Antibody data
- Antibody Data
- Antigen structure
- References [20]
- Comments [0]
- Validations
- Immunocytochemistry [6]
- Immunohistochemistry [1]
- Flow cytometry [5]
- Other assay [5]
Submit
Validation data
Reference
Comment
Report error
- Product number
- MA1-21818 - Provider product page

- Provider
- Invitrogen Antibodies
- Product name
- Ku70/Ku80 Monoclonal Antibody (162)
- Antibody type
- Monoclonal
- Antigen
- Other
- Description
- MA1-21818 detects Ku70/Ku80 in human, non-human primate, mouse, rat and xenopus laevis samples. It reacts with a conformational epitope of the p70/p80 dimer, which is destroyed during Western blot. This antibody is not recommended for Western blot procedures and will not cross-react with bovine or rabbit.
- Reactivity
- Human, Mouse, Rat, Xenopus
- Host
- Mouse
- Isotype
- IgG
- Antibody clone number
- 162
- Vial size
- 500 µL
- Concentration
- 0.2 mg/mL
- Storage
- 4° C
Submitted references Rad52 competes with Ku70/Ku86 for binding to S-region DSB ends to modulate antibody class-switch DNA recombination.
The DNA damage/repair cascade in glioblastoma cell lines after chemotherapeutic agent treatment.
Characterization of estrogen response element binding proteins as biomarkers of breast cancer behavior.
The DNA repair complex Ku70/86 modulates Apaf1 expression upon DNA damage.
Ku counteracts mobilization of PARP1 and MRN in chromatin damaged with DNA double-strand breaks.
The effect of DNA-dependent protein kinase on adeno-associated virus replication.
Accumulation of Ku80 proteins at DNA double-strand breaks in living cells.
Loading of the nonhomologous end joining factor, Ku, on protein-occluded DNA ends.
Interplay between Ku, Artemis, and the DNA-dependent protein kinase catalytic subunit at DNA ends.
Ongoing immunoglobulin class switch DNA recombination in lupus B cells: analysis of switch regulatory regions.
Ku70/Ku80 and DNA-dependent protein kinase catalytic subunit modulate RAG-mediated cleavage: implications for the enforcement of the 12/23 rule.
Involvement of poly(ADP-ribose) polymerase-1 and XRCC1/DNA ligase III in an alternative route for DNA double-strand breaks rejoining.
Biochemical evidence for Ku-independent backup pathways of NHEJ.
Possible anti-recombinogenic role of Bloom's syndrome helicase in double-strand break processing.
Differential DNA binding of Ku antigen determines its involvement in DNA replication.
In vivo association of Ku with mammalian origins of DNA replication.
Ku entry into DNA inhibits inward DNA transactions in vitro.
Replication-mediated DNA damage by camptothecin induces phosphorylation of RPA by DNA-dependent protein kinase and dissociates RPA:DNA-PK complexes.
The repressor which binds the -75 GATA motif of the GPB promoter contains Ku70 as the DNA binding subunit.
Physical interaction between epidermal growth factor receptor and DNA-dependent protein kinase in mammalian cells.
Zan H, Tat C, Qiu Z, Taylor JR, Guerrero JA, Shen T, Casali P
Nature communications 2017 Feb 8;8:14244
Nature communications 2017 Feb 8;8:14244
The DNA damage/repair cascade in glioblastoma cell lines after chemotherapeutic agent treatment.
Annovazzi L, Caldera V, Mellai M, Riganti C, Battaglia L, Chirio D, Melcarne A, Schiffer D
International journal of oncology 2015;46(6):2299-308
International journal of oncology 2015;46(6):2299-308
Characterization of estrogen response element binding proteins as biomarkers of breast cancer behavior.
Kruer TL, Cummins TD, Powell DW, Wittliff JL
Clinical biochemistry 2013 Nov;46(16-17):1739-46
Clinical biochemistry 2013 Nov;46(16-17):1739-46
The DNA repair complex Ku70/86 modulates Apaf1 expression upon DNA damage.
