Antibody data
- Antibody Data
- Antigen structure
- References [0]
- Comments [0]
- Validations
- Flow cytometry [1]
- Other assay [69]
Submit
Validation data
Reference
Comment
Report error
- Product number
- 53-0037-41 - Provider product page

- Provider
- Invitrogen Antibodies
- Product name
- CD3 Monoclonal Antibody (OKT3), Alexa Fluor™ 488, eBioscience™
- Antibody type
- Monoclonal
- Antigen
- Other
- Description
- Description: The OKT3 monoclonal antibody reacts with an epitope on the epsilon-subunit within the human CD3 complex. The OKT3 antibody has been reported to have potent immunosuppressive properties in vivo and has been proven effective in the treatment of renal, heart and liver allograft rejection. The CD3 subunits, gamma, delta, and epsilon chains, are required for proper assembly, trafficking and surface expression of the TCR complex. CD3 is expressed by thymocytes in a developmentally regulated manner and by all mature T cells. Crosslinking of TCR initiates an intracellular biochemical pathway resulting in cellular activation and proliferation. Applications Reported: This OKT3 antibody has been reported for use in flow cytometric analysis. Applications Tested: This OKT3 antibody has been pre-titrated and tested by flow cytometric analysis of normal human peripheral blood cells. This can be used at 5 µL (1 µg) per test. A test is defined as the amount (µg) of antibody that will stain a cell sample in a final volume of 100 µL. Cell number should be determined empirically but can range from 10^5 to 10^8 cells/test. Excitation: 488 nm; Emission: 519 nm; Laser: Blue Laser. Filtration: 0.2 µm post-manufacturing filtered.
- Reactivity
- Human
- Host
- Mouse
- Conjugate
- Green dye
- Isotype
- IgG
- Antibody clone number
- OKT3
- Vial size
- 25 Tests
- Concentration
- 5 µL/Test
- Storage
- 4° C, store in dark, DO NOT FREEZE!
No comments: Submit comment
Supportive validation
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- Staining of normal human peripheral blood cells with Anti-Human CD19 PE (Product # 12-0199-80) and staining buffer (autofluorescence) (left) or Anti-Human CD3 Alexa Fluor® 488 (right). Cells in the lymphocyte gate were used for analysis.
- Conjugate
- Green dye
Supportive validation
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
- NULL
- Conjugate
- Green dye
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
- NULL
- Conjugate
- Green dye
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
- NULL
- Conjugate
- Green dye
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
- NULL
- Conjugate
- Green dye
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
- NULL
- Conjugate
- Green dye
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
- NULL
- Conjugate
- Green dye
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
- NULL
- Conjugate
- Green dye
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
- NULL
- Conjugate
- Green dye
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
- NULL
- Conjugate
- Green dye
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
- NULL
- Conjugate
- Green dye
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
- NULL
- Conjugate
- Green dye
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
- NULL
- Conjugate
- Green dye
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
- NULL
- Conjugate
- Green dye
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
- NULL
- Conjugate
- Green dye
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
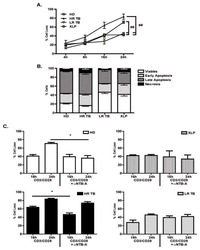
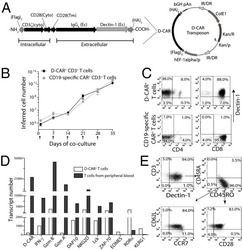
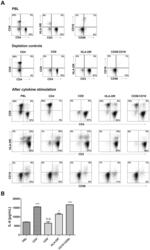
- Figure 9 IL-6 produced by PBLs depleted for CD4 + , CD8 + , HLA-DR + or CD56 + /CD16 + cells. (A) Freshly purified PBLs stained for CD4, CD8, CD3, HLA-DR, CD16, CD56. Depletion controls for PBLs depleted for CD4 + , CD8 + , HLA-DR + or CD16 + /CD56 + cells. PBLs and depletions were stimulated for 7 days with combinations of IL-12 (4 ng/mL), IL-15 (10 ng/mL), IL-18 (40 ng/mL) and TL1A (100 ng/mL) and stained for CD4, CD8, CD3, HLA-DR, CD16 and CD56 expression. (B) IL-6 production after 7 days by PBLs and depletions stimulated with IL-12, IL-15, IL-18 and TL1A as described above. Error bars represent the SEM of two measurements. Statistically significant differences by t-test: *** = p
- Conjugate
- Green dye
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
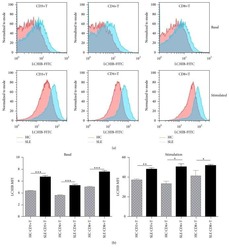
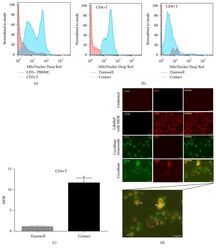
- Figure 7 UC-MSCs transferred mitochondria to activate T cells. PBMCs from SLE patients were labeled with carboxyfluorescein succinimidyl amino ester (CFSE) and treated with anti-CD3/CD28 antibodies for two days. Then they were cocultured with UC-MSCs for 12 h, which had been prelabeled with respiratory mitochondrion specific probe Mitotracker Deep Red (MDR). PBMCs were cultured with UC-MSCs through transwell as control. Then PBMCs were stained with anti-CD3, anti-CD4, or anti-CD8 dye and detected for MDR fluorescence with flow cytometry. Fluorescence microscopy was carried out similarly with anti-CD3/28 stimulation for 12 h and then cocultured for 6 h. All experiments were performed in triplicate. (a) T cells (CD3 positive) rather than non-T cells (CD3 negative) got MDR staining. (b-d) T cells cultured with UC-MSCs directly rather than through transwell got transferred mitochondria ( n = 3). Arrows indicated transferred mitochondria within lymphocytes. * p < 0.05.
- Conjugate
- Green dye
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
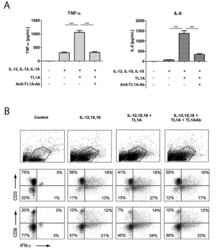
- Figure 2 TL1A induces IL-6 and TNF-alpha. Freshly purified PBMCs were incubated with IL-12 (2 ng/mL), IL-15 (10 ng/mL), IL-18 (10 ng/mL), TL1A (100 ng/mL) and TL1AAb (1 ug/mL, blocking antibody). Extra IL-15 (2 ng/mL) was added on day 3. ( A ) After 6 days, supernatants were collected and different cytokines were measured by bead-based ELISA. Error bars represent the SEM of eight measurements. Statistically significant differences are indicated by ***(t-test, P
- Conjugate
- Green dye
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
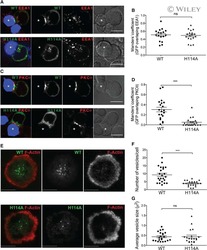
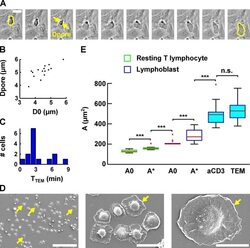
- FIGURE 7: Evolution of lymphoblast membrane surface area during transendothelial migration and cell spreading. (A) Time lapse of a lymphoblast transmigrating between human aortic endothelial cells. Scale bar, 20 mum. Images were taken every 15 s. The projected surface area S proj is represented in yellow before (leftmost image) and after (rightmost image) transendothelial migration. The pore diameter, D pore , is estimated by taking the image in which the lymphoblast width is identical above and below the pore (yellow arrows). (B) D pore during transendothelial migration as a function of the lymphoblast's projected diameter before transendothelial migration (computed using D 0 = S proj /2pi, and D 0 is an equivalent diameter for a sphere whose projected area is S proj ). (C) Histogram of the duration of transmigration. The mean duration is 3 +- 2 min (mean +- SD). (D) Scanning electron microscopy images of lymphoblasts spreading on a substrate coated with anti-CD3 plus anti-CD28 activating antibodies. Scale bars, 100 mum, 20 mum, 10 mum (left to right). Yellow arrows indicate spread cells. (E) Boxplots of the apparent membrane surface area of T-lymphocytes under both passive (white-filled box) and active (blue-filled box) deformations. The bottom and top of the box indicate the 25th and 75th percentiles, respectively. Red plus signs indicate outliers. From left to right, resting T-lymphocytes initially (column 1, A 0 , n = 14) and at rupture (column 2, A *, n = 14) aspir
- Conjugate
- Green dye
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
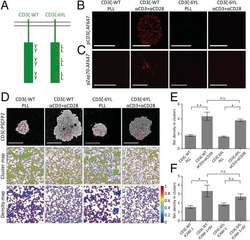
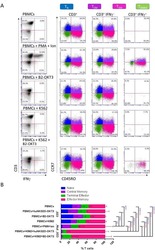
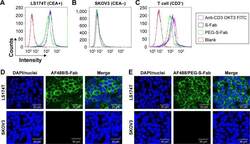
- Figure 4 PEG-S-Fab can bind CEA on tumor cells and CD3 + on T cells. Notes: Flow cytometry analysis of PEG-S-Fab and S-Fab on CEA-positive LS174T cells ( A ), CEA-negative SKOV3 cells ( B ), and CD3 + T cells ( C ). The positive control anti-CD3 antibody OKT3 was used for T-cell flow cytometry. Confocal microscopy of immunofluorescence staining as described in the ""Materials and methods"" section. S-Fab ( D ) and PEG-S-Fab ( E ) on LS174T cells (upper panel) and SKOV3 cells (lower panel), respectively. The scale bar represents 30 mum. AF488, stained by the goat antihuman IgG (H + L)-AlexaFluor 488 antibody; LS174T, human colorectal cell line LS174T; SKOV3, human ovarian cancer cell line SKOV3. Abbreviations: CEA, carcinoembryonic antigen; DAPI, 4',6-diamidino-2-phenylindole; IgG, immunoglobulin G; PEG, polyethylene glycol; PEG-S-Fab, PEGylated S-Fab; S-Fab, single-domain antibody-linked Fab; FITC, fluoresceine isothiocyanate.
