Antibody data
- Antibody Data
- Antigen structure
- References [10]
- Comments [0]
- Validations
- Western blot [2]
- Other assay [11]
Submit
Validation data
Reference
Comment
Report error
- Product number
- 37-1700 - Provider product page

- Provider
- Invitrogen Antibodies
- Product name
- EphB2 Monoclonal Antibody (1A6C9)
- Antibody type
- Monoclonal
- Antigen
- Synthetic peptide
- Reactivity
- Human, Chicken/Avian
- Host
- Mouse
- Isotype
- IgG
- Antibody clone number
- 1A6C9
- Vial size
- 100 µg
- Concentration
- 0.5 mg/mL
- Storage
- -20°C
Submitted references The ephrin receptor EphB2 regulates the connectivity and activity of enteric neurons.
A T cell redirection platform for co-targeting dual antigens on solid tumors.
Positive surface charge of GluN1 N-terminus mediates the direct interaction with EphB2 and NMDAR mobility.
Receptor-specific interactome as a hub for rapid cue-induced selective translation in axons.
Role of forward and reverse signaling in Eph receptor and ephrin mediated cell segregation.
Ubiquitin ligase SPSB4 diminishes cell repulsive responses mediated by EphB2.
YES oncogenic activity is specified by its SH4 domain and regulates RAS/MAPK signaling in colon carcinoma cells.
The Eph tyrosine kinase receptors EphB2 and EphA2 are novel proteolytic substrates of tissue factor/coagulation factor VIIa.
Neuron glia-related cell adhesion molecule (NrCAM) promotes topographic retinocollicular mapping.
The EphA4 receptor regulates neuronal morphology through SPAR-mediated inactivation of Rap GTPases.
Bodin R, Paillé V, Oullier T, Durand T, Aubert P, Le Berre-Scoul C, Hulin P, Neunlist M, Cissé M
The Journal of biological chemistry 2021 Nov;297(5):101300
The Journal of biological chemistry 2021 Nov;297(5):101300
A T cell redirection platform for co-targeting dual antigens on solid tumors.
Enderle L, Shalaby KH, Gorelik M, Weiss A, Blazer LL, Paduch M, Cardarelli L, Kossiakoff A, Adams JJ, Sidhu SS
mAbs 2021 Jan-Dec;13(1):1933690
mAbs 2021 Jan-Dec;13(1):1933690
Positive surface charge of GluN1 N-terminus mediates the direct interaction with EphB2 and NMDAR mobility.
Washburn HR, Xia NL, Zhou W, Mao YT, Dalva MB
Nature communications 2020 Jan 29;11(1):570
Nature communications 2020 Jan 29;11(1):570
Receptor-specific interactome as a hub for rapid cue-induced selective translation in axons.
Koppers M, Cagnetta R, Shigeoka T, Wunderlich LC, Vallejo-Ramirez P, Qiaojin Lin J, Zhao S, Jakobs MA, Dwivedy A, Minett MS, Bellon A, Kaminski CF, Harris WA, Flanagan JG, Holt CE
eLife 2019 Nov 20;8
eLife 2019 Nov 20;8
Role of forward and reverse signaling in Eph receptor and ephrin mediated cell segregation.
Wu Z, Ashlin TG, Xu Q, Wilkinson DG
Experimental cell research 2019 Aug 1;381(1):57-65
Experimental cell research 2019 Aug 1;381(1):57-65
Ubiquitin ligase SPSB4 diminishes cell repulsive responses mediated by EphB2.
Okumura F, Joo-Okumura A, Obara K, Petersen A, Nishikimi A, Fukui Y, Nakatsukasa K, Kamura T
Molecular biology of the cell 2017 Nov 15;28(24):3532-3541
Molecular biology of the cell 2017 Nov 15;28(24):3532-3541
YES oncogenic activity is specified by its SH4 domain and regulates RAS/MAPK signaling in colon carcinoma cells.
Dubois F, Leroy C, Simon V, Benistant C, Roche S
American journal of cancer research 2015;5(6):1972-87
American journal of cancer research 2015;5(6):1972-87
The Eph tyrosine kinase receptors EphB2 and EphA2 are novel proteolytic substrates of tissue factor/coagulation factor VIIa.
Eriksson O, Ramström M, Hörnaeus K, Bergquist J, Mokhtari D, Siegbahn A
The Journal of biological chemistry 2014 Nov 21;289(47):32379-91
The Journal of biological chemistry 2014 Nov 21;289(47):32379-91
Neuron glia-related cell adhesion molecule (NrCAM) promotes topographic retinocollicular mapping.
Dai J, Buhusi M, Demyanenko GP, Brennaman LH, Hruska M, Dalva MB, Maness PF
PloS one 2013;8(9):e73000
PloS one 2013;8(9):e73000
The EphA4 receptor regulates neuronal morphology through SPAR-mediated inactivation of Rap GTPases.