De Zio D, Bordi M, Tino E, Lanzuolo C, Ferraro E, Mora E, Ciccosanti F, Fimia GM, Orlando V, Cecconi F
Cell death and differentiation 2011 Mar;18(3):516-27
Cell death and differentiation 2011 Mar;18(3):516-27
Ku counteracts mobilization of PARP1 and MRN in chromatin damaged with DNA double-strand breaks.
Cheng Q, Barboule N, Frit P, Gomez D, Bombarde O, Couderc B, Ren GS, Salles B, Calsou P
Nucleic acids research 2011 Dec;39(22):9605-19
Nucleic acids research 2011 Dec;39(22):9605-19
The effect of DNA-dependent protein kinase on adeno-associated virus replication.
Choi YK, Nash K, Byrne BJ, Muzyczka N, Song S
PloS one 2010 Dec 20;5(12):e15073
PloS one 2010 Dec 20;5(12):e15073
Accumulation of Ku80 proteins at DNA double-strand breaks in living cells.
Koike M, Koike A
Experimental cell research 2008 Mar 10;314(5):1061-70
Experimental cell research 2008 Mar 10;314(5):1061-70
Loading of the nonhomologous end joining factor, Ku, on protein-occluded DNA ends.
Roberts SA, Ramsden DA
The Journal of biological chemistry 2007 Apr 6;282(14):10605-13
The Journal of biological chemistry 2007 Apr 6;282(14):10605-13
Interplay between Ku, Artemis, and the DNA-dependent protein kinase catalytic subunit at DNA ends.
Drouet J, Frit P, Delteil C, de Villartay JP, Salles B, Calsou P
The Journal of biological chemistry 2006 Sep 22;281(38):27784-93
The Journal of biological chemistry 2006 Sep 22;281(38):27784-93
Ongoing immunoglobulin class switch DNA recombination in lupus B cells: analysis of switch regulatory regions.
Liu S, Cerutti A, Casali P, Crow MK
Autoimmunity 2004 Sep-Nov;37(6-7):431-443
Autoimmunity 2004 Sep-Nov;37(6-7):431-443
Ku70/Ku80 and DNA-dependent protein kinase catalytic subunit modulate RAG-mediated cleavage: implications for the enforcement of the 12/23 rule.
Sawchuk DJ, Mansilla-Soto J, Alarcon C, Singha NC, Langen H, Bianchi ME, Lees-Miller SP, Nussenzweig MC, Cortes P
The Journal of biological chemistry 2004 Jul 9;279(28):29821-31
The Journal of biological chemistry 2004 Jul 9;279(28):29821-31
Involvement of poly(ADP-ribose) polymerase-1 and XRCC1/DNA ligase III in an alternative route for DNA double-strand breaks rejoining.
Audebert M, Salles B, Calsou P
The Journal of biological chemistry 2004 Dec 31;279(53):55117-26
The Journal of biological chemistry 2004 Dec 31;279(53):55117-26
Biochemical evidence for Ku-independent backup pathways of NHEJ.
Wang H, Perrault AR, Takeda Y, Qin W, Wang H, Iliakis G
Nucleic acids research 2003 Sep 15;31(18):5377-88
Nucleic acids research 2003 Sep 15;31(18):5377-88
Possible anti-recombinogenic role of Bloom's syndrome helicase in double-strand break processing.
Onclercq-Delic R, Calsou P, Delteil C, Salles B, Papadopoulo D, Amor-Guéret M
Nucleic acids research 2003 Nov 1;31(21):6272-82
Nucleic acids research 2003 Nov 1;31(21):6272-82
Differential DNA binding of Ku antigen determines its involvement in DNA replication.
Schild-Poulter C, Matheos D, Novac O, Cui B, Giffin W, Ruiz MT, Price GB, Zannis-Hadjopoulos M, Haché RJ
DNA and cell biology 2003 Feb;22(2):65-78
DNA and cell biology 2003 Feb;22(2):65-78
In vivo association of Ku with mammalian origins of DNA replication.
