Antibody data
- Antibody Data
- Antigen structure
- References [18]
- Comments [0]
- Validations
- Western blot [1]
- Immunoprecipitation [2]
- Immunohistochemistry [2]
Submit
Validation data
Reference
Comment
Report error
- Product number
- NB100-565 - Provider product page

- Provider
- Novus Biologicals
- Proper citation
- Novus Cat#NB100-565, RRID:AB_2105207
- Product name
- Rabbit Polyclonal FUS Antibody
- Antibody type
- Polyclonal
Submitted references PARP-1 Activation Directs FUS to DNA Damage Sites to Form PARG-Reversible Compartments Enriched in Damaged DNA.
Heterogeneous nuclear ribonucleoproteins R and Q accumulate in pathological inclusions in FTLD-FUS.
FUS (fused in sarcoma) is a component of the cellular response to topoisomerase I-induced DNA breakage and transcriptional stress.
Lymphoblastoid cell lines as a model to understand amyotrophic lateral sclerosis disease mechanisms.
The Effect of Different Types of Nanoparticles on FUS and TDP-43 Solubility and Subcellular Localization.
Modelling FUS Mislocalisation in an In Vitro Model of Innervated Human Muscle.
ALS/FTD-associated FUS activates GSK-3β to disrupt the VAPB-PTPIP51 interaction and ER-mitochondria associations.
The presence of heterogeneous nuclear ribonucleoproteins in frontotemporal lobar degeneration with FUS-positive inclusions.
Recursive splicing in long vertebrate genes.
Widespread binding of FUS along nascent RNA regulates alternative splicing in the brain.
FUS/TLS-immunoreactive neuronal and glial cell inclusions increase with disease duration in familial amyotrophic lateral sclerosis with an R521C FUS/TLS mutation.
Progressive neuronal inclusion formation and axonal degeneration in CHMP2B mutant transgenic mice.
A comparative clinical, pathological, biochemical and genetic study of fused in sarcoma proteinopathies.
Globular glial tauopathies (GGT) presenting with motor neuron disease or frontotemporal dementia: an emerging group of 4-repeat tauopathies.
A Drosophila model of FUS-related neurodegeneration reveals genetic interaction between FUS and TDP-43.
Distinct pathological subtypes of FTLD-FUS.
Intracellular localization and splicing regulation of FUS/TLS are variably affected by amyotrophic lateral sclerosis-linked mutations.
A new subtype of frontotemporal lobar degeneration with FUS pathology.
Singatulina AS, Hamon L, Sukhanova MV, Desforges B, Joshi V, Bouhss A, Lavrik OI, Pastré D
Cell reports 2019 May 7;27(6):1809-1821.e5
Cell reports 2019 May 7;27(6):1809-1821.e5
Heterogeneous nuclear ribonucleoproteins R and Q accumulate in pathological inclusions in FTLD-FUS.
Gittings LM, Foti SC, Benson BC, Gami-Patel P, Isaacs AM, Lashley T
Acta neuropathologica communications 2019 Feb 12;7(1):18
Acta neuropathologica communications 2019 Feb 12;7(1):18
FUS (fused in sarcoma) is a component of the cellular response to topoisomerase I-induced DNA breakage and transcriptional stress.
Martinez-Macias MI, Moore DA, Green RL, Gomez-Herreros F, Naumann M, Hermann A, Van Damme P, Hafezparast M, Caldecott KW
Life science alliance 2019 Apr;2(2)
Life science alliance 2019 Apr;2(2)
Lymphoblastoid cell lines as a model to understand amyotrophic lateral sclerosis disease mechanisms.
Pansarasa O, Bordoni M, Drufuca L, Diamanti L, Sproviero D, Trotti R, Bernuzzi S, La Salvia S, Gagliardi S, Ceroni M, Cereda C
Disease models & mechanisms 2018 Mar 26;11(3)
Disease models & mechanisms 2018 Mar 26;11(3)
The Effect of Different Types of Nanoparticles on FUS and TDP-43 Solubility and Subcellular Localization.
Lojk J, Prpar Mihevc S, Bregar VB, Pavlin M, Rogelj B
Neurotoxicity research 2017 Oct;32(3):325-339
Neurotoxicity research 2017 Oct;32(3):325-339
Modelling FUS Mislocalisation in an In Vitro Model of Innervated Human Muscle.
