GTX20080
antibody from GeneTex
Targeting: CDKN2A
ARF, CDK4I, CDKN2, CMM2, INK4, INK4a, MLM, MTS1, p14, p14ARF, p16, p16INK4a, p19, p19Arf
Antibody data
- Antibody Data
- Antigen structure
- References [12]
- Comments [0]
- Validations
- Western blot [3]
- Immunocytochemistry [1]
Submit
Validation data
Reference
Comment
Report error
- Product number
- GTX20080 - Provider product page

- Provider
- GeneTex
- Proper citation
- GeneTex Cat#GTX20080, RRID:AB_848642
- Product name
- p19ARF antibody
- Antibody type
- Polyclonal
- Reactivity
- Human, Mouse, Rat
- Host
- Rabbit
Submitted references High glucose induces bone marrow-derived mesenchymal stem cell senescence by upregulating autophagy.
Inactivation of p53 is insufficient to allow B cells and B-cell lymphomas to survive without Dicer.
The p53 Codon 72 Polymorphism Modifies the Cellular Response to Inflammatory Challenge in the Liver.
Suppression of Ras/Mapk pathway signaling inhibits Myc-induced lymphomagenesis.
Egr1 mediates p53-independent c-Myc-induced apoptosis via a noncanonical ARF-dependent transcriptional mechanism.
MicroRNA biogenesis is required for Myc-induced B-cell lymphoma development and survival.
ARF induces autophagy by virtue of interaction with Bcl-xl.
The ARF tumor suppressor can promote the progression of some tumors.
ARF directly binds DP1: interaction with DP1 coincides with the G1 arrest function of ARF.
Myc-ARF (alternate reading frame) interaction inhibits the functions of Myc.
Foxm1b transcription factor is essential for development of hepatocellular carcinomas and is negatively regulated by the p19ARF tumor suppressor.
Differential regulation of E2F1, DP1, and the E2F1/DP1 complex by ARF.
Chang TC, Hsu MF, Wu KK
PloS one 2015;10(5):e0126537
PloS one 2015;10(5):e0126537
Inactivation of p53 is insufficient to allow B cells and B-cell lymphomas to survive without Dicer.
Adams CM, Eischen CM
Cancer research 2014 Jul 15;74(14):3923-34
Cancer research 2014 Jul 15;74(14):3923-34
The p53 Codon 72 Polymorphism Modifies the Cellular Response to Inflammatory Challenge in the Liver.
Leu JI, Murphy ME, George DL
Journal of liver 2013;2(1)
Journal of liver 2013;2(1)
Suppression of Ras/Mapk pathway signaling inhibits Myc-induced lymphomagenesis.
Gramling MW, Eischen CM
Cell death and differentiation 2012 Jul;19(7):1220-7
Cell death and differentiation 2012 Jul;19(7):1220-7
Egr1 mediates p53-independent c-Myc-induced apoptosis via a noncanonical ARF-dependent transcriptional mechanism.
Boone DN, Qi Y, Li Z, Hann SR
Proceedings of the National Academy of Sciences of the United States of America 2011 Jan 11;108(2):632-7
Proceedings of the National Academy of Sciences of the United States of America 2011 Jan 11;108(2):632-7
MicroRNA biogenesis is required for Myc-induced B-cell lymphoma development and survival.
Arrate MP, Vincent T, Odvody J, Kar R, Jones SN, Eischen CM
Cancer research 2010 Jul 15;70(14):6083-92
Cancer research 2010 Jul 15;70(14):6083-92
ARF induces autophagy by virtue of interaction with Bcl-xl.
Pimkina J, Humbey O, Zilfou JT, Jarnik M, Murphy ME
The Journal of biological chemistry 2009 Jan 30;284(5):2803-10
The Journal of biological chemistry 2009 Jan 30;284(5):2803-10
The ARF tumor suppressor can promote the progression of some tumors.
Humbey O, Pimkina J, Zilfou JT, Jarnik M, Dominguez-Brauer C, Burgess DJ, Eischen CM, Murphy ME
Cancer research 2008 Dec 1;68(23):9608-13
Cancer research 2008 Dec 1;68(23):9608-13
ARF directly binds DP1: interaction with DP1 coincides with the G1 arrest function of ARF.
Datta A, Sen J, Hagen J, Korgaonkar CK, Caffrey M, Quelle DE, Hughes DE, Ackerson TJ, Costa RH, Raychaudhuri P
Molecular and cellular biology 2005 Sep;25(18):8024-36
Molecular and cellular biology 2005 Sep;25(18):8024-36
Myc-ARF (alternate reading frame) interaction inhibits the functions of Myc.
Datta A, Nag A, Pan W, Hay N, Gartel AL, Colamonici O, Mori Y, Raychaudhuri P
The Journal of biological chemistry 2004 Aug 27;279(35):36698-707
The Journal of biological chemistry 2004 Aug 27;279(35):36698-707
Foxm1b transcription factor is essential for development of hepatocellular carcinomas and is negatively regulated by the p19ARF tumor suppressor.
Kalinichenko VV, Major ML, Wang X, Petrovic V, Kuechle J, Yoder HM, Dennewitz MB, Shin B, Datta A, Raychaudhuri P, Costa RH
Genes & development 2004 Apr 1;18(7):830-50
Genes & development 2004 Apr 1;18(7):830-50
Differential regulation of E2F1, DP1, and the E2F1/DP1 complex by ARF.
Datta A, Nag A, Raychaudhuri P
Molecular and cellular biology 2002 Dec;22(24):8398-408
Molecular and cellular biology 2002 Dec;22(24):8398-408
No comments: Submit comment
Supportive validation
- Submitted by
- GeneTex (provider)
- Main image

- Experimental details
- GTX20080 antibody was tested on NIH 3T3 cells (which have a deletion of the p19ARF gene) as a negative control (-) and NIH 3T3 cells engineered to express HA-tagged p19ARF as a positive control (+). Total cell lysate (70μg/lane) was probed with 70 μg/mL of GTX20080, which was detected using an anti-rabbit IgG-HRP conjμgate.
- Submitted by
- GeneTex (provider)
- Main image
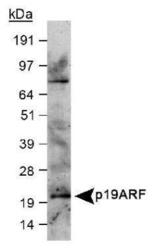
- Experimental details
- Detection of p19ARF in western blot of mouse embryonic fibroblast (MEF) lysate (25μg) using p19ARF Antibody [GTX20080] (2μg/ml).
- Submitted by
- GeneTex (provider)
- Main image
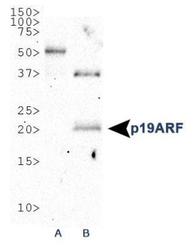
- Experimental details
- Western Blot: p19ARF Antibody (GTX20080) - WB analysis of p19ARF in 1. NIH/3T3 and B. MEF cell lysate.
Supportive validation
- Submitted by
- GeneTex (provider)
- Main image
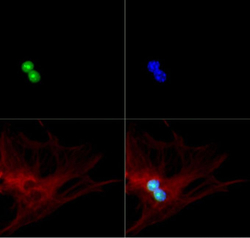
- Experimental details
- ICC/IF analysis of MEF cells using p19ARF antibody (green), alpha Tubulin (red) and DAPI (blue).
 Explore
Explore Validate
Validate Learn
Learn Western blot
Western blot ELISA
ELISA