Novac O, Matheos D, Araujo FD, Price GB, Zannis-Hadjopoulos M
Molecular biology of the cell 2001 Nov;12(11):3386-401
Molecular biology of the cell 2001 Nov;12(11):3386-401
Ku entry into DNA inhibits inward DNA transactions in vitro.
Frit P, Li RY, Arzel D, Salles B, Calsou P
The Journal of biological chemistry 2000 Nov 17;275(46):35684-91
The Journal of biological chemistry 2000 Nov 17;275(46):35684-91
Replication-mediated DNA damage by camptothecin induces phosphorylation of RPA by DNA-dependent protein kinase and dissociates RPA:DNA-PK complexes.
Shao RG, Cao CX, Zhang H, Kohn KW, Wold MS, Pommier Y
The EMBO journal 1999 Mar 1;18(5):1397-406
The EMBO journal 1999 Mar 1;18(5):1397-406
The repressor which binds the -75 GATA motif of the GPB promoter contains Ku70 as the DNA binding subunit.
Camara-Clayette V, Thomas D, Rahuel C, Barbey R, Cartron JP, Bertrand O
Nucleic acids research 1999 Apr 1;27(7):1656-63
Nucleic acids research 1999 Apr 1;27(7):1656-63
Physical interaction between epidermal growth factor receptor and DNA-dependent protein kinase in mammalian cells.
Bandyopadhyay D, Mandal M, Adam L, Mendelsohn J, Kumar R
The Journal of biological chemistry 1998 Jan 16;273(3):1568-73
The Journal of biological chemistry 1998 Jan 16;273(3):1568-73
No comments: Submit comment
Supportive validation
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- Immunofluorescent analysis of Ku70+Ku80 (green) showing staining in the nucleus of A549 cells (right) compared to a negative control without primary antibody (left). Formalin-fixed cells were permeabilized with 0.1% Triton X-100 in TBS for 5-10 minutes and blocked with 3% BSA-PBS for 30 minutes at room temperature. Cells were probed with a Ku70+Ku80 monoclonal antibody (Product # MA1-21818) in 3% BSA-PBS at a dilution of 1:50 and incubated overnight at 4ºC in a humidified chamber. Cells were washed with PBST and incubated with a DyLight-conjugated secondary antibody in PBS at room temperature in the dark. Actin was stained using Alexa Fluor 554 (red) and nuclei were stained with Hoechst or DAPI (blue). Images were taken at a magnification of 60x.
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
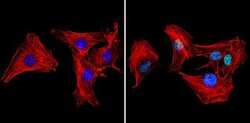
- Experimental details
- Immunofluorescent analysis of Ku70+Ku80 (green) showing staining in the nucleus of C2C12 cells (right) compared to a negative control without primary antibody (left). Formalin-fixed cells were permeabilized with 0.1% Triton X-100 in TBS for 5-10 minutes and blocked with 3% BSA-PBS for 30 minutes at room temperature. Cells were probed with a Ku70+Ku80 monoclonal antibody (Product # MA1-21818) in 3% BSA-PBS at a dilution of 1:20 and incubated overnight at 4ºC in a humidified chamber. Cells were washed with PBST and incubated with a DyLight-conjugated secondary antibody in PBS at room temperature in the dark. Actin was stained using Alexa Fluor 554 (red) and nuclei were stained with Hoechst or DAPI (blue). Images were taken at a magnification of 60x.
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
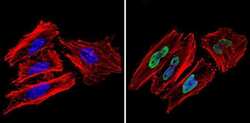
- Experimental details
- Immunofluorescent analysis of Ku70+Ku80 (green) showing staining in the nucleus of Hela cells (right) compared to a negative control without primary antibody (left). Formalin-fixed cells were permeabilized with 0.1% Triton X-100 in TBS for 5-10 minutes and blocked with 3% BSA-PBS for 30 minutes at room temperature. Cells were probed with a Ku70+Ku80 monoclonal antibody (Product # MA1-21818) in 3% BSA-PBS at a dilution of 1:50 and incubated overnight at 4ºC in a humidified chamber. Cells were washed with PBST and incubated with a DyLight-conjugated secondary antibody in PBS at room temperature in the dark. Actin was stained using Alexa Fluor 554 (red) and nuclei were stained with Hoechst or DAPI (blue). Images were taken at a magnification of 60x.