- Conjugate
- Green dye
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
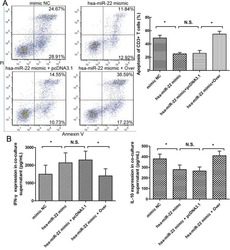
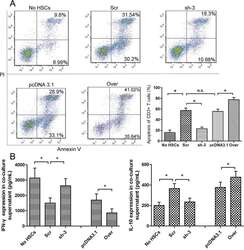
- Figure 4 The expression of galectin-1 in HSCs promotes HSC-induced T cell apoptosis and Th1/Th2 cytokine balance skewing A. Flow cytometry (annexin V-FITC apoptosis detection) analyses to detect T cell apoptosis in CD3+ T cells, cultured alone or co-cultured with HSCs subjected to different pre-treatments (cell transfection for galectin-1 knockdown and overexpression: sh-3 group versus Scr group; Over group versus pcDNA3.1 group) for 48 hours at a ratio of 10:1 (T:HSC), B. ELISA showing the levels of cytokines (IFN-gamma and IL-10) in the supernatant. Data are shown as the means (+- SD) of triplicates (n = 7). *P < 0.05. NC, negative control group; Scr, non-targeting scrambled sequence group; sh, small hairpin RNA sequence transfection group; pcDNA3.1, negative control group; Over, galectin-1 overexpression group; No HSCs, T cells cultured alone.
- Conjugate
- Green dye
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- Figure 2 Cell surface expression of SAIL in CLL, AML and MM patient samples and normal BMMC and PBMC controls. ( a ) Three CLL specimens analyzed by flow cytometry. CLL cells were identified as CD19/CD5 double-positive cells. The histograms present SAIL (filled) and isotype control (open) staining in the live-cell and the CLL population. ( b ) Flow cytometry analysis of three AML specimens. SAIL expression is assessed in live-cells, CD33-positive and CD34-positive cells. ( c ) Flow cytometry analysis of three MM specimens. CD38 high cells with CD56 expression were gated for MM cells. SAIL expression is assessed in the live-cell and the MM population. ( d and e ) Flow cytometry analysis of SAIL expression in BMMC ( d ) and PBMC ( e ) via co-staining with CD19, CD3, CD14, CD56, CD33, CD34 and a cocktail of lineage (LN) markers. Numbers in histograms are median-fluorescence-intensity fold-change values relative to the isotype control. Three and two representative examples are shown for the tumor and normal samples, respectively.
- Conjugate
- Green dye
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
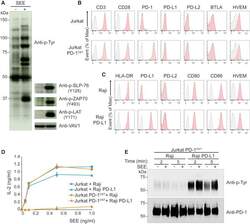
- Figure 3 Outcome of PD-1-PD-L1 Engagement at T Cell-APC Interface (A) Jurkat-PD-1 OST cells were stimulated with Raji cells that have been preincubated in the absence (-) or presence (+) of 200 ng/mL SEE and lysed 2 min after the initiation of cell-cell contact. Immunoblot analysis of equal amounts of lysates from the specified conditions probed with antibody to phosphorylated proteins (Anti-p-Tyr) or with phospho-tyrosine-specific antibodies directed against SLP76 pY128, ZAP70 pY493, LAT pY171 or VAV1 (loading control). Left margin, molecular size in kilodaltons (kDa). Data are representative of three independent experiments. (B) Expression of CD3, CD28, PD-1, PD-L1, PD-L2, BTLA, and HVEM at the surface of Jurkat cells and Jurkat-PD-1 OST cells, analyzed using flow cytometry. (C) Expression of HLA-DR, PD-L1, PD-L2, CD80, CD86, and HVEM at the surface of Raji cells and Raji-PD-L1 cells, analyzed using flow cytometry. In (B) and (C), gray shaded curves correspond to isotype-matched control antibody (negative control), and data are representative of two independent experiments. (D) IL-2 production by Jurkat and Jurkat-PD-1 OST cells stimulated for 24 h with either Raji or Raji-PD-L1 cells in the absence (0) or presence of the specified amounts of SEE. Data are representative of three independent experiments, and mean and SEM are shown. (E) Jurkat-PD-1 OST cells stimulated at 37degC with Raji or Raji-PD-L1 cells preincubated in the absence (-) or presence (+) of 200 ng/mL SEE an
- Conjugate
- Green dye
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
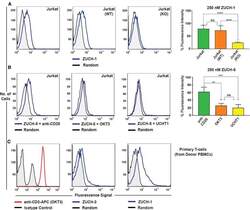
- Figure 6 Characterization of Aptamer Specificity against TCR-CD3epsilon (A) Flow-cytometric analyses of binding of the highest affinity aptamer, ZUCH-1, against Jurkat.E6 cells used in SELEX (left), against wild-type Jurkat cells used for generating CRISPR knockout cell lines (middle), against CRISPR double-knockout Jurkat cells (right), and the overall conclusion from six independent specificity analyses (far right). Aptamer ZUCH-1 does not bind to knockout cells, thereby demonstrating epitope specificity (ordinary one-way ANOVA, using Tukey''s multiple comparisons test performed on GraphPad Prism to obtain statistical significance: ****p
- Conjugate
- Green dye
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
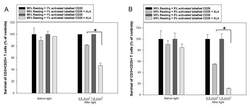
- Figure 7 ALA-PDT of mixed populations of resting and activated cells. Healthy donor PBMCs were activated in vitro with anti-CD3/CD28 antibodies for three days. The activated T cells were then labeled with anti-human CD25-FITC antibody. The resting and CD25-FITC labeled activated T cells were mixed in certain ratios as indicated. The mixed cells were incubated with 3 mM ALA for 1 h at 37 degC and then irradiated with the LED blue light at 0.9 J/cm 2 or 1.8 J/cm 2 . The cell survivals were measured 2 h after light irradiation with flow cytometry as described in Figure 3 . The control samples without light are also included. ( A ) Mixture of 1% CD25-FITC labelled activated T cells with 99% resting PBMCs. ( B ) Mixture of 5% CD25-FITC labelled activated T cells with 95% resting PBMCs. * p < 0.05.
- Conjugate
- Green dye
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
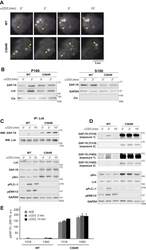
- Figure 3 Acylation-deficient ZAP-70 exhibits increased phosphorylation at Y319 and Y493. A , TIRF imaging of P116 (ZAP-70 -/-) Jurkat T cells transiently transfected with mCherry-tagged WT or acylation-deficient C564R ZAP70. Both WT ZAP-70 and C564R ZAP-70 showed similar localization patterns by forming puncta at the surface of the resting or stimulated cells. Shown are representative cells from three independent experiments. B , WT and C564R ZAP-70 were analyzed for membrane association by subcellular fractionation. GAPDH and calnexin were used as loading controls for cytosolic (S100) and membrane (P100) fractions, respectively. C , Coimmunoprecipitation of Lck and ZAP-70. P116 Jurkat T cells stably expressing WT or C564R ZAP-70 were stimulated with anti-CD3 antibody for the indicated time points. Lck was immunoprecipitated from the lysates, and the presence of ZAP-70 was assessed in eluates. D , Western blot analysis of ZAP-70 phosphorylation at Y319 and Y493. P116 Jurkat T cells stably expressing WT ZAP-70 or C564R ZAP-70 were stimulated with anti-CD3 antibody for the indicated time points, and phosphorylation of ZAP-70 was analyzed by immunoblotting. E , Quantified data showing phosphorylation of WT or C564R ZAP-70. Data shown are representative of three independent biological repeats and represented as mean +- SEM, normalized to total ZAP-70.
- Conjugate
- Green dye
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
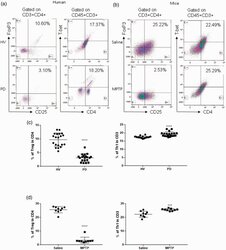
- Figure 1. Patients with Parkinson's disease (PD) and mice with experimental PD exhibit decreased regulatory T (Treg) and increased T helper 1 (Th1) cell numbers in the blood. Treg cells were defined as CD3+CD4+CD25+FoxP3+ cells, whereas Th1 cells were identified as CD3+CD4+T-bet+ cells. (a) Representative plots of Treg and Th1 cells in patients with PD and healthy volunteers (HVs). (b) Representative plots of Treg and Th1 cells in control mice (Saline) and 1-methyl-4-phenyl-1,2,3,6-tetrahydropyridine (MPTP)-treated mice. (c) Patients with PD exhibited lower Treg cell levels and greater Th1 cell levels in the blood than HVs. n = 20. ****, p < 0.0001 by a two-tailed Student's t tests. (d) MPTP-induced experimental PD mice had fewer Treg and more Th1 cells in the circulation than saline-treated mice. n = 9/group in each experiment performed in triplicate. ***, p < 0.001; ****, p < 0.0001, according to Student's t test.
