Antibody data
- Antibody Data
- Antigen structure
- References [41]
- Comments [0]
- Validations
- Western blot [2]
- Immunohistochemistry [1]
- Other assay [10]
Submit
Validation data
Reference
Comment
Report error
- Product number
- MA1-26418 - Provider product page

- Provider
- Invitrogen Antibodies
- Product name
- RPA2 Monoclonal Antibody (9H8)
- Antibody type
- Monoclonal
- Antigen
- Other
- Description
- Recommended positive controls: Tonsil.
- Reactivity
- Human
- Host
- Mouse
- Isotype
- IgG
- Antibody clone number
- 9H8
- Vial size
- 500 µL
- Storage
- 4° C
Submitted references NSMF promotes the replication stress-induced DNA damage response for genome maintenance.
CARM1 regulates replication fork speed and stress response by stimulating PARP1.
DDX11 loss causes replication stress and pharmacologically exploitable DNA repair defects.
APE1 senses DNA single-strand breaks for repair and signaling.
A phosphorylation-and-ubiquitylation circuitry driving ATR activation and homologous recombination.
Restoration of Replication Fork Stability in BRCA1- and BRCA2-Deficient Cells by Inactivation of SNF2-Family Fork Remodelers.
ATR kinase inhibition induces unscheduled origin firing through a Cdc7-dependent association between GINS and And-1.
Functions of Replication Protein A as a Sensor of R Loops and a Regulator of RNaseH1.
PC4 promotes genome stability and DNA repair through binding of ssDNA at DNA damage sites.
DNA end resection is needed for the repair of complex lesions in G1-phase human cells.
Gap-filling and bypass at the replication fork are both active mechanisms for tolerance of low-dose ultraviolet-induced DNA damage in the human genome.
Scaffold functions of 14-3-3 adaptors in B cell immunoglobulin class switch DNA recombination.
BLM SUMOylation regulates ssDNA accumulation at stalled replication forks.
Mammalian Exo1 encodes both structural and catalytic functions that play distinct roles in essential biological processes.
Zinc finger protein 668 interacts with Tip60 to promote H2AX acetylation after DNA damage.
Replication stress links structural and numerical cancer chromosomal instability.
Differential contribution of HP1 proteins to DNA end resection and homology-directed repair.
Cyclin-dependent kinase suppression by WEE1 kinase protects the genome through control of replication initiation and nucleotide consumption.
REV1 and polymerase ζ facilitate homologous recombination repair.
ATM modulates the loading of recombination proteins onto a chromosomal translocation breakpoint hotspot.
A PP4 phosphatase complex dephosphorylates RPA2 to facilitate DNA repair via homologous recombination.
Sensitization of ovarian carcinoma cells to the atypical retinoid ST1926 by the histone deacetylase inhibitor, RC307: enhanced DNA damage response.
ATM- and ATR-mediated response to DNA damage induced by a novel camptothecin, ST1968.
Deficiency of FANCD2-associated nuclease KIAA1018/FAN1 sensitizes cells to interstrand crosslinking agents.
Human Fbh1 helicase contributes to genome maintenance via pro- and anti-recombinase activities.
The cohesin complex is required for the DNA damage-induced G2/M checkpoint in mammalian cells.
Ionizing radiation-dependent and independent phosphorylation of the 32-kDa subunit of replication protein A during mitosis.
Gamma-glutamyltransferase-dependent resistance to arsenic trioxide in melanoma cells and cellular sensitization by ascorbic acid.
The NER proteins are differentially expressed in ever smokers and in never smokers with lung adenocarcinoma.
Human HMGB1 directly facilitates interactions between nucleotide excision repair proteins on triplex-directed psoralen interstrand crosslinks.
Distinct functions of POT1 at telomeres.
Cellular response to oxidative stress and ascorbic acid in melanoma cells overexpressing gamma-glutamyltransferase.
Intracellular accumulation and DNA damage persistence as determinants of human squamous cell carcinoma hypersensitivity to the novel camptothecin ST1968.
Mobility and distribution of replication protein A in living cells using fluorescence correlation spectroscopy.
Autoantibodies against the replication protein A complex in systemic lupus erythematosus and other autoimmune diseases.