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
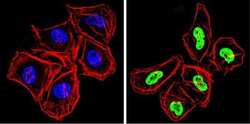
- Experimental details
- Immunofluorescent analysis of Ku (p70/p80) (green) showing staining in the nucleus of HeLa cells. Formalin-fixed cells were permeabilized with 0.1% Triton X-100 in TBS for 5-10 minutes and blocked with 3% BSA-PBS for 30 minutes at room temperature. Cells were probed with a Ku (p70/p80) monoclonal antibody (Product # MA1-21818) in 3% BSA-PBS at a dilution of 1:200 and incubated overnight at 4 ºC in a humidified chamber. Cells were washed with PBST and incubated with a DyLight-conjugated secondary antibody in PBS at room temperature in the dark. F-actin (red) was stained with a fluorescent red phalloidin and nuclei (blue) were stained with Hoechst or DAPI. Images were taken at a magnification of 60x.
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- Immunofluorescent analysis of Ku (p70/p80) (green) showing staining in the nucleus of MCF-7 cells. Formalin-fixed cells were permeabilized with 0.1% Triton X-100 in TBS for 5-10 minutes and blocked with 3% BSA-PBS for 30 minutes at room temperature. Cells were probed with a Ku (p70/p80) monoclonal antibody (Product # MA1-21818) in 3% BSA-PBS at a dilution of 1:200 and incubated overnight at 4 ºC in a humidified chamber. Cells were washed with PBST and incubated with a DyLight-conjugated secondary antibody in PBS at room temperature in the dark. F-actin (red) was stained with a fluorescent red phalloidin and nuclei (blue) were stained with Hoechst or DAPI. Images were taken at a magnification of 60x.
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
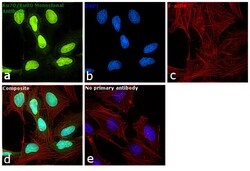
- Experimental details
- Immunofluorescence analysis of Ku70/Ku80 was performed using 70% confluent log phase HeLa cells. The cells were fixed with 4% paraformaldehyde for 10 minutes, permeabilized with 0.1% Triton™ X-100 for 15 minutes, and blocked with 1% BSA for 1 hour at room temperature. The cells were labeled with Ku70/Ku80 Monoclonal Antibody (162) (Product # MA1-21818) at 1:100 dilution in 0.1% BSA, incubated at 4 degree Celsius overnight and then labeled with Goat anti-Mouse IgG (H+L) Superclonal™ Secondary Antibody, Alexa Fluor® 488 conjugate (Product # A28175) at a dilution of 1:2000 for 45 minutes at room temperature (Panel a: green). Nuclei (Panel b: blue) were stained with ProLong™ Diamond Antifade Mountant with DAPI (Product # P36962). F-actin (Panel c: red) was stained with Rhodamine Phalloidin (Product # R415, 1:300). Panel d represents the merged image showing nuclear localization. Panel e represents control cells with no primary antibody to assess background. The images were captured at 60X magnification.
Supportive validation
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- Immunohistochemical analysis of formalin-fixed, paraffin-embedded human tonsil stained using a Ku monoclonal antibody (Product # MA1-21818) with a peroxidase-conjugate and AEC chromogen. Note nuclear staining of the cells.
Supportive validation
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- Flow cytometry analysis of Ku70+Ku80 in A549 cells (green) compared to an isotype control (blue). Cells were harvested, adjusted to a concentration of 1-5x10^6 cells/mL, fixed with 2% paraformaldehyde and washed with PBS. Cells were blocked with a 2% solution of BSA-PBS for 30 min at room temperature and incubated with a Ku70+Ku80 monoclonal antibody (Product # MA1-21818) at a dilution of 1 µg/test for 40 min at room temperature. Cells were then incubated for 40 min at room temperature in the dark using a Dylight 488-conjugated secondary antibody and re-suspended in PBS for FACS analysis.
