Antibody data
- Antibody Data
- Antigen structure
- References [6]
- Comments [0]
- Validations
- Flow cytometry [1]
- Other assay [4]
Submit
Validation data
Reference
Comment
Report error
- Product number
- 62-1239-42 - Provider product page

- Provider
- Invitrogen Antibodies
- Product name
- CD123 Monoclonal Antibody (6H6), Super Bright™ 436, eBioscience™
- Antibody type
- Monoclonal
- Antigen
- Other
- Description
- The 6H6 monoclonal antibody reacts with human CD123, the alpha chain of the IL-3 receptor. This 60-70 kDa transmembrane protein binds to IL-3 with low affinity by itself, and when associated with CD131 (common beta chain) binds IL-3 with high affinity. CD123 is expressed by myeloid precursors, macrophages, dendritic cells, mast cells, basophils, and megakaryocytes.
- Antibody clone number
- 6H6
- Concentration
- 5 µL/Test
Submitted references Impaired IL-23-dependent induction of IFN-γ underlies mycobacterial disease in patients with inherited TYK2 deficiency.
Unsupervised Analysis of Flow Cytometry Data in a Clinical Setting Captures Cell Diversity and Allows Population Discovery.
Mucosal-associated invariant T cell responses differ by sex in COVID-19.
Neutrophils Inhibit Synthesis of Mineralized Extracellular Matrix by Human Bone Marrow-Derived Stromal Cells In Vitro.
Malaria-induced NLRP12/NLRP3-dependent caspase-1 activation mediates inflammation and hypersensitivity to bacterial superinfection.
Frailty in old age is associated with decreased interleukin-12/23 production in response to toll-like receptor ligation.
Ogishi M, Arias AA, Yang R, Han JE, Zhang P, Rinchai D, Halpern J, Mulwa J, Keating N, Chrabieh M, Lainé C, Seeleuthner Y, Ramírez-Alejo N, Nekooie-Marnany N, Guennoun A, Muller-Fleckenstein I, Fleckenstein B, Kilic SS, Minegishi Y, Ehl S, Kaiser-Labusch P, Kendir-Demirkol Y, Rozenberg F, Errami A, Zhang SY, Zhang Q, Bohlen J, Philippot Q, Puel A, Jouanguy E, Pourmoghaddas Z, Bakhtiar S, Willasch AM, Horneff G, Llanora G, Shek LP, Chai LYA, Tay SH, Rahimi HH, Mahdaviani SA, Nepesov S, Bousfiha AA, Erdeniz EH, Karbuz A, Marr N, Navarrete C, Adeli M, Hammarstrom L, Abolhassani H, Parvaneh N, Al Muhsen S, Alosaimi MF, Alsohime F, Nourizadeh M, Moin M, Arnaout R, Alshareef S, El-Baghdadi J, Genel F, Sherkat R, Kiykim A, Yücel E, Keles S, Bustamante J, Abel L, Casanova JL, Boisson-Dupuis S
The Journal of experimental medicine 2022 Oct 3;219(10)
The Journal of experimental medicine 2022 Oct 3;219(10)
Unsupervised Analysis of Flow Cytometry Data in a Clinical Setting Captures Cell Diversity and Allows Population Discovery.
Baumgaertner P, Sankar M, Herrera F, Benedetti F, Barras D, Thierry AC, Dangaj D, Kandalaft LE, Coukos G, Xenarios I, Guex N, Harari A
Frontiers in immunology 2021;12:633910
Frontiers in immunology 2021;12:633910
Mucosal-associated invariant T cell responses differ by sex in COVID-19.
Yu C, Littleton S, Giroux NS, Mathew R, Ding S, Kalnitsky J, Yang Y, Petzold E, Chung HA, Rivera GO, Rotstein T, Xi R, Ko ER, Tsalik EL, Sempowski GD, Denny TN, Burke TW, McClain MT, Woods CW, Shen X, Saban DR
Med (New York, N.Y.) 2021 Jun 11;2(6):755-772.e5
Med (New York, N.Y.) 2021 Jun 11;2(6):755-772.e5
Neutrophils Inhibit Synthesis of Mineralized Extracellular Matrix by Human Bone Marrow-Derived Stromal Cells In Vitro.
Bastian OW, Croes M, Alblas J, Koenderman L, Leenen LPH, Blokhuis TJ
Frontiers in immunology 2018;9:945
Frontiers in immunology 2018;9:945
Malaria-induced NLRP12/NLRP3-dependent caspase-1 activation mediates inflammation and hypersensitivity to bacterial superinfection.