The dispersal of replication proteins after Etoposide treatment requires the cooperation of Nbs1 with the ataxia telangiectasia Rad3-related/Chk1 pathway.
Development of resistance to the atypical retinoid, ST1926, in the lung carcinoma cell line H460 is associated with reduced formation of DNA strand breaks and a defective DNA damage response.
Mcm10 regulates the stability and chromatin association of DNA polymerase-alpha.
Induction of apoptosis and stress response in ovarian carcinoma cell lines treated with ST1926, an atypical retinoid.
The 32-kilodalton subunit of replication protein A interacts with menin, the product of the MEN1 tumor suppressor gene.
The 32-kilodalton subunit of replication protein A interacts with menin, the product of the MEN1 tumor suppressor gene.
Ju MK, Shin KJ, Lee JR, Khim KW, A Lee E, Ra JS, Kim BG, Jo HS, Yoon JH, Kim TM, Myung K, Choi JH, Kim H, Chae YC
Nucleic acids research 2021 Jun 4;49(10):5605-5622
Nucleic acids research 2021 Jun 4;49(10):5605-5622
CARM1 regulates replication fork speed and stress response by stimulating PARP1.
Genois MM, Gagné JP, Yasuhara T, Jackson J, Saxena S, Langelier MF, Ahel I, Bedford MT, Pascal JM, Vindigni A, Poirier GG, Zou L
Molecular cell 2021 Feb 18;81(4):784-800.e8
Molecular cell 2021 Feb 18;81(4):784-800.e8
DDX11 loss causes replication stress and pharmacologically exploitable DNA repair defects.
Jegadesan NK, Branzei D
Proceedings of the National Academy of Sciences of the United States of America 2021 Apr 27;118(17)
Proceedings of the National Academy of Sciences of the United States of America 2021 Apr 27;118(17)
APE1 senses DNA single-strand breaks for repair and signaling.
Lin Y, Raj J, Li J, Ha A, Hossain MA, Richardson C, Mukherjee P, Yan S
Nucleic acids research 2020 Feb 28;48(4):1925-1940
Nucleic acids research 2020 Feb 28;48(4):1925-1940
A phosphorylation-and-ubiquitylation circuitry driving ATR activation and homologous recombination.
Dubois JC, Yates M, Gaudreau-Lapierre A, Clément G, Cappadocia L, Gaudreau L, Zou L, Maréchal A
Nucleic acids research 2017 Sep 6;45(15):8859-8872
Nucleic acids research 2017 Sep 6;45(15):8859-8872
Restoration of Replication Fork Stability in BRCA1- and BRCA2-Deficient Cells by Inactivation of SNF2-Family Fork Remodelers.
Taglialatela A, Alvarez S, Leuzzi G, Sannino V, Ranjha L, Huang JW, Madubata C, Anand R, Levy B, Rabadan R, Cejka P, Costanzo V, Ciccia A
Molecular cell 2017 Oct 19;68(2):414-430.e8
Molecular cell 2017 Oct 19;68(2):414-430.e8
ATR kinase inhibition induces unscheduled origin firing through a Cdc7-dependent association between GINS and And-1.
Moiseeva T, Hood B, Schamus S, O'Connor MJ, Conrads TP, Bakkenist CJ
Nature communications 2017 Nov 9;8(1):1392
Nature communications 2017 Nov 9;8(1):1392
Functions of Replication Protein A as a Sensor of R Loops and a Regulator of RNaseH1.
Nguyen HD, Yadav T, Giri S, Saez B, Graubert TA, Zou L
Molecular cell 2017 Mar 2;65(5):832-847.e4
Molecular cell 2017 Mar 2;65(5):832-847.e4
PC4 promotes genome stability and DNA repair through binding of ssDNA at DNA damage sites.
Mortusewicz O, Evers B, Helleday T
Oncogene 2016 Feb 11;35(6):761-70
Oncogene 2016 Feb 11;35(6):761-70
DNA end resection is needed for the repair of complex lesions in G1-phase human cells.