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- Flow cytometry analysis of Ku70+Ku80 in C2C12 cells (green) compared to an isotype control (blue). Cells were harvested, adjusted to a concentration of 1-5x10^6 cells/mL, fixed with 2% paraformaldehyde and washed with PBS. Cells were blocked with a 2% solution of BSA-PBS for 30 min at room temperature and incubated with a Ku70+Ku80 monoclonal antibody (Product # MA1-21818) at a dilution of 1 µg/test for 40 min at room temperature. Cells were then incubated for 40 min at room temperature in the dark using a Dylight 488-conjugated secondary antibody and re-suspended in PBS for FACS analysis.
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- Flow cytometry analysis of Ku70+Ku80 in Hela cells (green) compared to an isotype control (blue). Cells were harvested, adjusted to a concentration of 1-5x10^6 cells/mL, fixed with 2% paraformaldehyde and washed with PBS. Cells were blocked with a 2% solution of BSA-PBS for 30 min at room temperature and incubated with a Ku70+Ku80 monoclonal antibody (Product # MA1-21818) at a dilution of 1 µg/test for 40 min at room temperature. Cells were then incubated for 40 min at room temperature in the dark using a Dylight 488-conjugated secondary antibody and re-suspended in PBS for FACS analysis.
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- Flow cytometry analysis of Ku (p70/p80) in HepG2 cells (green) compared to an isotype control (blue). Cells were harvested, adjusted to a concentration of 1-5x10^6 cells/mL, fixed with 2% paraformaldehyde and washed with PBS. Cells were blocked with a 2% solution of BSA-PBS for 30 min at room temperature and incubated with a Ku (p70/p80) monoclonal antibody (Product # MA1-21818) at a dilution of 0.5 µg/test for 60 min at room temperature. Cells were then incubated for 40 min at room temperature in the dark using a Dylight 488-conjugated goat anti-mouse IgG (H+L) secondary antibody and re-suspended in PBS for FACS analysis.
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- Flow cytometry analysis of Ku (p70/p80) in MCF-7 cells (green) compared to an isotype control (blue). Cells were harvested, adjusted to a concentration of 1-5x10^6 cells/mL, fixed with 2% paraformaldehyde and washed with PBS. Cells were blocked with a 2% solution of BSA-PBS for 30 min at room temperature and incubated with a Ku (p70/p80) monoclonal antibody (Product # MA1-21818) at a dilution of 0.5 µg/test for 60 min at room temperature. Cells were then incubated for 40 min at room temperature in the dark using a Dylight 488-conjugated goat anti-mouse IgG (H+L) secondary antibody and re-suspended in PBS for FACS analysis.
Supportive validation
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
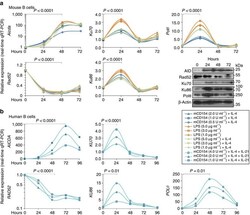
- Figure 9 Aicda ( AICDA )/CSR-inducing stimuli decrease expression of Rad52 ( RAD52 ) and increase expression of Ku70/Ku86 ( KU70/KU86 ) and Poltheta ( POLtheta ) in mouse and human B cells. ( a ) C57BL/6 B cells were stimulated with different amounts of LPS only, or LPS or mCD154 plus IL-4 for 0, 24, 48 and 72 h. Aicda , Rad52 , Poltheta and Ku70 / Ku86 transcripts were analysed by real-time qRT-PCR. Expression of Aicda normalized to Cd79b expression, expression of Rad52 , Ku70/Ku86 and Poltheta normalized to Gapdh expression and depicted relative to the expression in unstimulated B cells, set as 1. Expression of AID, Rad52, Ku70, Ku86, Poltheta and beta-Actin proteins in B cells stimulated with LPS (3.0 mug ml -1 ) plus IL-4 were analysed by immunoblotting. ( b ) Human naive B cells were stimulated with different amounts of mCD154 plus human IL-4 and human IL-21 for 0, 24, 48, 72 or 96 h. AICDA , RAD52 , KU70 / KU86 and POLtheta transcripts were analysed by real-time qRT-PCR. Expression normalized to CD79b expression and depicted relative to the expression in unstimulated B cells, set as 1. P -values determined using a paired Student's t -test. Data are from one representative of three independent experiments.