- Conjugate
- Green dye
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
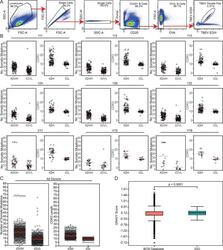
- Figure S2. Sorting strategy and antibody sequence analysis. (A) Sorting strategy. Forward and side scatter (FSC and SSC, respectively) were used to gate on single lymphocytes. Dump channel included CD3, CD8, CD14, CD16, and a viability dye. CD20 + B cells that failed to bind OVA (OVA - ) but did bind to the TBEV EDIII bait coupled with both PE and AF647 fluorophores were purified. (B) For each donor, the number of V gene somatic nucleotide mutations is shown on the left and the amino acid length of the CDR3 is shown on the right. (C) As in B, but for all donors combined. For B and C, horizontal red lines indicate the mean. (D) Distribution of hydrophobicity GRAVY scores at the IGH CDR3 of antibodies from all donors combined and compared with human repertoire (). P < 0.0001 was determined using the Wilcoxon nonparametric test. Related to Fig. 2 , Table S2 , and Table S3 .
- Conjugate
- Green dye
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
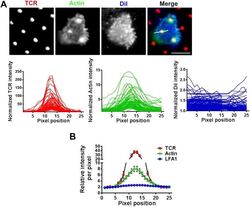
- Figure 5--figure supplement 2. F-actin foci on microdots are not enriched in phospholipid membranes. ( A ) F-actin enrichment at the microdot is not a consequence of cell morphology around the microdot. Human CD4 T cells were labeled with CM-DiI according to manufacturer's protocol and were then incubated with glass coverslip coated with patterned Alexa647 tagged anti-CD3 and ICAM1 for 5 min, fixed and stained with Alexa488-phalloidin. Cells were then imaged using TIRF microscopy. The graphs on the bottom panels show line-scan profiles of anti-CD3 (TCR, red, left), actin (Green, center) and DiI (blue, right) acquired from 70 different microdots. The TCR, actin and DiI intensities were measured from identical pixel positions for a given microdot, were then normalized by the lowest pixel intensity per microdot for a given fluorescent channel, and plotted. Note that while actin shows enrichment at the microdot site, DiI is not enriched at the same position. ( B ) The same procedure as described above, except with the use of Cy3-anti- LFA1 Fab fragments instead of DiI, was carried out in T cells, and the normalized line-scan intensities were plotted as average value per pixel position +-SEM. The graph represents mean of normalized intensities across 25 pixels, acquired using 74 different microdots. In both these experiments ( A , B ), n > 20. DOI: http://dx.doi.org/
- Conjugate
- Green dye
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
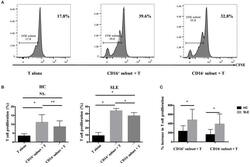
- Figure 6 CD16 + monocytes promoted T-cell proliferation in SLE . CFSE-labeled CD4 + T cells isolated from freshly collected SLE blood or blood bank collected HC blood buffy coat were cocultured with CD16 + and CD16 - monocytes for 60 h in the presence of anti-CD3 (1 mug/mL) and anti-CD 28 (1 mug/mL) antibodies and M-CSF (50 ng/mL). CFSE histograms depicted the number of events ( y -axis) and the fluorescence intensity ( x -axis), with proliferating cells displaying a progressive loss in fluorescence intensity following cell division, indicative of proliferating cells. (A) Representative histogram plots from a patient with SLE displayed the frequencies of CFSE low T cells in monocyte/T-cell cocultures and T cells cultured alone. (B) Percentages of CFSE low T cells in CD4 + T cells after coculture with each monocyte subset from five healthy donors and six patients with SLE. (C) The percentage increase in CFSE low T cells in cocultures of monocytes and T cells compared with CD4 + T cells cultured alone was compared between five healthy individuals and six patients with SLE. Data were expressed as mean +- SD and analyzed by non-parametric paired t test and Mann-Whitney U test. * P < 0.05, ** P < 0.01; NS, no significance.
- Conjugate
- Green dye
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
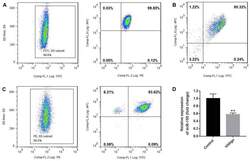
- Figure 1. Purity of CD3 + CD4 + CD45RA + T cells, CD3 + CD8 + T cells and CD4 + CD25 + FoxP3 + Treg cells. CD3 + CD4 + CD45RA + T cells and CD3 + CD8 + T cells were purified by magnetic cell sorting, and their purity was determined by flow cytometry. (A) The purity of CD3 + CD4 + CD45RA + T cells was 99.45% (CD3 + T cells, 99.6%; CD4 + CD45RA + T cells, 99.85%). (B) The purity of CD3 + CD8 + T cells was 95.32%. (C) The purity of CD4 + CD25 + FoxP3 + Treg cells was 93.15% (CD4 + T cells, 99.5%; CD25 + FoxP3 + T cells, 93.62%). (D) miR-155 expression in T cells of the patients with vitiligo and healthy donor was detected by reverse transcription quantitative polymerase chain reaction. **P
- Conjugate
- Green dye
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
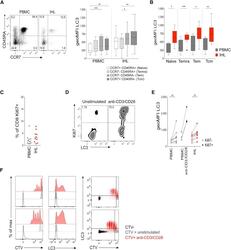
- Figure 2 High Autophagy Level of Intrahepatic T Cells Is Not a Result of a Difference in Differentiation Status or Recent Proliferation (A) Example plot of CD45RA versus CCR7 staining (CD8 + T cells) from a PBMC or IHL sample and summary data for LC3 staining of CD8 + T cell memory subsets (PBMC, 9; and IHL, 15, biological replicates; box whisker, Tukey). (B) Comparison of LC3 staining of CD8 + T cell memory subsets between paired PBMC and IHL samples (9 biological replicates; box whisker, Tukey; outliers shown as dots). (C) Ex vivo CD8 + T cell Ki67 expression. (D and E) Example plots (CD8 + T cells, PBMC) (D) and summary data for LC3 staining on Ki67 - and Ki67 + CD8 + T cells (E) in PBMCs and IHLs ex vivo (10 biological replicates) or after anti-CD3/CD28 stimulation (overnight, 0.5 mug/mL each; three biological replicates) in PBMCs. (F) Histograms showing the dilution of CellTrace Violet (CTV), LC3 staining, and co-staining of LC3 and CTV on CD8 + T cells after 5 days of stimulation with anti-CD3/CD28 (red), compared with that without stimulation (gray) or without CTV staining (black; two representative biological replicates of five, PBMCs). Cells were treated with bafA1 (A-F). Friedman test (ANOVA) with Dunn's post hoc test for pairwise multiple comparisons (A and B). Mann-Whitney t test (C and E). * p < 0.05, ** p < 0.005, *** p < 0.001.