Averbeck NB, Ringel O, Herrlitz M, Jakob B, Durante M, Taucher-Scholz G
Cell cycle (Georgetown, Tex.) 2014;13(16):2509-16
Cell cycle (Georgetown, Tex.) 2014;13(16):2509-16
Gap-filling and bypass at the replication fork are both active mechanisms for tolerance of low-dose ultraviolet-induced DNA damage in the human genome.
Quinet A, Vessoni AT, Rocha CR, Gottifredi V, Biard D, Sarasin A, Menck CF, Stary A
DNA repair 2014 Feb;14:27-38
DNA repair 2014 Feb;14:27-38
Scaffold functions of 14-3-3 adaptors in B cell immunoglobulin class switch DNA recombination.
Lam T, Thomas LM, White CA, Li G, Pone EJ, Xu Z, Casali P
PloS one 2013;8(11):e80414
PloS one 2013;8(11):e80414
BLM SUMOylation regulates ssDNA accumulation at stalled replication forks.
Ouyang KJ, Yagle MK, Matunis MJ, Ellis NA
Frontiers in genetics 2013;4:167
Frontiers in genetics 2013;4:167
Mammalian Exo1 encodes both structural and catalytic functions that play distinct roles in essential biological processes.
Schaetzlein S, Chahwan R, Avdievich E, Roa S, Wei K, Eoff RL, Sellers RS, Clark AB, Kunkel TA, Scharff MD, Edelmann W
Proceedings of the National Academy of Sciences of the United States of America 2013 Jul 2;110(27):E2470-9
Proceedings of the National Academy of Sciences of the United States of America 2013 Jul 2;110(27):E2470-9
Zinc finger protein 668 interacts with Tip60 to promote H2AX acetylation after DNA damage.
Hu R, Wang E, Peng G, Dai H, Lin SY
Cell cycle (Georgetown, Tex.) 2013 Jul 1;12(13):2033-41
Cell cycle (Georgetown, Tex.) 2013 Jul 1;12(13):2033-41
Replication stress links structural and numerical cancer chromosomal instability.
Burrell RA, McClelland SE, Endesfelder D, Groth P, Weller MC, Shaikh N, Domingo E, Kanu N, Dewhurst SM, Gronroos E, Chew SK, Rowan AJ, Schenk A, Sheffer M, Howell M, Kschischo M, Behrens A, Helleday T, Bartek J, Tomlinson IP, Swanton C
Nature 2013 Feb 28;494(7438):492-496
Nature 2013 Feb 28;494(7438):492-496
Differential contribution of HP1 proteins to DNA end resection and homology-directed repair.
Soria G, Almouzni G
Cell cycle (Georgetown, Tex.) 2013 Feb 1;12(3):422-9
Cell cycle (Georgetown, Tex.) 2013 Feb 1;12(3):422-9
Cyclin-dependent kinase suppression by WEE1 kinase protects the genome through control of replication initiation and nucleotide consumption.
Beck H, Nähse-Kumpf V, Larsen MS, O'Hanlon KA, Patzke S, Holmberg C, Mejlvang J, Groth A, Nielsen O, Syljuåsen RG, Sørensen CS
Molecular and cellular biology 2012 Oct;32(20):4226-36
Molecular and cellular biology 2012 Oct;32(20):4226-36
REV1 and polymerase ζ facilitate homologous recombination repair.
Sharma S, Hicks JK, Chute CL, Brennan JR, Ahn JY, Glover TW, Canman CE
Nucleic acids research 2012 Jan;40(2):682-91
Nucleic acids research 2012 Jan;40(2):682-91
ATM modulates the loading of recombination proteins onto a chromosomal translocation breakpoint hotspot.
Sun J, Oma Y, Harata M, Kono K, Shima H, Kinomura A, Ikura T, Suzuki H, Mizutani S, Kanaar R, Tashiro S
PloS one 2010 Oct 27;5(10):e13554
PloS one 2010 Oct 27;5(10):e13554
A PP4 phosphatase complex dephosphorylates RPA2 to facilitate DNA repair via homologous recombination.
Lee DH, Pan Y, Kanner S, Sung P, Borowiec JA, Chowdhury D
Nature structural & molecular biology 2010 Mar;17(3):365-72
Nature structural & molecular biology 2010 Mar;17(3):365-72
Sensitization of ovarian carcinoma cells to the atypical retinoid ST1926 by the histone deacetylase inhibitor, RC307: enhanced DNA damage response.