Ataide MA, Andrade WA, Zamboni DS, Wang D, Souza Mdo C, Franklin BS, Elian S, Martins FS, Pereira D, Reed G, Fitzgerald KA, Golenbock DT, Gazzinelli RT
PLoS pathogens 2014 Jan;10(1):e1003885
PLoS pathogens 2014 Jan;10(1):e1003885
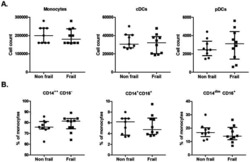
Frailty in old age is associated with decreased interleukin-12/23 production in response to toll-like receptor ligation.
Compté N, Zouaoui Boudjeltia K, Vanhaeverbeek M, De Breucker S, Tassignon J, Trelcat A, Pepersack T, Goriely S
PloS one 2013;8(6):e65325
PloS one 2013;8(6):e65325
No comments: Submit comment
Supportive validation
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- Staining of normal human peripheral blood cells with Anti-Human CD85g (ILT7) APC (Product # 17-5179-42) and Mouse IgG1 K Isotype Control Super Bright 436 (Product # 62-4714-82) (left) or Anti-Human CD123 Super Bright 436 (right). Cells in the monocyte gate were used for analysis.
Supportive validation
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
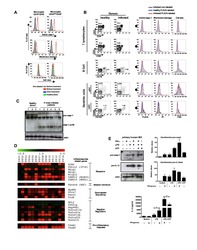
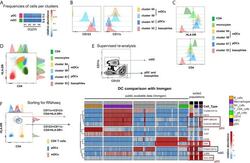
- Figure 5 Identification of multiple mDC and pDC subsets by MegaClust. (A) Comparison of the different clusters identified by MegaClust as mDCs and pDCs (example for one patient for four time points). (B) Expression level of CD123 and CD11c of the two mDC clusters and the two pDC clusters identified by MegaClust. (C) Expression levels of HLA-DR and CD4 for the two pDC clusters and the two mDC clusters identified by MegaClust. Cluster 53 could be assigned as basophils due to the lack of expression of CD4 and HLA-DR (). The expression of CD4 is shown for CD4 T cells and CD14 + monocyte clusters for comparison. (D) Overlay of CD4 and HLA-DR of MegaClust identified clusters for mDC, pDC, basophils, CD4 and monocytes. (E) mDCs and pDCs were discriminated according to CD11c and CD123 expression in the supervised flow cytometry re-analysis according to the new gating strategy described in Supplementary Figure 5 (one representative patient 0QZW). (F) Representative illustration of HLA-DR and CD4 co-expression on mDCs and pDCs prior to FACS sorting CD11c+CD123 - CD4 + HLA-DR+ mDCs and CD123 + CD11c - CD4 + HLA-DR + were FACS sorted (dotted lines). (G) Gene expression profiles of flow cytometry sorted CD11c + CD123 - CD4 + HLA-DR + mDCs, CD123 + CD11c - CD4 + HLA-DR + CD4 + HLA-DR + pDC and CD14 + CD16 - monocytes from 3 healthy donors are shown in comparison to the RNA expression profile of public available RNAseq datasets (Immgen) indicating the gene expression profile of B cells, mac
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
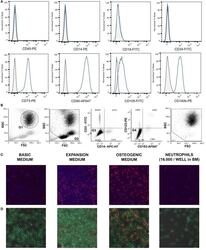
- Figure 1 (A) Surface antigen expression of bone marrow stromal cells (BMSCs) isolated from the talus bone marrow using flow cytometry. >95% of cells were negative for CD45 and CD14, and >99% of cells were negative for CD19 and CD34. In addition, >95% were positive for CD73, CD90, CD105, and CD140b. Since plastic adherence is a well-established and validated technique to isolate multipotent stromal cells (MSCs), we have only characterized one BMSC donor using flowcytometry instead of all donors. The blue lines are stained cells and the gray lines are negative (unstained) controls. Adapted from Croes et al. ( 25 ). (B) Fluorescence-activated cell sorting (FACS) gating strategy used to isolate granulocytes/neutrophils from peripheral blood leukocytes. Granulocytes were either isolated from unlabeled leukocytes using gate 1 (G1) within the forward/sideward scatter (FSC/SSC). Alternatively, leukocytes were stained using CD3, CD14, CD193, and CD123. Within the FSC/SSC of these labeled cells, debris was first excluded [gate 2 (G2)]. Subsequently, CD3+ cells (lymphocytes) and CD14+ (monocytes) were excluded [gate 3 (G3)]. In addition, CD193+ cells (eosinophils) and CD123+ cells (basophils) were excluded [gate 4 (G4)]. The remaining CD3- CD14- CD193- CD123- cells were defined FACS-sorted neutrophils (G2+, G3+, G4+ sorted neutrophils). Re-analysis of FACS-sorted neutrophils shows adequate exclusion of lymphocytes and monocytes based on their FSC/SSC. (C) Images of BMSCs obtained by arr
 Explore
Explore Validate
Validate Learn
Learn Flow cytometry
Flow cytometry