32-4600
antibody from Invitrogen Antibodies
Targeting: SRSF1
ASF, MGC5228, SF2, SF2p33, SFRS1, SRp30a
Antibody data
- Antibody Data
- Antigen structure
- References [27]
- Comments [0]
- Validations
- Western blot [1]
- Immunocytochemistry [1]
- Other assay [19]
Submit
Validation data
Reference
Comment
Report error
- Product number
- 32-4600 - Provider product page

- Provider
- Invitrogen Antibodies
- Product name
- SRSF1 Monoclonal Antibody (103)
- Antibody type
- Monoclonal
- Antigen
- Recombinant full-length protein
- Reactivity
- Human, Mouse, Rat
- Host
- Mouse
- Isotype
- IgG
- Antibody clone number
- 103
- Vial size
- 100 µg
- Concentration
- 0.5 mg/mL
- Storage
- -20°C
Submitted references RNA splicing is a key mediator of tumour cell plasticity and a therapeutic vulnerability in colorectal cancer.
Mechanism of action of hepatitis B virus S antigen transport-inhibiting oligonucleotide polymer, STOPS, molecules.
BRD4 prevents the accumulation of R-loops and protects against transcription-replication collision events and DNA damage.
Splicing events in the control of genome integrity: role of SLU7 and truncated SRSF3 proteins.
SAMMSON fosters cancer cell fitness by concertedly enhancing mitochondrial and cytosolic translation.
5-methylcytosine promotes mRNA export - NSUN2 as the methyltransferase and ALYREF as an m(5)C reader.
Human satellite-III non-coding RNAs modulate heat-shock-induced transcriptional repression.
Comparative expression patterns and diagnostic efficacies of SR splicing factors and HNRNPA1 in gastric and colorectal cancer.
Regulation of gene expression via retrotransposon insertions and the noncoding RNA 4.5S RNAH.
Splicing factor SRSF1 negatively regulates alternative splicing of MDM2 under damage.
The adipogenic transcriptional cofactor ZNF638 interacts with splicing regulators and influences alternative splicing.
Nuclear ARVCF protein binds splicing factors and contributes to the regulation of alternative splicing.
The splicing factor FUBP1 is required for the efficient splicing of oncogene MDM2 pre-mRNA.
A METTL3-METTL14 complex mediates mammalian nuclear RNA N6-adenosine methylation.
RRP1B is a metastasis modifier that regulates the expression of alternative mRNA isoforms through interactions with SRSF1.
ALKBH5 is a mammalian RNA demethylase that impacts RNA metabolism and mouse fertility.
CTD serine-2 plays a critical role in splicing and termination factor recruitment to RNA polymerase II in vivo.
Adaptive alternative splicing correlates with less environmental risk of parkinsonism.
Hypoxia is a modifier of SMN2 splicing and disease severity in a severe SMA mouse model.
Malat1 is not an essential component of nuclear speckles in mice.
Cell-to-cell variability of alternative RNA splicing.
SAM68 regulates neuronal activity-dependent alternative splicing of neurexin-1.
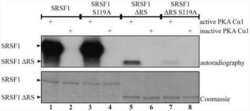
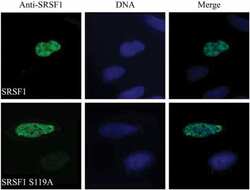
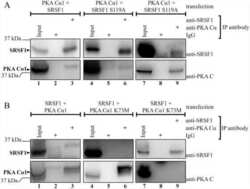
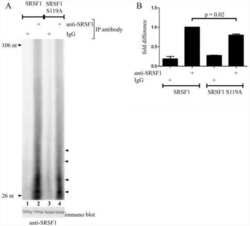
Protein kinase a-dependent phosphorylation of serine 119 in the proto-oncogenic serine/arginine-rich splicing factor 1 modulates its activity as a splicing enhancer protein.
S1-1 nuclear domains: characterization and dynamics as a function of transcriptional activity.
Three-dimensional positioning of genes in mouse cell nuclei.
The switch in alternative splicing of cyclic AMP-response element modulator protein CREM{tau}2{alpha} (activator) to CREM{alpha} (repressor) in human myometrial cells is mediated by SRp40.
F-actin-dependent insolubility of chromatin-modifying components.
Hall AE, Pohl SÖ, Cammareri P, Aitken S, Younger NT, Raponi M, Billard CV, Carrancio AB, Bastem A, Freile P, Haward F, Adams IR, Caceres JF, Preyzner P, von Kriegsheim A, Dunlop MG, Din FV, Myant KB
Nature communications 2022 May 19;13(1):2791
Nature communications 2022 May 19;13(1):2791
Mechanism of action of hepatitis B virus S antigen transport-inhibiting oligonucleotide polymer, STOPS, molecules.