Zuco V, Benedetti V, De Cesare M, Zunino F
International journal of cancer 2010 Mar 1;126(5):1246-55
International journal of cancer 2010 Mar 1;126(5):1246-55
ATM- and ATR-mediated response to DNA damage induced by a novel camptothecin, ST1968.
Zuco V, Benedetti V, Zunino F
Cancer letters 2010 Jun 28;292(2):186-96
Cancer letters 2010 Jun 28;292(2):186-96
Deficiency of FANCD2-associated nuclease KIAA1018/FAN1 sensitizes cells to interstrand crosslinking agents.
Kratz K, Schöpf B, Kaden S, Sendoel A, Eberhard R, Lademann C, Cannavó E, Sartori AA, Hengartner MO, Jiricny J
Cell 2010 Jul 9;142(1):77-88
Cell 2010 Jul 9;142(1):77-88
Human Fbh1 helicase contributes to genome maintenance via pro- and anti-recombinase activities.
Fugger K, Mistrik M, Danielsen JR, Dinant C, Falck J, Bartek J, Lukas J, Mailand N
The Journal of cell biology 2009 Sep 7;186(5):655-63
The Journal of cell biology 2009 Sep 7;186(5):655-63
The cohesin complex is required for the DNA damage-induced G2/M checkpoint in mammalian cells.
Watrin E, Peters JM
The EMBO journal 2009 Sep 2;28(17):2625-35
The EMBO journal 2009 Sep 2;28(17):2625-35
Ionizing radiation-dependent and independent phosphorylation of the 32-kDa subunit of replication protein A during mitosis.
Stephan H, Concannon C, Kremmer E, Carty MP, Nasheuer HP
Nucleic acids research 2009 Oct;37(18):6028-41
Nucleic acids research 2009 Oct;37(18):6028-41
Gamma-glutamyltransferase-dependent resistance to arsenic trioxide in melanoma cells and cellular sensitization by ascorbic acid.
Giommarelli C, Corti A, Supino R, Favini E, Paolicchi A, Pompella A, Zunino F
Free radical biology & medicine 2009 Jun 1;46(11):1516-26
Free radical biology & medicine 2009 Jun 1;46(11):1516-26
The NER proteins are differentially expressed in ever smokers and in never smokers with lung adenocarcinoma.
Planchard D, Domont J, Taranchon E, Monnet I, Tredaniel J, Caliandro R, Validire P, Besse B, Soria JC, Fouret P
Annals of oncology : official journal of the European Society for Medical Oncology 2009 Jul;20(7):1257-63
Annals of oncology : official journal of the European Society for Medical Oncology 2009 Jul;20(7):1257-63
Human HMGB1 directly facilitates interactions between nucleotide excision repair proteins on triplex-directed psoralen interstrand crosslinks.
Lange SS, Reddy MC, Vasquez KM
DNA repair 2009 Jul 4;8(7):865-72
DNA repair 2009 Jul 4;8(7):865-72
Distinct functions of POT1 at telomeres.
Barrientos KS, Kendellen MF, Freibaum BD, Armbruster BN, Etheridge KT, Counter CM
Molecular and cellular biology 2008 Sep;28(17):5251-64
Molecular and cellular biology 2008 Sep;28(17):5251-64
Cellular response to oxidative stress and ascorbic acid in melanoma cells overexpressing gamma-glutamyltransferase.
Giommarelli C, Corti A, Supino R, Favini E, Paolicchi A, Pompella A, Zunino F
European journal of cancer (Oxford, England : 1990) 2008 Mar;44(5):750-9
European journal of cancer (Oxford, England : 1990) 2008 Mar;44(5):750-9
Intracellular accumulation and DNA damage persistence as determinants of human squamous cell carcinoma hypersensitivity to the novel camptothecin ST1968.
Pisano C, Zuco V, De Cesare M, Benedetti V, Vesci L, Foderà R, Bucci F, Aulicino C, Penco S, Carminati P, Zunino F
European journal of cancer (Oxford, England : 1990) 2008 Jun;44(9):1332-40
European journal of cancer (Oxford, England : 1990) 2008 Jun;44(9):1332-40
Mobility and distribution of replication protein A in living cells using fluorescence correlation spectroscopy.