- Conjugate
- Green dye
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
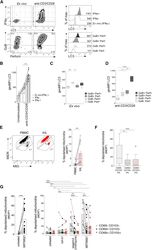
- Figure 4 Enhanced Autophagy Levels Are Linked to Effector Function and Mitochondrial Fitness in Human T Cells (A) Example plots of IFN-gamma, LC3, granzyme B (GzB), and perforin (perf; gated on CD8 + T cells) and histograms of LC3 staining for PBMC ex vivo or after anti-CD3/CD28 stimulation (3 days; see also Figure S3 ). (B) LC3 staining of CD8 + T cells from unstimulated PBMCs (IFN-gamma - ), IFN-gamma - and IFN-gamma + CD8 + T cells after anti-CD3/CD28 stimulation (3 days; eight biological replicates). (C and D) LC3 staining on GzB and perf-expressing CD8 + T cells ex vivo (C) and after anti-CD3/CD28 stimulation (D) (3 days; eight biological replicates; box whisker, Tukey). (E) Example mitochondrial staining of CD8 + T cells in blood (PBMCs; black) and liver (IHLs; red) and summary data for the ex vivo percentage of total CD8 + T cells with depolarized mitochondria (mitoTracker deep red [MtDR] lo ; see also Figure S4 ; PBMCs, 10; and IHLs, 15 biological replicates). (F) Ex vivo percentage of CD8 + T RM cell subsets in the liver with depolarized mitochondria (14 biological replicates; box whisker, Tukey; outliers shown as dots). (G) The percentage of total CD8 + T cells or CD8 + T RM cell subsets with depolarized mitochondria after overnight culture of IHLs with DMSO (untreated), MRT68921 dihydrochloride (10 muM), bafA1 (0.1 muM), or reagent A (chloroquine diphosphate, 1:1000, FlowCellect LC3 kit; 13-15 biological replicates). Cells were treated with bafA1 (A-D). Bars at mea
- Conjugate
- Green dye
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
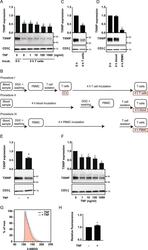
- Figure 1 TNF induces TXNIP down-regulation and increased glucose up-take in T cells. ( A ) Representative Western blot (lower panel) and quantification (upper panel) of TXNIP with CD3zeta as loading control from T cells lysed immediately after isolation (0 h, Procedure I, B ) and after 4 hours of incubation (4 h T cells, Procedure I, B ) with TNF (0-1000 ng/ml) as indicated. ( B ) Overview of the different T cell isolation and incubation procedures used. The text in the red frames gives the nomenclature for the T cell lysates analyzed and shown in the figures. ( C ) Representative Western blot (lower panel) and quantification (upper panel) of TXNIP with CD3zeta as loading control from T cells incubated for 0 hours (0 h, Procedure I, B ) and 4 hours (4 h T cells, Procedure I, B ) before lysis. ( D ) Representative Western blot (lower panel) and quantification (upper panel) of TXNIP with CD3zeta as loading control from T cells immediately isolated and lysed (0 h, Procedure I, B ), isolated and lysed after incubation of blood samples for 4 hours (4 h blood, Procedure II, B ), and isolated and lysed after incubation of PBMC for 4 hours (4 h PBMC, Procedure III, B ). ( E ) Representative Western blot (lower panel) and quantification (upper panel) of TXNIP with CD3zeta as loading control from T cells isolated from blood samples incubated for 4 hours (4 h blood, Procedure II, B ) in the absence (-) or presence (+) of TNF (10 ng/ml). ( F ) Representative Western blot (lower panel) an
- Conjugate
- Green dye
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
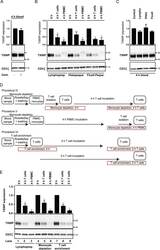
- Figure 2 DGC induces TXNIP down-regulation in T cells. ( A ) Representative Western blot (lower panel) and quantification (upper panel) of TXNIP with CD3zeta as loading control from T cells isolated from blood samples incubated for 4 hours (4 h blood, Procedure II, Fig. 1B ) after no treatment (-) or centrifugation (+). ( B ) Representative Western blot (lower panel) and quantification (upper panel) of TXNIP with CD3zeta as loading control from T cells immediately isolated and lysed (0 h, Procedure I, Fig. 1B ), isolated and incubated for 4 hours before lysis (4 h T cells, Procedure I, Fig. 1B ), and isolated and lysed after incubation of PBMC for 4 hours (4 h PBMC, Procedure III, Fig. 1B ) after DGC on either Lymphoprep, Histopaque or Ficoll-Paque as indicated. ( C ) Representative Western blot (lower panel) and quantification (upper panel) of TXNIP with CD3zeta as loading control from T cells isolated from blood samples incubated for 4 hours (4 h blood, Procedure II, Fig. 1B ) with no addition (Control) or with Lymphoprep, Histopaque or Ficoll-Paque as indicated. ( D ) Overview of different T cell isolation and incubation procedures used. The text in the red frames gives the procedure used and the nomenclature for the T cell lysates analyzed and shown in ( E) lanes 4-9. ( E ) Representative Western blot (lower panel) and quantification (upper panel) of TXNIP with CD3zeta as loading control from T cells immediately isolated and lysed (lane 1: 0 h, Procedure I, Fig. 1B ; lane
- Conjugate
- Green dye
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
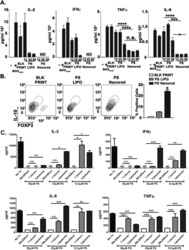
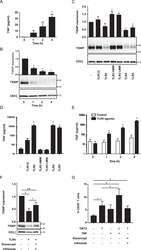
- Figure 3 DGC and TLR agonists induce TNF production and TXNIP down-regulation in T cells. ( A ) TNF in the supernatant of PBMC incubated for 0 to 4 hours (mean + SEM, n = 3). ( B ) Representative Western blot (lower panel) and quantification (upper panel) of TXNIP with CD3zeta as loading control from T cells isolated from PBMC incubated for 0 to 4 hours. ( C ) Representative Western blot (lower panel) and quantification (upper panel) of TXNIP with CD3zeta as loading control from T cells isolated from untreated blood and blood treated with TLR1-5 ligands (TLR1/2, 500 ng/ml PamCSK4; TLR2, 1 x 10 8 heat killed Listeria monocytogenes /ml; TLR3 HMW, 5 ug/ml Poly(I:C) high molecular weight; TLR3 LMW, 5 ug/ml Poly(I:C) low molecular weight; TLR4, 500 ng/ml E. coli K12 LPS; TLR5, 500 ng/ml S. typhimurium Flagellin) for 4 hours (4 h blood, Procedure II, Fig. 1B ) as indicated. ( D ) TNF in the supernatant of blood incubated with the TLR1-5 ligands as above for 4 hours (4 h blood, Procedure II, Fig. 1B ) as indicated. ( E ) TNF in the supernatant of blood incubated for 0 to 4 hours without (white columns) or with the TLR4 ligand (50 ng/ml) (black columns). ( F ) Representative Western blot (lower panel) and quantification (upper panel) of TXNIP with CD3zeta as loading control from T cells isolated from untreated blood and blood treated with the TLR4 ligand (50 ng/ml) in the absence or presence of the TNF inhibitors etanercept (10 ug/ml) and infliximab (10 ug/ml) for 4 hours (4 h blood,
- Conjugate
- Green dye
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
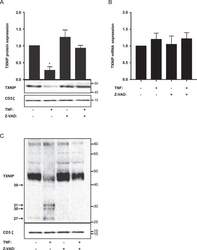
- Figure 4 TNF induces TXNIP down-regulation by a caspase-dependent mechanism. ( A ) Representative Western blot (lower panel) and quantification (upper panel) of TXNIP with CD3zeta as loading control from T cells isolated from untreated blood and blood treated with TNF (10 ng/ml) and the caspase inhibitor Z-VAD-FMK (Z-VAD) (10 ug/ml) for 4 hours (4 h blood, Procedure II, Fig. 1B ) as indicated. The Western blot is representative for Western blots obtained from 3 different biological experiments and the quantification shows the mean + SEM of the band densities of TXNIP from Western blots obtained from 3 different biological experiments. ( B ) Relative TXNIP mRNA expression in T cells isolated from untreated blood and blood treated with TNF (10 ng/ml) and the caspase inhibitor Z-VAD-FMK (Z-VAD) (10 ug/ml) for 4 hours (4 h blood, Procedure II, Fig. 1B ) as indicated (mean + SEM, n = 3). ( C ) Overexposed Western blot of TXNIP with CD3zeta as loading control from T cells isolated from untreated blood and blood treated with TNF (10 ng/ml) and Z-VAD-FMK (Z-VAD) (10 ug/ml) for 4 hours (4 h blood, Procedure II, Fig. 1B ) as indicated. The arrows indicate putative TXNIP fragments from caspase-mediated cleavage of TXNIP and their molecular weight in kDa. ( A,C ) The positions of the relevant molecular weight markers and their molecular weight in kDa are given to the right of each Western blot.
- Conjugate
- Green dye
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- Figure 1 Cdc42 and GRAF1 regulate the formation of tubular structures in activated T cells. ( A ) Jurkat T cells expressing GFP-GRAF1 (left) or GFP-GRAF1-BAR-PH (right) were activated on coverslips coated with functional antibodies against CD3epsilon and CD28 for 10 min and fixed with paraformaldehyde (PFA). Images are maximum intensity projection of confocal z-stacks ( B ) Left: Maximum intensity projection of a Jurkat T cell expressing GFP-GRAF1 and myc-Cdc42-Q61L and activated as in ( A ). Right: the same cell shown in a 3D reconstruction. ( C ) Blinded quantification of the percentage of cells expressing GFP-GRAF1 and mCherry-tagged Cdc42 variants as indicated and having at least one GRAF1-positive tubule. Each data point represents the mean of an individual experiment, 75-136 cells per experiment. Small horizontal lines indicate mean (+-SEM), * p < 0.05; *** p < 0.001; unpaired, two-tailed student's t-test. ( D ) Jurkat T cells expressing GFP-Cdc42-Q61L, activated and fixed as in ( A ) and stained with an antibody against endogenous GRAF1. Arrows indicate GRAF1-positive tubules. Images representative of at least three independent experiments. Scale bar: 5 mum.
- Conjugate
- Green dye
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
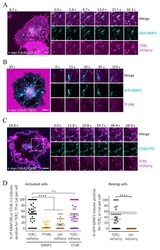
- Figure 2 GRAF1- and CTxB-positive tubules contain TCRzeta but not transferrin. ( A ) Representative time series of Jurkat T cells expressing myc-Cdc42-Q61L, GFP-GRAF1, and TCRzeta-mCherry, and activated on coverslips coated with functional antibodies against CD3epsilon and CD28 and imaged live at 37@C. ( B , C ) Jurkat T cells expressing myc-Cdc42-Q61L and GFP-GRAF1 ( B ) or TCRzeta-mCherry ( C ) were incubated with Alexa546-Tf ( B ) or CTxB-FITC ( C ) for 10 min and imaged live at 37 degC. ( D ) Quantification of the cells as shown in ( A ), ( B ), and ( D ). Percentage of GRAF1- or CTxB-positive tubules that were also positive for TCRzeta-mCherry, Alexa546-Tf, or Lat-mCherry in activated (left, anti-CD3epsilon and anti-CD28) or resting (right, Poly-L-Lysine) cells. On the right, the dotted horizontal line represents the values for TCRzeta in activated cells. Data from at least 45 cells in at least 3 independent experiments. Each data point represents a cell. Small horizontal lines indicate mean (+-SEM), **** p < 0.0001, unpaired, n.s. not significant, two-tailed student's t-test. There were 11 outliers removed for Tf, 8 outliers were removed for Lat-mCherry (resting) using the robust regression and outlier removal (ROUT) method (Q = 0.1%). Scale bar: 5 mum.