Kao CC, Nie Y, Ren S, De Costa NTTS, Pandey RK, Hong J, Smith DB, Symons JA, Beigelman L, Blatt LM
Molecular therapy. Nucleic acids 2022 Mar 8;27:335-348
Molecular therapy. Nucleic acids 2022 Mar 8;27:335-348
BRD4 prevents the accumulation of R-loops and protects against transcription-replication collision events and DNA damage.
Lam FC, Kong YW, Huang Q, Vu Han TL, Maffa AD, Kasper EM, Yaffe MB
Nature communications 2020 Aug 14;11(1):4083
Nature communications 2020 Aug 14;11(1):4083
Splicing events in the control of genome integrity: role of SLU7 and truncated SRSF3 proteins.
Jiménez M, Urtasun R, Elizalde M, Azkona M, Latasa MU, Uriarte I, Arechederra M, Alignani D, Bárcena-Varela M, Álvarez-Sola G, Colyn L, Santamaría E, Sangro B, Rodriguez-Ortigosa C, Fernández-Barrena MG, Ávila MA, Berasain C
Nucleic acids research 2019 Apr 23;47(7):3450-3466
Nucleic acids research 2019 Apr 23;47(7):3450-3466
SAMMSON fosters cancer cell fitness by concertedly enhancing mitochondrial and cytosolic translation.
Vendramin R, Verheyden Y, Ishikawa H, Goedert L, Nicolas E, Saraf K, Armaos A, Delli Ponti R, Izumikawa K, Mestdagh P, Lafontaine DLJ, Tartaglia GG, Takahashi N, Marine JC, Leucci E
Nature structural & molecular biology 2018 Nov;25(11):1035-1046
Nature structural & molecular biology 2018 Nov;25(11):1035-1046
5-methylcytosine promotes mRNA export - NSUN2 as the methyltransferase and ALYREF as an m(5)C reader.
Yang X, Yang Y, Sun BF, Chen YS, Xu JW, Lai WY, Li A, Wang X, Bhattarai DP, Xiao W, Sun HY, Zhu Q, Ma HL, Adhikari S, Sun M, Hao YJ, Zhang B, Huang CM, Huang N, Jiang GB, Zhao YL, Wang HL, Sun YP, Yang YG
Cell research 2017 May;27(5):606-625
Cell research 2017 May;27(5):606-625
Human satellite-III non-coding RNAs modulate heat-shock-induced transcriptional repression.
Goenka A, Sengupta S, Pandey R, Parihar R, Mohanta GC, Mukerji M, Ganesh S
Journal of cell science 2016 Oct 1;129(19):3541-3552
Journal of cell science 2016 Oct 1;129(19):3541-3552
Comparative expression patterns and diagnostic efficacies of SR splicing factors and HNRNPA1 in gastric and colorectal cancer.
Park WC, Kim HR, Kang DB, Ryu JS, Choi KH, Lee GO, Yun KJ, Kim KY, Park R, Yoon KH, Cho JH, Lee YJ, Chae SC, Park MC, Park DS
BMC cancer 2016 Jun 10;16:358
BMC cancer 2016 Jun 10;16:358
Regulation of gene expression via retrotransposon insertions and the noncoding RNA 4.5S RNAH.
Ishida K, Miyauchi K, Kimura Y, Mito M, Okada S, Suzuki T, Nakagawa S
Genes to cells : devoted to molecular & cellular mechanisms 2015 Nov;20(11):887-901
Genes to cells : devoted to molecular & cellular mechanisms 2015 Nov;20(11):887-901
Splicing factor SRSF1 negatively regulates alternative splicing of MDM2 under damage.
Comiskey DF Jr, Jacob AG, Singh RK, Tapia-Santos AS, Chandler DS
Nucleic acids research 2015 Apr 30;43(8):4202-18
Nucleic acids research 2015 Apr 30;43(8):4202-18
The adipogenic transcriptional cofactor ZNF638 interacts with splicing regulators and influences alternative splicing.
Du C, Ma X, Meruvu S, Hugendubler L, Mueller E
Journal of lipid research 2014 Sep;55(9):1886-96
Journal of lipid research 2014 Sep;55(9):1886-96
Nuclear ARVCF protein binds splicing factors and contributes to the regulation of alternative splicing.