Prpar Mihevc S, Pavlin M, Darovic S, Živin M, Podbregar M, Rogelj B, Mars T
Journal of molecular neuroscience : MN 2017 Aug;62(3-4):318-328
Journal of molecular neuroscience : MN 2017 Aug;62(3-4):318-328
ALS/FTD-associated FUS activates GSK-3β to disrupt the VAPB-PTPIP51 interaction and ER-mitochondria associations.
Stoica R, Paillusson S, Gomez-Suaga P, Mitchell JC, Lau DH, Gray EH, Sancho RM, Vizcay-Barrena G, De Vos KJ, Shaw CE, Hanger DP, Noble W, Miller CC
EMBO reports 2016 Sep;17(9):1326-42
EMBO reports 2016 Sep;17(9):1326-42
The presence of heterogeneous nuclear ribonucleoproteins in frontotemporal lobar degeneration with FUS-positive inclusions.
Gami-Patel P, Bandopadhyay R, Brelstaff J, Revesz T, Lashley T
Neurobiology of aging 2016 Oct;46:192-203
Neurobiology of aging 2016 Oct;46:192-203
Recursive splicing in long vertebrate genes.
Sibley CR, Emmett W, Blazquez L, Faro A, Haberman N, Briese M, Trabzuni D, Ryten M, Weale ME, Hardy J, Modic M, Curk T, Wilson SW, Plagnol V, Ule J
Nature 2015 May 21;521(7552):371-375
Nature 2015 May 21;521(7552):371-375
Widespread binding of FUS along nascent RNA regulates alternative splicing in the brain.
Rogelj B, Easton LE, Bogu GK, Stanton LW, Rot G, Curk T, Zupan B, Sugimoto Y, Modic M, Haberman N, Tollervey J, Fujii R, Takumi T, Shaw CE, Ule J
Scientific reports 2012;2:603
Scientific reports 2012;2:603
FUS/TLS-immunoreactive neuronal and glial cell inclusions increase with disease duration in familial amyotrophic lateral sclerosis with an R521C FUS/TLS mutation.
Suzuki N, Kato S, Kato M, Warita H, Mizuno H, Kato M, Shimakura N, Akiyama H, Kobayashi Z, Konno H, Aoki M
Journal of neuropathology and experimental neurology 2012 Sep;71(9):779-88
Journal of neuropathology and experimental neurology 2012 Sep;71(9):779-88
Progressive neuronal inclusion formation and axonal degeneration in CHMP2B mutant transgenic mice.
Ghazi-Noori S, Froud KE, Mizielinska S, Powell C, Smidak M, Fernandez de Marco M, O'Malley C, Farmer M, Parkinson N, Fisher EM, Asante EA, Brandner S, Collinge J, Isaacs AM
Brain : a journal of neurology 2012 Mar;135(Pt 3):819-32
Brain : a journal of neurology 2012 Mar;135(Pt 3):819-32
A comparative clinical, pathological, biochemical and genetic study of fused in sarcoma proteinopathies.
Lashley T, Rohrer JD, Bandopadhyay R, Fry C, Ahmed Z, Isaacs AM, Brelstaff JH, Borroni B, Warren JD, Troakes C, King A, Al-Saraj S, Newcombe J, Quinn N, Ostergaard K, Schrøder HD, Bojsen-Møller M, Braendgaard H, Fox NC, Rossor MN, Lees AJ, Holton JL, Revesz T
Brain : a journal of neurology 2011 Sep;134(Pt 9):2548-64
Brain : a journal of neurology 2011 Sep;134(Pt 9):2548-64
Globular glial tauopathies (GGT) presenting with motor neuron disease or frontotemporal dementia: an emerging group of 4-repeat tauopathies.
Ahmed Z, Doherty KM, Silveira-Moriyama L, Bandopadhyay R, Lashley T, Mamais A, Hondhamuni G, Wray S, Newcombe J, O'Sullivan SS, Wroe S, de Silva R, Holton JL, Lees AJ, Revesz T
Acta neuropathologica 2011 Oct;122(4):415-28
Acta neuropathologica 2011 Oct;122(4):415-28
A Drosophila model of FUS-related neurodegeneration reveals genetic interaction between FUS and TDP-43.
Lanson NA Jr, Maltare A, King H, Smith R, Kim JH, Taylor JP, Lloyd TE, Pandey UB
Human molecular genetics 2011 Jul 1;20(13):2510-23
Human molecular genetics 2011 Jul 1;20(13):2510-23
Distinct pathological subtypes of FTLD-FUS.