Braet C, Stephan H, Dobbie IM, Togashi DM, Ryder AG, Földes-Papp Z, Lowndes N, Nasheuer HP
Experimental and molecular pathology 2007 Apr;82(2):156-62
Experimental and molecular pathology 2007 Apr;82(2):156-62
Autoantibodies against the replication protein A complex in systemic lupus erythematosus and other autoimmune diseases.
Yamasaki Y, Narain S, Hernandez L, Barker T, Ikeda K, Segal MS, Richards HB, Chan EK, Reeves WH, Satoh M
Arthritis research & therapy 2006;8(4):R111
Arthritis research & therapy 2006;8(4):R111
The dispersal of replication proteins after Etoposide treatment requires the cooperation of Nbs1 with the ataxia telangiectasia Rad3-related/Chk1 pathway.
Rossi R, Lidonnici MR, Soza S, Biamonti G, Montecucco A
Cancer research 2006 Feb 1;66(3):1675-83
Cancer research 2006 Feb 1;66(3):1675-83
Development of resistance to the atypical retinoid, ST1926, in the lung carcinoma cell line H460 is associated with reduced formation of DNA strand breaks and a defective DNA damage response.
Zuco V, Zanchi C, Lanzi C, Beretta GL, Supino R, Pisano C, Barbarino M, Zanier R, Bucci F, Aulicino C, Carminati P, Zunino F
Neoplasia (New York, N.Y.) 2005 Jul;7(7):667-77
Neoplasia (New York, N.Y.) 2005 Jul;7(7):667-77
Mcm10 regulates the stability and chromatin association of DNA polymerase-alpha.
Ricke RM, Bielinsky AK
Molecular cell 2004 Oct 22;16(2):173-85
Molecular cell 2004 Oct 22;16(2):173-85
Induction of apoptosis and stress response in ovarian carcinoma cell lines treated with ST1926, an atypical retinoid.
Zuco V, Zanchi C, Cassinelli G, Lanzi C, Supino R, Pisano C, Zanier R, Giordano V, Garattini E, Zunino F
Cell death and differentiation 2004 Mar;11(3):280-9
Cell death and differentiation 2004 Mar;11(3):280-9
The 32-kilodalton subunit of replication protein A interacts with menin, the product of the MEN1 tumor suppressor gene.
Sukhodolets KE, Hickman AB, Agarwal SK, Sukhodolets MV, Obungu VH, Novotny EA, Crabtree JS, Chandrasekharappa SC, Collins FS, Spiegel AM, Burns AL, Marx SJ
Molecular and cellular biology 2003 Jan;23(2):493-509
Molecular and cellular biology 2003 Jan;23(2):493-509
The 32-kilodalton subunit of replication protein A interacts with menin, the product of the MEN1 tumor suppressor gene.
Sukhodolets KE, Hickman AB, Agarwal SK, Sukhodolets MV, Obungu VH, Novotny EA, Crabtree JS, Chandrasekharappa SC, Collins FS, Spiegel AM, Burns AL, Marx SJ
Molecular and cellular biology 2003 Jan;23(2):493-509
Molecular and cellular biology 2003 Jan;23(2):493-509
No comments: Submit comment
Supportive validation
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- Western blot analysis was performed on nuclear enriched cell extracts (30 µg lysate) of HeLa (Lane 1), MDA-MB-231 (Lane 2), Hep G2 (Lane 3), MCF7 (Lane 4), U-2 OS (Lane 5), Jurkat (Lane 6) and HEK293T (Lane 7). The blot was probed with Anti-RPA2 Monoclonal Antibody (Product # MA1-26418, 1:500 dilution) and detected by chemiluminescence using Goat anti-Mouse IgG (H+L) Superclonal™ Secondary Antibody, HRP conjugate (Product # A28177, 0.25 µg/mL, 1:4000 dilution). A 32 kDa band corresponding to RPA2 was observed across the cell lines tested.