- Conjugate
- Green dye
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
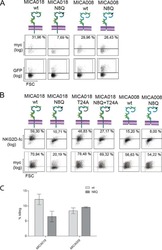
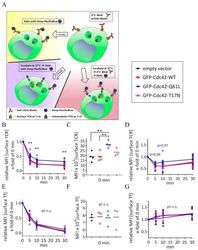
- Figure 4 Expression of Cdc42-Q61L selectively impairs internalization of the TCR-CD3 complex. ( A ) Schematic of the flow-cytometry based internalization assay; cells are labelled at 4 degC with a functional biotinylated anti-CD3epsilon and either (yellow) directly stained with Pacific Blue-streptavidin to measure surface expression of TCR-CD3 complex in resting cells, (blue) activated by incubation at 37 degC and stained with Pacific Blue-streptavidin to detect remaining TCR-CD3 at the cell surface after activation-induced internalization, or (red) activated by incubation at 37 degC, re-labelled with biotinylated anti-CD3epsilon and stained with Pacific Blue-streptavidin to detect total surface TCR-CD3 in activated cells. ( B ) Remaining TCR-CD3 at the cell surface detected by an antibody against CD3epsilon (clone OKT3) after activation-induced internalization in cells expressing an empty vector, GFP-WT-Cdc42, GFP-Cdc42-Q61L, or GFP-Cdc42-T17N. ( C ) Surface expression of TCR-CD3 complex in cells expressing the same constructs as in ( B ). ( D ) Total surface TCR-CD3 in activated cells transfected as in ( B ). ( E ) Internalization of biotinylated Tf detected with Pacific Blue-streptavidin after incubation at 37 degC as described for anti-CD3epsilon in ( A ), in cells activated by soluble anti-CD3epsilon and expressing the same constructs as in ( B ). ( F ) Surface expression of Tf in cells expressing the same constructs as in ( B ). ( G ) Total Tf at the cell surface detect
- Conjugate
- Green dye
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
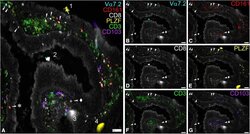
- Figure 7 MAIT Cells Can Be Found Close to and within the Colonic Epithelium (A-G) Representative images showing the expression of Va7.2, CD161, CD8, PLZF, CD3, and CD103 in the lamina propria and the epithelium of fixed samples of colonic polyp tissue. Samples were mounted on cytometer chips and iteratively stained with sets of three directly fluorochrome-conjugated antibodies as described in the methods section. Depicted are a merged picture (A) and all the individual stains for Va7.2 (B), CD161(C), CD8 (D), PLZF (E), CD3 (F), and CD103 (G). White arrows mark cells showing co-expression of Va7.2, CD161, PLZF, and CD3 that were defined as MAIT cells here. Note that while CD8 was co-expressed in most of them, CD8- MAITs (arrow + asterisk) could also be found. In contrast, CD103 was rarely co-expressed on MAITs (arrow + diamond). During the iterative staining process dust particles and other detritus can be picked up by the solution flowing over the tissue creating autofluorescent artifacts (1-4). While some of these get washed away after completion of the staining cycle (1, 4), others present during multiple imaging rounds (2, 3). Scale bars, 50mum.
- Conjugate
- Green dye
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
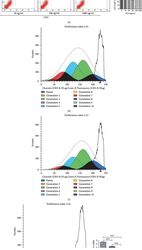
- Figure 3 Flow cytometric analysis of the proportion and proliferation ability of CD3 + CD8 + T lymphocytes. (a) Proportion of CD3 + CD8 + T lymphocytes. (b) Blank group. (c) Control group, lymphocytes cocultured with A-MSCs stimulated at 0 ng/mL PLP. (d) Treated group, lymphocytes cocultured with A-MSCs stimulated at 50 ng/mL PLP.
- Conjugate
- Green dye
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
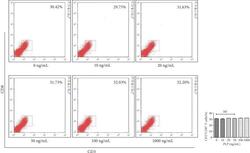
- Figure 13 Flow cytometric analysis of the proportion of CD3 + CD8 + T lymphocytes after A-MSCs coculture with TLR3 and TLR4 inhibition. There was no significant difference in the proportion of CD3 + CD8 + T lymphocytes between concentrations.
- Conjugate
- Green dye
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
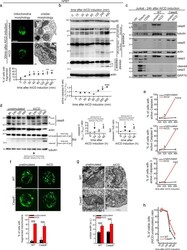
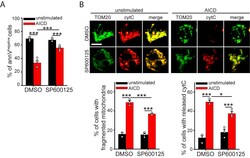
- Fig. 1 Caspase-8-dependent extrinsic cell death pathway is not involved in early AICD events. a hPBT cells representative z-stacks reconstructions of TOM20 staining (left panel) and representative electron micrographs (right panel), 30 min after AICD induction. Quantification of the cells with fragmented mitochondria at the indicated time after AICD induction is reported in the graph below ( n = 3). b Expression levels of the indicated proteins evaluated by western blot in hPBT cells at the indicated times after AICD induction. Relative quantification of the active caspase-8 (measured as the ratio between active p41/43, p30 and p18 isoforms compared with inactive p53/55 isoform) is reported in the graph below ( n = 3). Asterisks (*) indicate unspecific bands. c Expression levels of the indicated proteins evaluated by western blot in WT and caspase-8 -/- Jurkat cells 24 h after AICD induction, i.e., the starting point of mitochondria alteration detection, as shown in Fig. S1A (one experiment representative of three independent experiments). d Expression levels of the indicated proteins evaluated by western blot in Jurkat cells at the indicated time points after AICD induction. Relative quantifications of the active caspase-8 (measured as the ratio between active p41/43 compared with inactive p53/55 isoform) and active Bid (measured as cleaved/full length ratio) are reported in the graphs on the right (caspase-8: n = 3; Bid: n = 4). e Percentage of control or AICD-stimulated Ju
- Conjugate
- Green dye
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
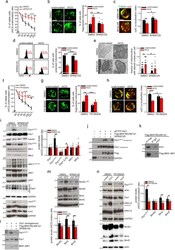
- Fig. 2 Treatment with SP600125 and FR180204 inhibitors prevents mitochondrial alterations upon AICD in hPBT cells. a-e hPBT cells have been stimulated for AICD induction in presence or not of SP600125. In a is reported the relative viability (AICD:unstimulated ratio) assessed by flow cytometry (percentage of annexinV negative 7AAD negative cells assessed by flow cytometry) ( n = 5). In b are reported representative confocal z-stack reconstructions of mitochondria (TOM20) in hPBT cells 2 h after AICD induction ( n = 4). In c are reported representative images of viable and dying hPBT cells showing mitochondria (TOM20 in green, confocal z-stack reconstruction) and cytochrome-C (red) localization, 4 h after AICD induction. Quantification of the cyt-C co-localization index with mitochondria (see Methods for details) in presence or not of SP600125 4 h after AICD induction is reported in the graph on the right ( n = 4). In d is reported the TMRE profile by flow cytometry 2 h after AICD induction. Percentages of cells with polarized mitochondria (indicated with red bars on plots) are quantified in the graph on the right ( n = 3). In e are reported representative electron micrographs of hPBT cells 4 h after AICD induction. Quantification of the fraction of mitochondria with altered cristae per cell (range 0-1) in each condition is reported in the graph on the right (at least 30 cells per condition from n = 3 independent experiments). f Relative viability (AICD:unstimulated ratio) of
- Conjugate
- Green dye
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
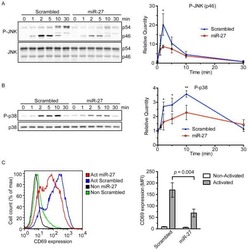
- Fig. 3 Dissection of JNK1 and ERK1/2 roles during AICD progression. Jurkat cells have been transfected with the indicated siRNA or with Flag-MKK7B2JNK1a1 plasmid (only in panel h ). 24 h after transfection, Jurkat cells have been simulated for AICD. a Representative western blot showing transfection efficiency in Jurkat cells ( n = 4). b Relative viability (AICD:unstimulated ratio) of transfected Jurkat cells stimulated for AICD induction (percentage of annexinV neg 7AAD neg cells assessed by flow cytometry) ( n = 6). c On the left: representative images of mitochondria morphology (TOM20, confocal z-stack reconstructions) 24 h after AICD induction in Jurkat cells transfected with siNEG. On the right: quantification of the percentage of Jurkat cells with fragmented mitochondria upon AICD induction and after transfection with the indicated siRNAs ( n = 5). d Representative images of viable and dying Jurkat cells transfected with siNEG showing mitochondria (TOM20 in green, confocal z-stack reconstructions) and cytochrome-C (red) localization 28 h after AICD induction. Quantification of the percentage of cells with cyt-C released from mitochondria upon AICD induction and after transfection with the indicated siRNAs is reported in the graph on the right ( n = 5). e-j Expression levels of the indicated (phospho)-proteins in transfected Jurkat cells. In e , f , i , j cells have been analyzed 24 h after AICD induction. In g , h cells have been analyzed 24 h after transfection without