Mackenzie IR, Munoz DG, Kusaka H, Yokota O, Ishihara K, Roeber S, Kretzschmar HA, Cairns NJ, Neumann M
Acta neuropathologica 2011 Feb;121(2):207-18
Acta neuropathologica 2011 Feb;121(2):207-18
Intracellular localization and splicing regulation of FUS/TLS are variably affected by amyotrophic lateral sclerosis-linked mutations.
Kino Y, Washizu C, Aquilanti E, Okuno M, Kurosawa M, Yamada M, Doi H, Nukina N
Nucleic acids research 2011 Apr;39(7):2781-98
Nucleic acids research 2011 Apr;39(7):2781-98
A new subtype of frontotemporal lobar degeneration with FUS pathology.
Neumann M, Rademakers R, Roeber S, Baker M, Kretzschmar HA, Mackenzie IR
Brain : a journal of neurology 2009 Nov;132(Pt 11):2922-31
Brain : a journal of neurology 2009 Nov;132(Pt 11):2922-31
No comments: Submit comment
Supportive validation
- Submitted by
- Novus Biologicals (provider)
- Main image
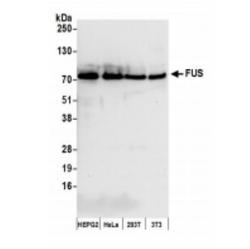
- Experimental details
- Western Blot: FUS Antibody [NB100-565] - Whole cell lysate (50 ug) from HepG2, HeLa, 293T, and mouse NIH3T3 cells prepared using NETN lysis buffer. Affinity purified rabbit anti-FUS antibody used for WB at 0.04 ug/mL. Detection: chemiluminescence with an exposure time of 10 seconds.
Supportive validation
- Submitted by
- Novus Biologicals (provider)
- Main image
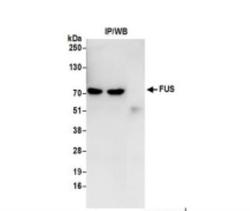
- Experimental details
- Immunoprecipitation: FUS Antibody [NB100-565] - Whole cell lysate (0.5 or 1.0 mg per IP reaction; 20% of IP loaded) from 293T cells prepared using NETN lysis buffer. Antibodies: Affinity purified rabbit antiFUS antibody used for IP at 6 ug per reaction. FUS was also immunoprecipitated by a previous lot of this antibody. For blotting immunoprecipitated FUS was used at 1 ug/ml. Detection: Chemiluminescence with an exposure time of 3 seconds.
- Submitted by
- Novus Biologicals (provider)
- Main image
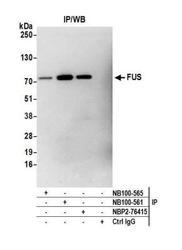
- Experimental details
- Immunoprecipitation: FUS Antibody [NB100-565] - Detection of human FUS by western blot of immunoprecipitates. Samples: Whole cell lysate (1.0 mg per IP reaction; 20% of IP loaded) from HEK293T cells prepared using NETN lysis buffer. Antibodies: Affinity purified rabbit anti-FUS antibody NB100-565 used for IP at 3 ug per reaction. FUS was also immunoprecipitated by rabbit anti-FUS recombinant monoclonal antibody [BLR023E] (NBP2-76415) and rabbit anti-FUS antibody NB100-561. For blotting immunoprecipitated FUS, NB100-565 was used at 1:1000. Detection: Chemiluminescence with an exposure time time of 3 minutes.
Supportive validation
- Submitted by
- Novus Biologicals (provider)
- Main image
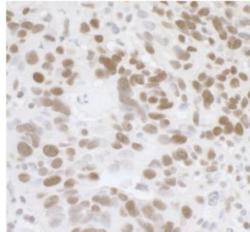
- Experimental details
- Immunohistochemistry-Paraffin: FUS Antibody [NB100-565] - FFPE section of human ovarian carcinoma. Affinity purified rabbit anti-FUS used at a dilution of 1:1,000 (1 ug/mL). Detection: DAB
- Submitted by
- Novus Biologicals (provider)
- Main image
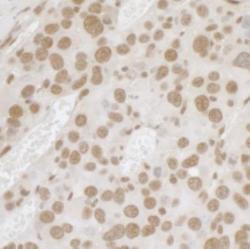
- Experimental details
- Immunohistochemistry-Paraffin: FUS Antibody [NB100-565] - FFPE section of mouse renal cells carcinoma. Affinity purified rabbit anti-FUS used at a dilution of 1:1,000 (1 ug/mL). Detection: DA
 Explore
Explore Validate
Validate Learn
Learn Western blot
Western blot Immunocytochemistry
Immunocytochemistry