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
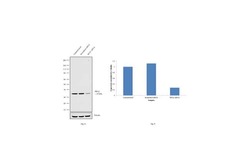
- Knockdown of RPA2 was achieved by transfecting HeLa with RPA2 specific siRNAs (Silencer® select Product # s12132, s12130). Western blot analysis (Fig. a) was performed using whole cell extracts (1% SDS) from the RPA2 knockdown cells (lane 3), non-specific scrambled siRNA transfected cells (lane 2) and untransfected cells (lane 1). The blots were probed with RPA2 Monoclonal Antibody (9H8) (Product # MA1-26418, 1:500 dilution) and Goat anti-Mouse IgG (H+L) Superclonal™ Secondary Antibody, HRP conjugate (Product # A28177, 0.25 µg/mL, 1:4000 dilution). Densitometric analysis of this western blot is shown in histogram (Fig. b). Decrease in signal upon siRNA mediated knock down confirms that antibody is specific to RPA2.
Supportive validation
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
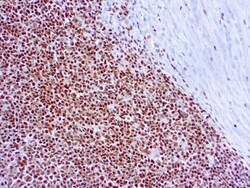
- Experimental details
- Immunohistochemical (Paraffin) analysis of human tonsil tissue using (Product # MA1-26418) RPA2 Monoclonal Antibody (9H8).
Supportive validation
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
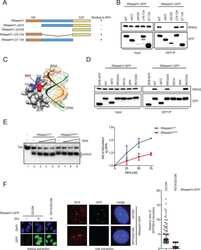
- Fig. 7. DDX11 facilitates resection and RPA loading to ssDNA substrates. ( A ) Quantification and representative micrographs of RPA32 foci in U2OS Ctrl and DDX11 KO cells recovering from an acute treatment with cisplatin (2.5 muM for 1 h) after 24 h. (Scale bar, 10 mum.) n = 2. Statistical analysis of foci was performed using Student''s t test. Error bars show average +- SD. ( B ) U2OS DIvA cells were transfected with siCtrl and si DDX11 followed by the addition of 4OHT (300 nM) to induce AsiSI-mediated DSBs after 48 h of post-transfection. Resection assay at the two different regions (KDELR3 and ASXL1) were analyzed after 4 h of 4OHT by real-time PCR. Values were normalized against the amount of ssDNA detected in control cells prior to 4OHT treatment, n >= 3. Corresponding Western blot and immunofluorescence for AsiSI-induced DSBs is shown. (Scale bar, 10 mum.) Statistical analysis was performed using Student''s t test. Error bars show average +- SEM. ( C ) Model for the DDX11 proposed role in resolving secondary structures upon DSB end resection to facilitate RPA loading and subsequently RAD51 nucleofilament formation required for homologous recombination-mediated DSB repair (see text for details).
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
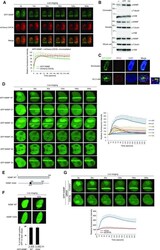
- Figure 2. NSMF localizes to DNA lesions via the central residues of NSMF. ( A ) The kinetics of GFP-NSMF and mCherry-CDC5L recruitment to DNA lesions were examined by live cell imaging. The relative signal intensities are quantified and summarized in the bottom panel. The highest intensity on any lesion was set at 100% for each cell, and the recruitment kinetics were plotted. The average intensity for >=10 cells for each condition is presented graphically. Data represent the mean +- SD. ( B ) HeLa cells were treated with 1 muM CPT for 2 h or with 2 mM HU for 16 h and then harvested as whole-cell extracts (WCEs) or fractionated into soluble or chromatin-bound protein fractions. Each fraction was analyzed by western blot with the indicated antibodies. ( C ) HeLa cells transfected with GFP-NSMF expression plasmids were treated with 2 mM HU for 16 h and then incubated in fresh media for 3 h. Colocalization of GFP-NSMF and RPA2 was determined by immunofluorescence with an RPA2 antibody. Magnified insets show colocalization of NSMF and RPA2 foci. ( D ) HeLa cells were transfected with expression plasmids for GFP-NSMF WT or GFP-NMSF deletion mutants. After 24 h, the cells were laser microirradiated, and recruitment of GFP-NSMF proteins to DNA lesions was examined by live cell imaging. Relative intensities of the lesions were quantified, and the recruitment kinetics are plotted in the right panel. The average intensity for >=10 cells for each condition is presented graphically. Data