Richter M, Murai KK, Bourgin C, Pak DT, Pasquale EB
The Journal of neuroscience : the official journal of the Society for Neuroscience 2007 Dec 19;27(51):14205-15
The Journal of neuroscience : the official journal of the Society for Neuroscience 2007 Dec 19;27(51):14205-15
No comments: Submit comment
Supportive validation
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
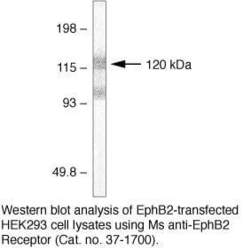
- Experimental details
- Western blot analysis of EphB2-transfected HEK293 cell lysates using mouse anti-EphB2 receptor monoclonal antibody (Product # 37-1700).
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
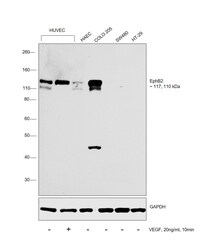
- Experimental details
- Western blot was performed using Anti-EphB2 Monoclonal Antibody (1A6C9) (Product # 37-1700) and 117, 110kDa bands corresponding to EphB2 was increased upon VEGF treatment in HUVEC. Also, the expression was highest in COLO 205 in comparison to SW480 and HT-29 as reported. Membrane enriched cell extracts (30 ug lysate) of HUVEC (Lane 1), HUVEC treated with VEGF (20ng/ml, 10 min) (Lane 2), COLO 205 (Lane 3), SW480 (Lane 4) and HT-29 (Lane 5) were electrophoresed using Novex® NuPAGE® 4-12 % Bis-Tris gel (Product # NP0322BOX). Resolved proteins were then transferred onto a nitrocellulose membrane (Product # IB23001) by iBlot® 2 Dry Blotting System (Product # IB21001). The blot was probed with the primary antibody (1:500 dilution) and detected by chemiluminescence with Goat anti-Mouse IgG (H+L), Superclonal™ Recombinant Secondary Antibody, HRP (Product # A28177, 1:4000 dilution) using the iBright FL 1000 (Product # A32752). Chemiluminescent detection was performed using Novex® ECL Chemiluminescent Substrate Reagent Kit (Product # WP20005)..
Supportive validation
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
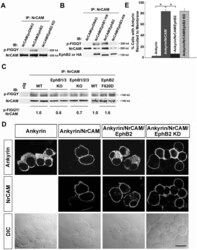
- Figure 6 EphB receptors Induce Tyrosine Phosphorylation of NrCAM at the FIGQY Motif, and Inhibit Ankyrin Recruitment in HEK293 cells. A. NrCAM was expressed with or without EphB2 or EphB2 kinase dead mutant (EphB2 KD) in transfected HEK293 cells, followed by immunoprecipitation with NrCAM antibodies and immunoblotting with phospho-FIGQY antibodies. EphB2 induced phosphorylation of NrCAM (200 kD) at FIGQY (p-FIGQY) compared to NrCAM alone or to co-expression of the EphB2 kinase dead mutant. Blots were stripped and reprobed with antibodies to NrCAM to indicate relative levels of NrCAM in the immunoprecipitates. B. NrCAM was expressed with or without EphB1, EphB2, or EphB3 in transfected HEK293 cells, followed by immunoprecipitation with NrCAM antibodies and immunoblotting with phospho-FIGQY antibodies. EphB1 (HA-tagged) or EphB2 (not HA-tagged) induced effective phosphorylation of NrCAM at FIGQY, whereas EphB3 (HA-tagged) had a weak effect. Blots were stripped and reprobed with antibodies to EphB2 or to the HA-epitope on EphB1 and EphB3. Each EphB receptor was co-immunoprecipitated efficiently with NrCAM. C. Lysates of the superior colliculus from WT, EphB1/3 double , EphB1/2/3 triple and constitutively active EphB2 (F620D) homozygous mice (P2-P3) were immunoprecipitated with antibodies to NrCAM, and immunoblotted with phospho-FIGQY or NrCAM antibodies. Bands were quantified using the threshold function of ImageJ. The numbers under the blots represent the normalized av
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
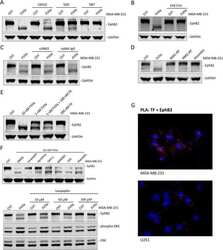
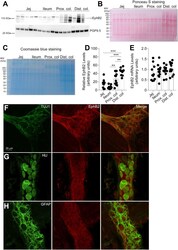
- Figure 1 EphB2 expression in different segments of the rat GI tract. A - C , EphB2 expression was examined by western blot analysis of full-thickness tissue sections from indicated regions of the rat GI tract. Pgp9.5 was used as a loading control ( A ). Note that pgp9.5 is expressed at higher levels in the distal colon. Furthermore, we observed the apparition of putative EphB2-derived bands in the colon that might be degradation products. Ponceau S ( B ) and Coomassie ( C ) stainings are provided as addition to the traditional loading control. Although equal amount of proteins was loaded for analysis, those stainings show the variability in protein migration profil and relative protein