +/- lymphocytes with low and high autophagy in three healthy donors (mean +- SEM, **p = 0.0029, *p = 0.0273). (C) Bar graph showing mean gammaH2AX foci/cell in low and high autophagy CD8 + CD57 + , CD8 + CD57 - populations (geometric mean +- SEM, **p = 0.0057 and **p = 0.0087, respectively). (D) Representative ImageStream images of gammaH2AX foci in CD8 + lymphocytes from young and old cells.
 Explore
Explore Validate
Validate Learn
Learn Flow cytometry
Flow cytometry