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
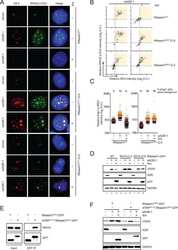
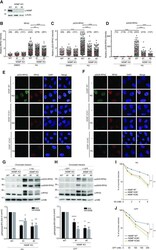
- Figure 3. NSMF is involved in the replication stress response. ( A ) The NMSF gene was deleted from HeLa cells using CRISPR/Cas9. Expression of NSMF in WT and NSMF KO HeLa-cell extracts was analyzed by western blot. ( B - F ) Localization of RPA2 and phospho-RPA2 (S4/S8 or S32) was determined by immunofluorescence. NSMF WT and KO HeLa cells were treated with 2 mM HU for 16 h, fixed and stained with the indicated antibodies. RPA2 and phospho-RPA2 intensities per nucleus were quantified for each sample using ZEN Blue software (Carl Zeiss). The numbers above each sample indicate the number of nuclei analyzed. Data are presented as the mean +- SD; *** P < 0.001, **** P < 0.0001. ( G and H ) NMSF WT and KO HeLa cells were treated with either 2 mM HU for 16 h or 1 muM CPT for 2 h. The cells were fractionated, and phosphorylation of RPA2 in the chromatin-bound protein fraction was determined by western blot with the indicated antibodies. The levels of phospho-RPA2 (S4/S8 or S32) and total RPA were quantified using ImageJ software. Quantification of the western blots represents an average of three independent experiments. Data represent the mean +- SEM of three independent experiments; * P < 0.05, ** P < 0.01, **** P < 0.0001. ( I and J ) Sensitivity of NSMF KO HeLa cells to DNA replication-blocking agents. NSMF WT and KO HeLa cells were plated and treated with the indicated concentrations of HU or CPT. The numbers of surviving colonies were counted 14 days after treatment. Data repr
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- Figure 5. NSMF directs recruitment of ATRIP-ATR to DNA lesions by mediating the interaction between RPA2 and ATRIP-ATR. ( A ) NSMF WT and KO HeLa cells were transfected with GFP-ATRIP expression plasmids. After 24 h, the cells were microirradiated, and recruitment of GFP-ATRIP to DNA lesions was examined by live cell imaging. ( B ) NSMF WT and KO HeLa cells were treated with or without 2 mM HU for 16 h. The cells were fractionated, and the chromatin-bound protein fraction was subjected to western blot with the indicated antibodies. ( C ) Cotransfection of GFP-ATRIP with the mCherry-vec, mCherry-NSMF-WT or mCherry-NSMF-D4 mutant plasmid in NSMF KO#1 and KO#2 HeLa cells. After 24 h, the transfected cells were microirradiated, and recruitment of ATRIP and NSMF to DNA lesions was examined by live cell imaging. ( D and E ) HeLa-cell lysates were immunoprecipitated with anti-IgG, anti-ATRIP or anti-RPA2 antibodies. Endogenous interactions between NSMF and ATRIP or RPA2 were determined by western blot with the indicated antibodies. ( F and G ) HEK293T cells were cotransfected with a Flag-NSMF-deletion mutant and either GFP-ATRIP or Myc-RPA2. Immunoprecipitation using an anti-FLAG antibody and western blot with the indicated antibodies were performed to identify the NSMF regions that interact with ATRIP (F) or RPA2 (G). ( H ) HEK293T cells were transfected with GFP-ATRIP or Myc-RPA2 and either FLAG-NSMF WT or FLAG-NSMF D2. Immunoprecipitation with an anti-FLAG antibody and western bl
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
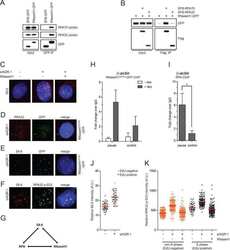
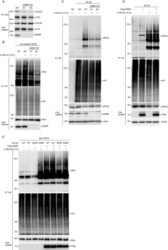