- Conjugate
- Green dye
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
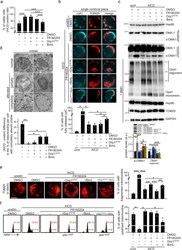
- Fig. 4 Drp1 and Bim mediate AICD progression downstream of JNK1 and ERK1/2. Jurkat cells have been transfected with the indicated plasmids (pEYFP-Drp1-S616E and Flag-Bim-L). 24 h after transfection, Jurkat cells have been stimulated for AICD in presence or not of FR180204, as indicated. a Relative viability (AICD:unstimulated ratio) of YFP-Drp1 and Bim-L transfected Jurkat cells after 32 h from AICD induction (percentage of annexinV neg 7AAD neg cells assessed by flow cytometry). Cells have been gated on YFP-positive cells when transfected with YFP-Drp1-expressing plasmid ( n = 5). b Representative single z-stack images of mitochondria morphology (TOM20) and cytochrome-C (light blue) 28 h after AICD induction in Jurkat cells transfected with YFP-Drp1 and Bim-L. Quantification of the percentage of cells with fully released cyt-C is reported in the graph below ( n = 6). c Expression levels of the indicated (phospho)-proteins 28 h after AICD induction in Jurkat cells transfected with YFP-Drp1 and Bim-L. Opa1, Hsp90, TOM20 and GAPDH have been evaluated in samples treated with BMH (+BMH) to isolate Opa1 oligomers, which are indicated with an asterisk (*). Quantifications of the amount of s-OMA1 and Opa1 oligomers are reported in the graph below ( n = 5). One-way ANOVA have been performed to analyze s-OMA1 levels. One-way ANOVA on repeated measurements have been performed to analyze Opa-1 oligomers. d Representative electron micrographs of Jurkat cells transfected with YFP-Drp1 and
- Conjugate
- Green dye
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- Fig. 5 Bim is required for AICD progression in normal and T-ALL cells. a Expression levels of the indicated proteins in naive hPBT stimulated once, for 3 h (I stim), or twice (II stim) with anti-CD3 (plate-coated) and anti-CD28 antibodies. For the second stimulation, cells have been pre-activated with anti-CD3 (plate-coated) and anti-CD28 antibodies for 48 h and expanded for 6 days in IL2-containing medium. The quantification of the ratio between Bim-L/S protein isoforms and Bcl2 protein is reported for each condition in the graph on the right ( n = 3). b-f AICD has been induced in Jurkat cells silenced for Bim (checked by western blot in B, n = 3). Relative viability (AICD:unstimulated ratio) at 30 h upon AICD induction is indicated in C ( n = 6), western blot analysis of the Opa1 oligomers at 30 h in d ( n = 6; relative quantification on the right), TMRE staining at 26 h by flow cytometry in e ( n = 3; relative quantification on the right), and immunofluorescence analysis of mitochondria morphology (TOM20) and cyt-C localization at 30 h, in single confocal plane, in f ( n = 3; relative quantifications of percentage of cells with fragmented mitochondria, or released cyt-C, on the right). g Expression levels of the indicated genes from T-ALL and healthy bone marrows (HBM) obtained from Leukemia MILE Dataset and analyzed through BloodSpot (HBM: n = 73; T-ALL: n = 174). h Western blot analysis of Bim levels in healthy human Teff cells (stimulated once, and expanded 6 days with
- Conjugate
- Green dye
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
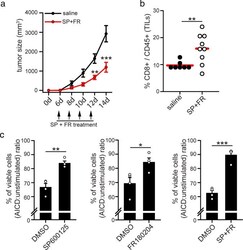
- Fig. 6 Drugs targeting MAPK proteins reduces tumor growth in vivo and increases T cell survival. a, b 5*10 5 MC38 tumor cells have been inoculated s.c. into the right flank of WT mice. On the indicated days, mice have been injected i.p with SP600125 and FR18024, and tumor volume assessed ( a , n = 7 saline; n = 9 SP + FR-treated). After 14 days, TILs have been isolated from tumor mass and the percentage of CD8 + T cells among all CD45 + TILs quantified by flow cytometry ( b , n = 7 saline; n = 9 SP + FR-treated). c T cells have been isolated from the spleen of 14 days-old MC38-derived tumor-bearing WT mice and cultured in vitro for 10 days in presence of UV-irradiated MC38 cells, plus IL2 and IL15, to expand tumor-reactive T cells. After 10 days, CD8 + T cells have been magnetically purified and re-stimulated for the indicated time with plate-coated anti-CD3 antibodies to induce AICD in presence or not of SP600125 and/or FR180204. Graphs show the relative viabilities (AICD:unstimulated ratio) of murine T cells stimulated 6 h for AICD (percentage of annexinV neg 7AAD neg cells assessed by flow cytometry) ( n = 4). Data are shown as mean +- SEM. Significance is indicated as follows: * p < 0.05; *** p < 0.001.
- Conjugate
- Green dye
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
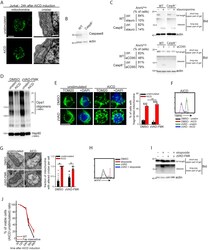
- Supplemental Figure 1
- Conjugate
- Green dye
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- Supplemental Figure 2
- Conjugate
- Green dye
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
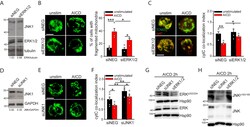
- Supplemental Figure 3
- Conjugate
- Green dye
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
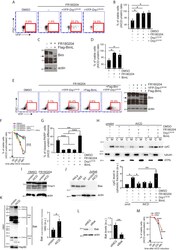
- Supplemental Figure 4
- Conjugate
- Green dye
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- Supplemental Figure 5
- Conjugate
- Green dye
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
- Supplemental Figure 6
- Conjugate
- Green dye
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
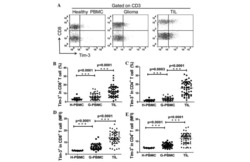
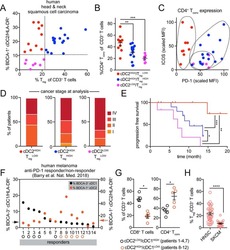
- 1 Figure Characterization of TILs and PBMCs from colorectal and breast cancer patients. (A) Expression of PD-1 in CD8 + TILs and PBMCs from colorectal cancers and breast cancers. (Left) Representative flow cytometry data. TILs and PBMCs were stained with CD3, CD8, and PD-1 antibodies, and PD-1 expression in CD3 + CD8 + T cells was analyzed by flow cytometry. (Right) Percentages of PD-1 + and PD-1 - cells in CD8 + TILs and CD8 + PBMCs in colorectal cancer (CC) patients and breast cancer (BC) patients. The percentage of PD-1 + and PD-1 - cells in TILs were analyzed by flow cytometry with 34 CC patient's and 10 BC patient's samples. The percentages of PD-1 + and PD-1 - cells in PBMCs were analyzed with samples from 10 CC patients and 6 BC patients. Because of limited cell number, each patient's sample was analyzed once per experiment and mean +- SD is calculated. (B) The TCRs repertoire of PD-1 + CD8 + TILs from CC and BC patients. (Left) Representative TCR repertoires of PD-1 + and PD-1 - TILs from a CC patient and a BC patient. PD-1 + or PD-1 - CD8 + TILs were single cell sorted, and then TCR cDNA was amplified with single cell RT-PCR and sequenced directly. Each pie slice in gray indicates a T-cell population in which T cells expressed the same clonotypic TCRbeta. A white pie slice indicates a T cell population in which each T cell expressed a unique TCR. The total number of analyzed cells is shown in the center of each pie chart. (Right) Clonally expanded populations in CD8
- Conjugate
- Green dye
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
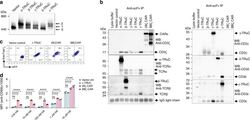
- Fig. 2 Incorporation of TRuCs in TCR complexes and functional analysis of endogenous TCR in Jurkat cells stably transduced with empty control vector or vectors encoding the indicated CAR or TRuC transgenes. The transgene expression was linked to GFP via a T2A sequence. a Cells were lysed and TCR complexes separated by Blue Native PAGE. After Western blot TCR complexes were stained using an anti-CD3zeta antibody. ""0"" denotes the natural TCR complex, ""1"" and ""2"" denote TCR complexes with one or two TRuCs, respectively. b Complex formation of TRuC variants with TCR subunits. Upon cell lysis, TRuCs and CARs were immunopurified using the anti-F(ab') 2 antibody, and then separated by reducing SDS-PAGE. As a control, the procedure was also applied to the lysis buffer alone. (Co)-purified proteins were detected using the described antibodies by western blot. Data in a and b show representative results of three experiments. c Transduced cells were stained with an anti-CD3epsilon antibody and analyzed by flow cytometry. d The effect of epsilon-TRuC on activation of an influenza HA-specific TCR. Jurkat T cells expressing the HA1.7 TCR (Jurkat cell line CH7C17) were transduced with a control vector (black) or vectors encoding for the epsilon-TRuC (light blue), the 28zeta (red) or BBzeta CAR (purple). Cells were co-cultured with DapDR1-ICAM1 antigen-presenting cells loaded with different amounts of the HA306-318 peptide. After 6 h, upregulation of the activation marker CD69 was quan