abundance between intestinal segments. Each data point represents the relative adjusted volume of protein band intensity measured by using a rectangle tool of constant volume on each lane of the WB membrane. Data are represented as relative values of protein density in arbitrary units. D , densitometric quantitation of western blot signals revealed that EphB2 is present in all segments of the gut (n = 12 sections per segment from 12 rats; jej., jejunum; prox. colon, proximal colon; dist. colon, distal colon; n = 12 sections per segment from 12 animals, F (3, 44) = 18.91, p = 0.00000005, ANOVA; Tukey's post-hoc test; ** p < 0.001, *** p < 0.0001). E , quantitative RT-PCR analysis of EphB2 mRNA levels in full-thickness tissue sections in the GI tract of adult rats. Note that there is no strict pr
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
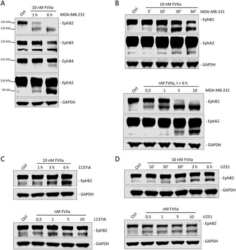
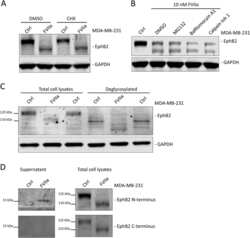
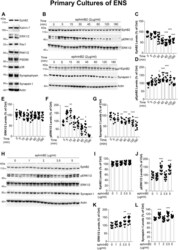
- Figure 3 EphB2 signaling in primary cultures of rat ENS. A , primary cultures of rat ENS were lysed at 12 DIV (days in vitro ) and analyzed by western blot for detection of EphB2 downstream signaling molecules as indicated. The two lanes show repeats of two samples or two wells obtained from two distinct primary cultures of ENS. B , kinetic of ephrinB2-dependent activation of EphB2. Primary cultures of E15 rat intestines were treated at 12 DIV with clustered ephrinB2-Fc (ephrinB2) at 2 mug/ml for indicated time points. Cell lysates were then subjected to immunoprecipitation and/or direct western blot analysis for detection of full-length or active EphB2 and signaling/interacting molecules as indicated. C - G , quantitation of western blot signals shown in ( B ). Phosphoproteins pEphB2 and pERK1/2 were normalized to their respective total protein levels (n = 7-18 wells per condition from at least three independent cultures, one-way ANOVA followed by Tukey's post hoc test). * p
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
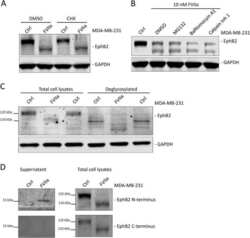
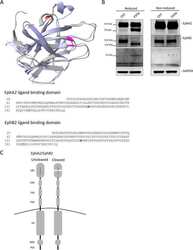
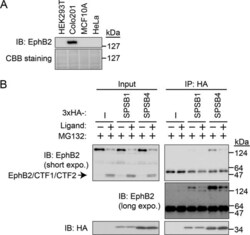
- FIGURE 3: Endogenous EphB2 interacts with SPSB1 and SPSB4. (A) Expression of endogenous EphB2 in Colo201 cells. The cell lysates of HEK293T, Colo201, MCF10A, and HeLa cells were subjected to immunoblotting with anti-EphB2 antibody. Coomassie brilliant blue (CBB) staining is shown as a loading control. (B) The interaction between endogenous EphB2 and 3x HA-SPSB4 in Colo201 cells. Colo201 cells stably expressing 3x HA-SPSB1 or 3x HA-SPSB4, and control cells were cultured in the presence of MG132 (10 muM for 1 h), and then stimulated with the ligand (clustered ephrin-B2-Fc) for 7 h in the presence of MG132. The cell lysates were lysed, immunoprecipitated (IP) with anti-HA antibody, and immunoblotted (IB) with anti-HA or anti-EphB2 antibody (recognizes the C-terminus of EphB2).
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
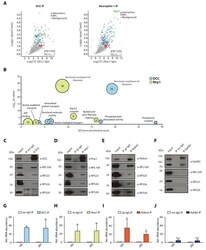
- Experimental details
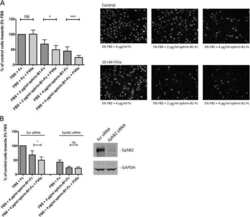
- Figure 1. Multiple guidance cue receptors interact with ribosomes. ( A ) Volcano plots showing statistically enriched proteins in DCC-IP and Nrp1-IP samples identified by permutation-based FDR-corrected t-test based on three biological replicates. The LFQ intensity of the DCC or Nrp1 pulldowns over IgG pulldowns are plotted against the -log10 p-value. FDR < 0.05; S0 = 2. ( B ) Gene enrichment analysis of statistically enriched proteins in the DCC and Nrp1 pulldown samples. The values in each circle denotes protein count. ( C-F ) Western blot validation of RP co-immunoprecipitation with DCC, Nrp1 and Robo2 but not with EphB2. Each Western blot was repeated 2 to 4 times, representative images are shown. ( G-J ) Relative 18S and 28S ribosomal RNA abundance after control (IgG) pulldown or receptors pulldowns shows enrichment of rRNA in DCC, Nrp1, and Robo2 but not EphB2 pulldowns (unpaired two-tailed t-test; three biological replicates). Bars indicate means, error bars indicate standard deviation; *p
 Explore
Explore Validate
Validate Learn
Learn Western blot
Western blot ELISA
ELISA