Rappe U, Schlechter T, Aschoff M, Hotz-Wagenblatt A, Hofmann I
The Journal of biological chemistry 2014 May 2;289(18):12421-34
The Journal of biological chemistry 2014 May 2;289(18):12421-34
The splicing factor FUBP1 is required for the efficient splicing of oncogene MDM2 pre-mRNA.
Jacob AG, Singh RK, Mohammad F, Bebee TW, Chandler DS
The Journal of biological chemistry 2014 Jun 20;289(25):17350-64
The Journal of biological chemistry 2014 Jun 20;289(25):17350-64
A METTL3-METTL14 complex mediates mammalian nuclear RNA N6-adenosine methylation.
Liu J, Yue Y, Han D, Wang X, Fu Y, Zhang L, Jia G, Yu M, Lu Z, Deng X, Dai Q, Chen W, He C
Nature chemical biology 2014 Feb;10(2):93-5
Nature chemical biology 2014 Feb;10(2):93-5
RRP1B is a metastasis modifier that regulates the expression of alternative mRNA isoforms through interactions with SRSF1.
Lee M, Dworkin AM, Gildea D, Trivedi NS, NISC Comparative Sequencing Program, Moorhead GB, Crawford NP
Oncogene 2014 Apr 3;33(14):1818-27
Oncogene 2014 Apr 3;33(14):1818-27
ALKBH5 is a mammalian RNA demethylase that impacts RNA metabolism and mouse fertility.
Zheng G, Dahl JA, Niu Y, Fedorcsak P, Huang CM, Li CJ, Vågbø CB, Shi Y, Wang WL, Song SH, Lu Z, Bosmans RP, Dai Q, Hao YJ, Yang X, Zhao WM, Tong WM, Wang XJ, Bogdan F, Furu K, Fu Y, Jia G, Zhao X, Liu J, Krokan HE, Klungland A, Yang YG, He C
Molecular cell 2013 Jan 10;49(1):18-29
Molecular cell 2013 Jan 10;49(1):18-29
CTD serine-2 plays a critical role in splicing and termination factor recruitment to RNA polymerase II in vivo.
Gu B, Eick D, Bensaude O
Nucleic acids research 2013 Feb 1;41(3):1591-603
Nucleic acids research 2013 Feb 1;41(3):1591-603
Adaptive alternative splicing correlates with less environmental risk of parkinsonism.
Benmoyal-Segal L, Soreq L, Ben-Shaul Y, Ben-Ari S, Ben-Moshe T, Aviel S, Bergman H, Soreq H
Neuro-degenerative diseases 2012;9(2):87-98
Neuro-degenerative diseases 2012;9(2):87-98
Hypoxia is a modifier of SMN2 splicing and disease severity in a severe SMA mouse model.
Bebee TW, Dominguez CE, Samadzadeh-Tarighat S, Akehurst KL, Chandler DS
Human molecular genetics 2012 Oct 1;21(19):4301-13
Human molecular genetics 2012 Oct 1;21(19):4301-13
Malat1 is not an essential component of nuclear speckles in mice.
Nakagawa S, Ip JY, Shioi G, Tripathi V, Zong X, Hirose T, Prasanth KV
RNA (New York, N.Y.) 2012 Aug;18(8):1487-99
RNA (New York, N.Y.) 2012 Aug;18(8):1487-99
Cell-to-cell variability of alternative RNA splicing.
Waks Z, Klein AM, Silver PA
Molecular systems biology 2011 Jul 5;7:506
Molecular systems biology 2011 Jul 5;7:506
SAM68 regulates neuronal activity-dependent alternative splicing of neurexin-1.
Iijima T, Wu K, Witte H, Hanno-Iijima Y, Glatter T, Richard S, Scheiffele P
Cell 2011 Dec 23;147(7):1601-14
Cell 2011 Dec 23;147(7):1601-14
Protein kinase a-dependent phosphorylation of serine 119 in the proto-oncogenic serine/arginine-rich splicing factor 1 modulates its activity as a splicing enhancer protein.
Aksaas AK, Eikvar S, Akusjärvi G, Skålhegg BS, Kvissel AK
Genes & cancer 2011 Aug;2(8):841-51
Genes & cancer 2011 Aug;2(8):841-51
S1-1 nuclear domains: characterization and dynamics as a function of transcriptional activity.
Inoue A, Tsugawa K, Tokunaga K, Takahashi KP, Uni S, Kimura M, Nishio K, Yamamoto N, Honda K, Watanabe T, Yamane H, Tani T
Biology of the cell 2008 Sep;100(9):523-35
Biology of the cell 2008 Sep;100(9):523-35
Three-dimensional positioning of genes in mouse cell nuclei.