- Conjugate
- Green dye
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
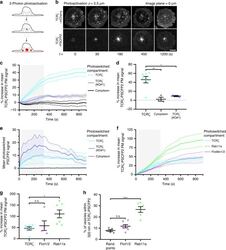
- Fig. 1 Flotillins mediate sorting of internalised TCRzeta into Rab5 and Rab11a endosomes. a Schematic depicting photoactivation to investigate sorting. Top: sorting of photoactivated endosomes (red) into the Rab compartment (green, yellow = merged), bottom: photoactivated endosomes not sorting into the Rab compartment. Dashed lines: quantified compartment mask. b Representative images of photoactivated (dashed region) TCRzeta-PAmCherry sorting into EGFP-Rab4 in WT (left) and FlotKO (right) Jurkat T cells. Cells photoactivated for five frames with 2.5 s intervals, then imaged every 2.5 s for 150 frames. c % endosomal TCRzeta-PAmCherry intensity in Rab4 in WT or FlotKO cells. Dashed line represents timepoint where TCRzeta-PAmCherry intensity significantly diverges between WT and FlotKO. d Percentage reduction in endosomal TCRzeta-PAmCherry intensity in Rab4 from the 50 to 55 s timepoint. Fitted linear regression line is bolded. n.s. indicates slopes are not significantly different. e Representative images of photoactivated (dashed regions) TCRzeta-PAmCherry sorting into EGFP-Rab5 in WT (left) and FlotKO (right) cells. Cells imaged as in b . f Percentage endosomal TCRzeta-PAmCherry in Rab5 in WT or FlotKO cells over time. g Percentage reduction in endosomal TCRzeta-PAmCherry intensity in Rab5 from 50 to 55 s. Fitted linear regression line is bolded. ***Indicates slopes are highly significantly different. h Representative images of photoactivated (dashed region) TCRzeta-PAmCherry
- Conjugate
- Green dye
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
- Fig. 2 TCRzeta recycles from an intracellular compartment labelled by flotillins and Rab11a. a Schematic depicting two-photon photoactivation to measure recycling from intracellular compartments. b Representative confocal images of TCRzeta-PSCFP2 photoactivated with 800 nm light from a two-photon laser, 2.5 um within the cell at the cell centre-localised TCRzeta-mCherry compartment displaying recycling to the plasma membrane over time. Dashed circle = photoactivation region. c % increase in photoactivated TCRzeta-PSCFP2 signal over time at the plasma membrane when activated from cell centre compartments labelled by TCRzeta-mCherry in WT or FlotKO Jurkat T cells or cytoplasmic photoactivation. Greyed region = time of photoactivation. d Mean TCRzeta-PSCFP2 signal from 900 to 1200 s timepoints when photoactivated from the indicated compartments. e Mean intenisty of photoactivated TCRzeta-PSCFP2 2 um within the cell from TCRzeta-mCherry in WT or FlotKO cells. Greyed region = time of photoactivation. f % increase in photoactivated TCRzeta-PSCFP2 signal over time at the plasma membrane when activated from cell centre compartments labelled by mCherry-Rab11a, flotillin1/2-mCherry or TCRzeta-mCherry in WT Jurkat T cells. Grey box = time of photoactivation. Greyed region = time of photoactivation. g Mean TCRzeta-PSCFP2 signal from 900 to 1200 s timepoints when photoactivated from the indicated intracellular compartments. h Percentage of photoactivated TCRzeta-PSCFP2 vesicles within 320
- Conjugate
- Green dye
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
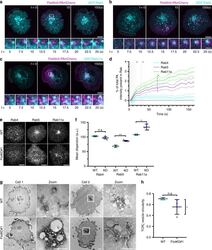
- Fig. 3 Flotillins regulate TCRzeta sorting into Rab11a vesicles for fusion with the plasma membrane. a Representative images of a TCRzeta-GFP vesicle fusion event. Zoomed images below represent the indicated fusion event (white box) at 200 ms intervals. Main image scale bar = 5 um, inset scale bar = 1 um. b TCRzeta-GFP vesicle fusion events per minute in WT and two FlotKO Jurkat T cell lines. c Representative images of TCRzeta-GFP (cyan) vesicle fusion events with either flotillin-1 and flotillin-2 mCherry (magenta, top two panels) or mCherry-Rab11a (magenta, bottom panel). Scale bar = 0.5 um. d Percentage of TCRzeta-GFP fusion events containing either flotillin1/2-mCherry or mCherry-Rab11a. e Representative images displaying a GFP-Rab11a vesicle fusion event. Zoomed images below represent the indicated fusion event (white box) at 200 ms intervals. Main image scale bar = 5 um, inset scale bar = 1 um. f GFP-Rab11a vesicle fusion events per minute in WT and two FlotKO Jurkat T cell lines. g Representative images of TCRzeta-GFP (cyan) vesicle fusion events with mCherry-Rab11a (magenta) in WT Jurkat T cells (top) or fusion with (middle) or without (bottom) mCherry-Rab11a in FlotKO Jurkat T cells. Scale bar = 0.5 um. h Percentage of TCRzeta-GFP fusion events containing mCherry-Rab11a in WT and two FlotKO Jurkat T cell lines. All images captured on a Zeiss ELYRA TIRF microscope. Data points indicate means of independent experiments, error bars indicate mean +- SEM. n.s. = not signi
- Conjugate
- Green dye
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
- Fig. 4 Flotillins rapidly interact with and maintain the organisation of Rab5 and Rab11a-positive endosomes. a Representative images of flotillin-2-PAmCherry photoactivated at the dashed region failing to interact with EGFP-Rab4 endosomes in WT Jurkat T cells. Cells were photoactivated for five frames with 2.5 s intervals, then imaged every 2.5 s for 60 frames. b Representative images of flotillin-2-PAmCherry photoactivated at the dashed region rapidly interacting with EGFP-Rab5 endosomes in WT Jurkat T cells. Cells were photoactivated and imaged as in a . c Representative images of flotillin-2-PAmCherry photoactivated at the dashed region rapidly interacting with EGFP-Rab11 endosomes in WT Jurkat T cells. Cells were photoactivated and imaged as in a . For a - c zoomed images below are indicated by a white box, main image scale bar = 5 um, inset scale bar = 1 um, and flotillin-2-PAmCherry was co-expressed with untagged flotillin-1. All cells imaged on a Zeiss 880 confocal, photoactivation performed at 405 nm. d Quantification of the percentage of photoactivated flotillin-2-PAmCherry present in Rab4, 5 or 11a. e Representative confocal images of endogenous Rab4, Rab5 and Rab11a detected with corresponding antibodies in WT (top) or FlotKO (bottom) Jurkat T cells. Scale bar = 5 um. f Mean fluorescence dispersion of the indicated markers. Data points indicate mean dispersion from n = 3 biologically independent experiments (Fig. 4a-d, Rab4: 10; 8; 5 cells, Rab5: 10; 12; 10 cells,
- Conjugate
- Green dye
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
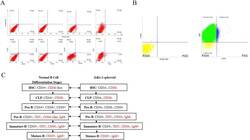
- Figure 2 Immunophenotype analysis of JeKo-1-parental and JeKo-1-spheroid cells. (A) The expression of CD22, CD34, CD10, CD5, CD38 and TDT in CD19 - population from JeKo-1-spheroid cells. (B) The expression of CD19 and IgM in JeKo-1-spheroid total cells. (C) Comparison of marker expression in JeKo-1-spheroid with the normal B cell differentiation stages. Red font indicates the same immunophenotype. Figure 2
- Conjugate
- Green dye
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
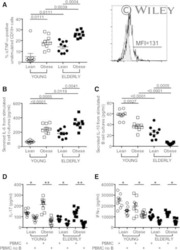
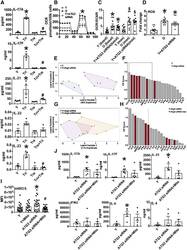
- Figure 2 Metformin Ameliorates OXPHOS and Promotes Non-mitochondrial Glycolysis in CD4 + T Cells from O Subjects (A) OCR in a mito stress test assayed by XF of CD4 + T cells following 40 h alphaCD3/alphaCD28 stimulation +- 100 mum MET as indicated. (B) OCR:ECAR ratio calculated by profiles in (A) and Figure S2 C. (C) Relative lactate production after 40-h stimulation per (A). (D) Proton leak calculated from (A) data. (E) MMP measured with TMRE after stimulation per (A). #p = 0.055 versus O. (F) MMP measured following addition of the mitochondrial uncoupler fluoro-carbonyl cyanide phenylhydrazone (FCCP) to unstimulated CD4 + T cells from Y or O subjects. n = 8-10 (A-E) and 12-13 (F). (G) LDH quantification on western blots. Top: representative blot and bottom averages n = 4-6 of group indicated beneath. * p < 0.05 versus Y, #p < 0.05 versus O. Data shown are mean +- SEM. (H) VIP scores, which rank cytokines as most (leftmost) or least (rightmost) important for differentiating overall cytokine profiles between CD4 + T cells from young subjects stimulated +- the LDH inhibitor OA in an orthagonalized model. A VIP score >1 (bracket) is considered important for differentiating inflammatory profiles between groups. All VIP cytokines indicated also differed in post hoc analyses (p < 0.05). Fold change is compared to Y or Y + FCCP. See also Figure S2 .