- Figure 6. NSMF mediates RPA2 ubiquitination. ( A ) Endogenous ATR in NSMF WT or KO HeLa cell lysates was immunoprecipitated with anti-ATR antibody and western blotting with the indicated antibodies was performed. ( B ) NSMF WT and KO HeLa cells were transfected with Myc-RAP2 and HA-ubiquitin and were treated with or without 2 mM HU for 3 h. Cells were lysed in denaturing conditions, diluted and ubiquitinated proteins were immunoprecipitated with anti-HA antibody and subjected to western blotting with the indicated antibodies. ( C ) NSMF WT and KO HeLa cells transfected with HA-ubiquitin were treated with or without 2 mM HU for 3 h. Cells were lysed in denaturing conditions, diluted, and ubiquitinated proteins were immunoprecipitated with anti-HA antibody. Endogenous RPA2 ubiquitination was detected by immunoblotting with the indicated antibodies. ( D ) HeLa cells transfected with HA-ubiquitin and Flag-NSMF were treated with or without 2 mM HU for 3 h. Cells were lysed in denaturing conditions, diluted, ubiquitinated proteins were immunoprecipitated with anti-HA antibody, and RPA2 ubiquitination was detected by immunoblotting with the indicated antibodies. ( E ) HeLa cells transfected with Myc-RPA2 and Flag-NSMF, plus the WT, lysine mutant K48R, or K63R form of HA-ubiquitin, were treated with or without 2 mM HU for 3 h. Cells were lysed in denaturing conditions, diluted and ubiquitinated proteins and immunoprecipitated with anti-HA antibody. RPA2 ubiquitination was detected by
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- Figure 7. NSMF participates in genome maintenance and mouse survival in response to in vivo DNA damage. ( A ) Diagram describing generation of the Nsmf KO allele using a gene targeting vector. The vector consists of a Flp-recombinase target (FRT)-flanked selection cassette inserted between the third and fourth exons of Nsmf . The Nsmf transcript is trapped through the splice acceptor (SA) element and truncated through the SV40 polyadenylation signal (pA). lacZ, beta-galactosidase; Neo, Neomycin resistance gene. The red arrow is the primer for genotyping. ( B ) Genotyping of Nsmf +/+, Nsmf +/- and Nsmf -/- mice using tail DNA. The sizes of the PCR products for the WT and mutant alleles are 343 and 292 bp, respectively. ( C and D ) Confirmation of Nsmf removal by qRT-PCR (C) and western blot analysis (D) using MEF mRNA and protein lysates, respectively. ( E ) Impaired interaction between RPA2 and ATRIP in NSMF KO MEF cells. Endogenous RPA2 in NSMF WT and KO MEF cell lysates was immunoprecipitated with anti-RPA2 antibody, and western blotting with the indicated antibodies was performed. ( F and G ) NSMF WT or KO cells were treated with 2 mM HU for 24 h, transferred to fresh media and subjected to metaphase spreading. (F) Representative images of spontaneous and HU-induced chromosome instability in NSMF WT and KO MEFs. The red dots indicate chromosomal abnormalities, such as chromosome breakage, fragments and radial figures. (G) The numbers of abnormal chromosomes per cell were p
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- Figure 3. RPA32 ubiquitylation is regulated by PI3K-like kinases. (A) Cells were transfected with a vector expressing Strep-HA-tagged ubiquitin, pre-treated for 1 h with VE-821 (ATRi, 10 muM), KU55933 (ATMi, 10 uM), NU7441 (DNAPKi, 2 muM) or a combination of all three inhibitors, treated with 1 muM CPT for 4 h and lysed under denaturing conditions. Strep-Tactin pulldown was performed to isolate ubiquitylated proteins. ( B ) Cell transfection with an SFB-PRP19 vector were treated as in A and streptavidin pulldown was performed to isolate PRP19 along with its interactors. Controls for the efficiency of the inhibitor treatments are provided in ( S2A ). ( C and D ) U2OS cells were pre-sensitized with 10 muM BrdU, treated with the indicated inhibitors along with DRB for 1 h and damaged by UV-laser microirradiation. Recruitment of PRP19 and RPA32 was monitored 2 h after damage by immunofluorescence.
 Explore
Explore Validate
Validate Learn
Learn Western blot
Western blot