Hepperger C, Mannes A, Merz J, Peters J, Dietzel S
Chromosoma 2008 Dec;117(6):535-51
Chromosoma 2008 Dec;117(6):535-51
The switch in alternative splicing of cyclic AMP-response element modulator protein CREM{tau}2{alpha} (activator) to CREM{alpha} (repressor) in human myometrial cells is mediated by SRp40.
Tyson-Capper AJ, Bailey J, Krainer AR, Robson SC, Europe-Finner GN
The Journal of biological chemistry 2005 Oct 14;280(41):34521-9
The Journal of biological chemistry 2005 Oct 14;280(41):34521-9
F-actin-dependent insolubility of chromatin-modifying components.
Andrin C, Hendzel MJ
The Journal of biological chemistry 2004 Jun 11;279(24):25017-23
The Journal of biological chemistry 2004 Jun 11;279(24):25017-23
No comments: Submit comment
Supportive validation
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
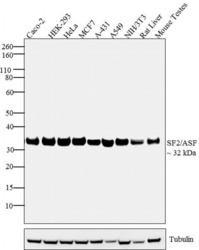
- Western blot analysis was performed on whole cell extracts of Caco-2 (Lane 1), HEK-293 (Lane 2), HeLa (Lane 3), MCF7 (Lane 4), A-431 (Lane 5), A549 (Lane 6), NIH/3T3 (Lane 7) and tissue lysates of Rat Liver (Lane 8) and Mouse Testes (Lane 9). The blots were probed with Anti- SF2/ASF Mouse Monoclonal Antibody (Product # 32-4600, 2µg/mL) and detected by chemiluminescence using Goat anti-Mouse IgG (H+L) Superclonal™ Secondary Antibody, HRP conjµgate (Product # A28177, 0.4 µg/mL, 1:2500 dilution). A ~ 32 kDa band corresponding to SF2/ASF was observed across all cell lines and tissues tested. Known quantity of protein samples were electrophoresed using Novex® NuPAGE® 4-12 % Bis-Tris gel (Product # NP0321BOX), XCell SureLock™ Electrophoresis System (Product # EI0002) and Novex® Sharp Pre-Stained Protein Standard (Product # LC5800). Resolved proteins were then transferred onto a nitrocellulose membrane with iBlot® 2 Dry Blotting System (Product # IB21001). The membrane was probed with the relevant primary and secondary Antibody following blocking with 5 % skimmed milk. Chemiluminescent detection was performed using Pierce™ ECL Western Blotting Substrate (Product # 32106).
Supportive validation
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- Immunofluorescence analysis of SF2/ASF was performed using 70% confluent log phase MCF-7 cells. The cells were fixed with 4% paraformaldehyde for 10 minutes, permeabilized with 0.1% Triton™ X-100 for 10 minutes, and blocked with 1% BSA for 1 hour at room temperature. The cells were labeled with SF2 / ASF (103) Mouse Monoclonal Antibody (Product # 32-4600) at 2µg/mL in 0.1% BSA and incubated for 3 hours at room temperature and then labeled with Goat anti-Mouse IgG (H+L) Superclonal™ Secondary Antibody, Alexa Fluor® 488 conjµgate (Product # A28175) at a dilution of 1:2000 for 45 minutes at room temperature (Panel a: green). Nuclei (Panel b: blue) were stained with SlowFade® Gold Antifade Mountant with DAPI (Product # S36938). F-actin (Panel c: red) was stained with Alexa Fluor® 555 Rhodamine Phalloidin (Product # R415, 1:300). Panel d represents the merged image showing nuclear localization. Panel e shows the no primary antibody control. The images were captured at 60X magnification.