- Conjugate
- Green dye
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
- Figure 5 Genetic Inhibition of Autophagy Recapitulates Respiratory Profiles of Cells from O Subjects (A) Th17-associated cytokine production by CD4 + T cells from Y subjects, with cells stimulated 40 h alphaCD3/alphaCD28 in the presence of trimetazidine, a fatty acid oxidation inhibitor; alone; or in combination with trehalose, an autophagy activator. n = 4. * p < 0.05 versus Y by one-way ANOVA. (B) Mito stress test XF profiles from 40 h alphaCD3/alphaCD28-stimulated CD4 + T cells from Y or O subjects as indicated. Autophagy dysfunction was induced in cells from Y subjects using siRNA-mediated ATG3 knockdown. n = 8-12. (C and D) OCR:ECAR ratio (C) and ROS generation (D) measured by DCFDA in CD4 + T cells manipulated as indicated. n = 8-12. * p < 0.05 versus Y by SHORE ( Nicholas et al., 2017 ) (B) or one-way ANOVA (C and D). * p < 0.05 versus Y (basal), $p < 0.05 versus Y (max) (C). (E and G) PLSDA analysis differentiated combinatorial ""inflammation"" of CD4 + cells from Y subjects (blue), Y with siRNA-induced autophagy dysfunction (purple), or autophagy dysfunction and metformin (met; orange). n = 8-9. (F and H) VIP scores rank cytokines important for differentiating data clouds in (E) and (G). Comparison of Figure 5 F with 1D highlights profiles that differentiate Y from either O or Y + ATG3 siRNA conditions. n = 8-9. (I) Mitochondrial ROS generation measured by MitoSOX in CD4 + T cells manipulated as indicated; 3 cells/field and 4 fields/slide were imaged using 40x in Zei
- Conjugate
- Green dye
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
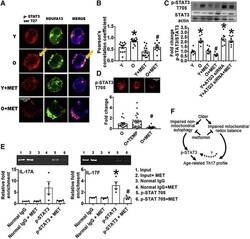
- Figure 6 Aging Promotes STAT3 Activation, Mitochondrial Localization, and IL17A/F Promoter Binding (A and B) Quantification of STAT3 ser727 and the mitochondrial complex 1 subunit NDUFA13 localization in CD4 + T cells stimulated for 40 h with alphaCD3/alphaCD28 +- metformin as indicated. n = 4 with multiple field readings shown per N. (C) Phospho (p)-STAT3 T705 expression assayed on western blots as indicated on x axis. n = 6-8. (D) p-STAT3 T705 expression relative to total STAT3 in O cells in the presence of TEMP or MET. n = 4-5. * p < 0.05 versus Y or control siRNA or #p < 0.05 versus O. Fold change is compared with Y or O. (A and D) 3 cells/field and 3-5 fields/slide were imaged using 63x oil immersion in Zeiss microscope. Average fluorescence/field is reported. (E) Chromatin immunoprecipitation assay showing fold-enrichment of p-STAT3 705 on (left) IL-17A or (right) IL-17F promoters. n = 4. All bar graphs show mean +- SEM. (F) Model for parallel metformin-sensitive pathways that drive T cell inflammaging. Activated T cells from O subjects displayed blunted autophagy, which was largely independent of changes in mitochondrial bioenergetics/excess ROS that promote STAT3-mediated activation of a CD4 + T cell inflammaging profile.
- Conjugate
- Green dye
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
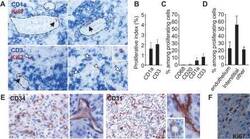
- Figure 1 Proliferating Cells in LCH Granuloma are Mostly Endothelial Cells, Fibroblasts, and T Cells Paraffin-embedded and frozen sections were stained with antibodies against Ki-67 (which label proliferating cells), CD1a (LCs), CD3 (T cells), CD20 (B cells), CD68, CD31, and CD34 (endothelial cells). (A) Double immunostaining of paraffin-embedded section from LCH eosinophilic granulomas with anti-Ki-67 Ab, (brown nuclear staining) and with anti-CD1a Ab (upper images, blue staining) or anti-CD3 Ab (lower images, blue staining). Open arrowheads indicate double-stained cells, black arrowheads indicate Ki-67 + cells with an endothelial morphology. (B) Histogram represents percentage of CD1a + cells and of CD3 + cells labeled with Ki-67 ( n = 15). (C) Histogram represents percentage of proliferating cells (Ki-67 + ) that express CD1a, CD3, CD20, or CD68 ( n = 15). (D) Histogram represents percentage of proliferating cells (Ki-67 + ) that are endothelial cells, interstitial cells (fibroblasts), and other types based on morphological examination. (E) Immunolabeling of blood vessels on paraffin-embedded section from LCH eosinophilic granulomas with CD34 (left) and CD31 (right) antibodies. (F) Proliferating Ki-67 + cells (brown nuclear staining) with a fibroblast-cell morphology in an eosinophilic granuloma.
- Conjugate
- Green dye
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
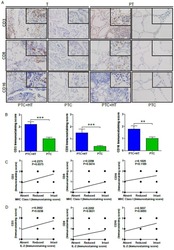
- Figure 3 UP-regulated CD3, CD8 and CD16 expression in PTC+HT tissues. A , IHC analysis of CD3, CD8, and CD16 expression in 53 PTC+HT, 91 PTC tumor (T) and their adjacent para-non-tumor tissue (PT). B , Immunostaining scores ofCD3, CD8, and CD16 expression in tumor tissues. C , The correlation between the immunostaining scores of HLA class I molecule and CD3, CD8 and CD16 expression. D , The correlation between the immunostaining scores of IL-2 and CD3, CD8 and CD16 expression. Data are representative images (magnification x 200 with a 400 inserter) or expressed as the mean +- SD of each group. * P
- Conjugate
- Green dye
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
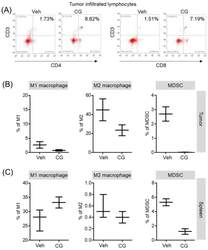
- Figure 6 The proportion of MDSCs and M1/M2 macrophage were changed by CG-745 treatment in Hepa1-6 syngeneic mouse model: (A-C) Hepa1-6 inoculated C57BL/6 mice were treated with vehicle or 15 mg/kg/day of CG (CG-745) by intraperitoneally for 5 days/week for 3 weeks. Tumor (A, B) and spleen (C) were harvested from each individual mouse. Cells were stained with fluorescence-conjugated antibodies specific to CD3, CD4, CD8, CD25 and Foxp3, and subjected to flow cytometry analysis. The graph shows means with error bars. Error bars represent standard deviations for 3 samples in each group. Results are representative of three separate experiments (A). p < 0.05.
- Conjugate
- Green dye
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
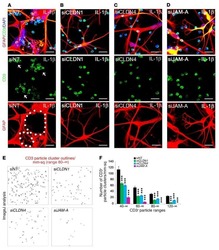
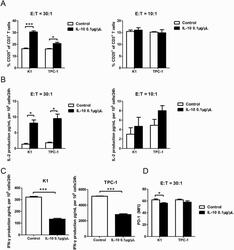
- Figure 4. IL-10 promotes T-cell activation in cocultures of CD3 + CD8 + CD25 + T cells and PTC cell lines. ( A ) Flow cytometry results of the CD25 + fraction of CD3 + T cells in coculture systems containing activated CD3 + CD8 + CD25 + T lymphocytes (effector cells, E) and pretreated PTC cells (K-1 and TPC-1 treated with 0.1 mug/muL of IL-10 for 24 hours; target cells, T) at various E:T ratios (30:1 and 10:1) (* P < 0.05, *** P < 0.001). ( B ) IL-2 production in coculture systems (details described above) measured by ELISA (* P < 0.05). ( C ) IFN-gamma production in coculture systems of CD3 + CD8 + CD25 + T lymphocytes and IL-10-stimulated PTC cells (K-1 and TPC-1) at a 30:1 ratio (*** P < 0.001). ( D ) Mean fluorescence intensity (MFI) of PD-1 expression in coculture systems of activated CD3 + CD8 + CD25 + T lymphocytes and IL-10 pretreated PTC cells at a 30:1 ratio (* P < 0.05). All tests were conducted in triplicate. ELISA, enzyme-linked immunosorbent assay; H, hours; IFN, interferon; IL, interleukin; PD-1, programmed death-1; PTC, papillary thyroid cancer.
- Conjugate
- Green dye
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
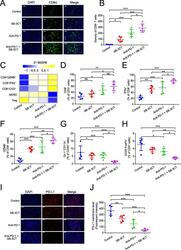
- Fig. 3 Immune features in tumors for B16F10 xenograft mouse model with SB-3CT treatment. a Fluorescence expression and b quantification of CD8 + T cells. c Heatmap of Z -score normalized percentage of immune cell populations ( d - h ) in TILs for B16F10 tumor-bearing mice treated with anti-PD-1 and SB-3CT in combination or alone. d - h In the implanted B16F10 tumors from mice treated with or without SB-3CT and PD-1 blockade, fluorescence-activated cell sorting (FACS) was used to measure d CD8 + in CD3 + T cells, e CD8 + IFNgamma + in CD8 + T cells, f CD8 + GZMB + in CD8 + T cells, g Gr-1 + CD11b + MDSCs in CD45 + cells, and h CD25 + FOXP3 + Treg in CD4 + cells. i Fluorescence expression and j quantification of PD-L1 in B16F10 tumor-bearing wild-type C57/BL6 mice treated with isotype, SB-3CT, anti-PD-1, or combination strategy. Sample size is 5 in each cohort. Scale bars, 50 mum. Results are mean +- s.d. ns, p > 0.05,* p < 0.05, ** p < 0.01, and *** p < 0.001, as determined by one-way ANOVA and Dunnett's multiple comparison test
- Conjugate
- Green dye
 Explore
Explore Validate
Validate Learn
Learn Flow cytometry
Flow cytometry