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
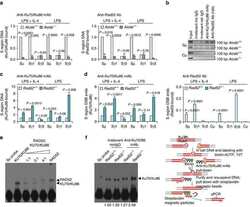
- Figure 1 Rad52 deficiency increases CSR. ( a ) B cells purified from Rad52 +/+ and Rad52 -/- C57BL/6 littermates were stimulated with mCD154 or LPS plus IL-4 (for CSR to IgG1), LPS alone (IgG3), LPS plus IFN-gamma (IgG2c) or LPS plus TGF-beta, IL-4, IL-5 and anti-delta mAb/dex (IgA). Purified Aicda +/+ and Aicda -/- B cells were stimulated with LPS plus IL-4. After 96 h of culture, the cells were analysed for surface IgG1, IgG3, IgG2c or IgA by flow cytometry. ( b ) Proliferation of Rad52 +/+ and Rad52 -/- B cells labelled with CFSE and stimulated for 72 h with different amounts of LPS or mCD154 plus IL-4. Progressive left shift of fluorescence intensity indicates B220 + B cell division. ( c ) Proliferation of Rad52 +/+ and Rad52 -/- B cells labelled with the cell division tracking fluorochrome CFSE and stimulated by mCD154 plus IL-4 or LPS plus IL-4 for 96 h. CFSE intensity and surface IgG1 expression analysed by flow cytometry. Proportion of surface IgG1 + B cells at each cell division indicated. P -values determined using a paired Student's t -test. Data are from one representative (left panels of each condition) or mean+-s.d. of three independent experiments (right panels of each condition). ( d ) Recombinant Smu-Sgamma1 or Smu-Sgamma3 DNAs analysed by digestion-circularization PCR (DC-PCR) using serially twofold diluted HindIII digested and T4 DNA ligase-ligated genomic DNA from Rad52 +/+ or Rad52 -/- B cells after stimulation with LPS or LPS plus IL-4 for 96 h. Gap
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
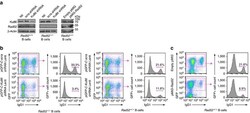
- Figure 8 Knockdown of Ku86 expression in Rad52 -/- B cells virtually ablates CSR and enforced expression of Rad52 in normal B cells significantly impairs CSR. Rad52 -/- and Rad52 +/+ B cells were transduced with pGFP-C-Ku86-shRNALenti lentiviral vector expressing Ku86 -specific shRNA and GFP, or pGFP-C-scr-shRNALenti lentiviral vector expressing scrambled shRNA and GFP. Rad52 +/+ B cells were activated with LPS for 12 h and transduced with empty pMIG-GFP retroviral vector or pMIG-GFP-Rad52 retroviral vector expressing recombinant Rad52. After lentiviral or retroviral transduction, B cells were cultured for 96 h with LPS plus IL-4. ( a ) B-cell expression of Ku86, Rad52 and beta-Actin proteins analysed by immunoblotting. ( b ) Proportions of surface IgG1 + B cells among pGFP-C-scr-shRNALenti or pGFP-C-Ku86-shRNALenti lentiviral vector-transduced (B220 + GFP + ) Rad52 -/- and Rad52 +/+ B cells were analysed by flow cytometry. Data are from one representative of three independent experiments. ( c ) Proportions of surface IgG1 + B cells among empty pMIG-GFP or pMIG-GFP-Rad52 retroviral vector-transduced (B220 + GFP + ) Rad52 +/+ B cells were analysed by flow cytometry. Data are from one representative of three independent experiments.
 Explore
Explore Validate
Validate Learn
Learn Immunocytochemistry
Immunocytochemistry