Supportive validation
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
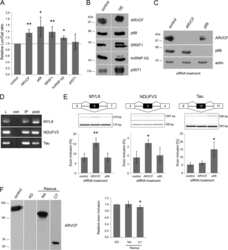
- Figure 6 Sources of cell-to-cell variability in fractional isoform abundance. ( A ) Schematic of sources of cell-to-cell variability in FIA. ( B - D ) Fluctuations in synthesis rate are a source of FIA variability, particularly for MKNK2, but cannot explain the increased variability in HeLa cells compared with Rpe1 cells. (B) Schematic demonstrating how synthesis rate fluctuations, in this case transcriptional bursting, together with differential isoform lifetimes can lead to FIA fluctuations and thus increase FIA variability. (C) Measurements of isoform lifetimes show differential isoform stability primarily for MKNK2. Data represent values from a fitted curve+-s.e.m. (D) Measured total cytoplasmic mRNA abundance variability is greater than that predicted in the absence of synthesis rate fluctuations (see Materials and methods and Supplementary Theory ). ( E ) Total cytoplasmic mRNA, but not FIA, is correlated with DAPI intensity. ( F - H ) SRSF1 knockdown causes a large increase in FIA variability beyond the theoretical minimum (binomial isoform partitioning). Efficient knockdown of SRSF1 was confirmed by western blotting (F), and led to decreased isoform 1 abundance and increased isoform 2 abundance (G). (H) FIA variability compared with binomial isoform partitioning. Bars in (D, E, G, and H) represent mean+-s.d. ( n =3). Source data is available for this figure at www.nature.com/msb . Source Data for Figure 6
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
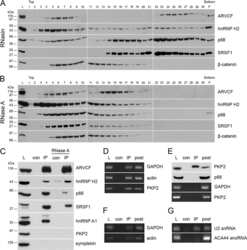
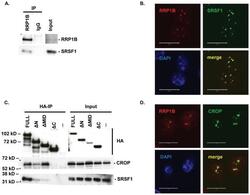
- Figure 1 Co-immunoprecipitation and co-localization of RRP1B with SRSF1 and CROP A. Western blot analysis of co-immunoprecipitation of endogenous RRP1B and SRSF1. B. Co-immunofluorescence of full-length HA-tagged RRP1B and endogenous SRSF1. Scale bar measures 10 muM. C. Co-immunoprecipitation of HA-tagged RRP1B constructs with endogenous SRSF1 and CROP. Lysates from 293T cells transfected with HA-tagged RRP1B were incubated with anti-HA for immunoprecipitation and blotted with anti-SRSF1 or anti-CROP. D. Co-immunofluorescence of full-length HA-tagged RRP1B and endogenous CROP. Scale bar measures 10 muM.
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- Figure 2 Inhibition of transcription enhances the interaction between RRP1B and SRSF1 A. Co-immunoprecipitation of RRP1B and SRSF1 with DRB or Act D treatment. 293T cells treated with DMSO, 25 mug/mL DRB, or 2.5 mug/mL and 5 mug/mL Act D as indicated were used for immunoprecipitation with anti-RRP1B. Normal rabbit IgG was used as a negative control. B. Co-immunofluorescence of RRP1B and SRSF1 after DMSO or 25 mug/mL DRB treatment. 293T cells were treated with 2 h DMSO or DRB and collected for immunofluorescence. Co-localization was confirmed using confocal fluorescence microscopy. Scale bar measures 10 muM.
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
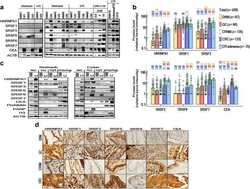
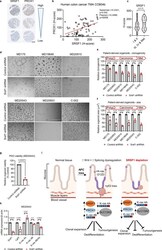
- Srsf1 depletion inhibits growth of human colorectal cancer-derived organoids. a A sample of cores taken from a human colon cancer tissue microarray (TMA), with the same respective core stained for SRSF1 or PROX1 on different sections. Scale bar 500 um. b Linear regression analysis showing the correlation of SRSF1 and PROX1 staining (based on histoscore) on the TMA shown in a , with each datapoint representing a core taken from a patient. c Relationship between SRSF1 immunohistochemistry staining and tumour stage in human patients, using TMA CO2081b, n = 54 vs 56 biologically independent tumour cores. d Representative images of patient-derived organoids (PDOs) treated with control or Srsf1 shRNAs. MD175 (polyp), MD20853 (CRC), MD20910 (CRC), MD19648 (FAP rectum Tumour), MD20043 (rectal carcinoma), C-002 (liver metastasis). Scale bar 1000 um. e Number of surviving organoid clones after shRNA treatment and f size of indicated PDOs, n = 3 vs 3 independent experiments. g Viability of PDO MD20043 after Srsf1 shRNA treatment and h mRNA expression (qPCR) of indicated genes, n = 3 vs 3 independent experiments. i Model outlining the role of SRSF1 in modulating tumour cell plasticity and invasion in colorectal cancer. Data in bar charts are represented as mean and error bars are SD with data analysed with two-tailed, unpaired t- tests, p values are indicated in figure panels. All biological replicates are shown as individual value plots and n > 3. See also Figure S7 .
 Explore
Explore Validate
Validate Learn
Learn Western blot
Western blot