Antibody data
- Antibody Data
- Antigen structure
- References [72]
- Comments [0]
- Validations
- Western blot [3]
- Immunocytochemistry [3]
- Flow cytometry [3]
- Other assay [9]
Submit
Validation data
Reference
Comment
Report error
- Product number
- MA5-12933 - Provider product page

- Provider
- Invitrogen Antibodies
- Product name
- Ku80 Monoclonal Antibody (111)
- Antibody type
- Monoclonal
- Antigen
- Other
- Description
- MA5-12933 targets Ku (p80) in FACS, ICC/IF, IP, and WB applications and shows reactivity with Human, Mouse and Non-human primate samples. This antibody does not cross-react with mouse in immunofluorescent applications, but does react with mouse in FACS applications. The MA5-12933 immunogen is human B cell nuclei from plasmacytoid 2p68 cells.
- Reactivity
- Human, Mouse
- Host
- Mouse
- Isotype
- IgG
- Antibody clone number
- 111
- Vial size
- 500 µL
- Concentration
- 0.2 mg/mL
- Storage
- 4° C
Submitted references The Ubiquitin Ligase RNF138 Cooperates with CtIP to Stimulate Resection of Complex DNA Double-Strand Breaks in Human G1-Phase Cells.
Genetic instability from a single S phase after whole-genome duplication.
Transcription-associated topoisomerase 2α (TOP2A) activity is a major effector of cytotoxicity induced by G-quadruplex ligands.
VAV2 is required for DNA repair and implicated in cancer radiotherapy resistance.
Reciprocal regulation of RIG-I and XRCC4 connects DNA repair with RIG-I immune signaling.
Systematic Characterization of Recurrent Genomic Alterations in Cyclin-Dependent Kinases Reveals Potential Therapeutic Strategies for Cancer Treatment.
LINP1 facilitates DNA damage repair through non-homologous end joining (NHEJ) pathway and subsequently decreases the sensitivity of cervical cancer cells to ionizing radiation.
Temporal DNA-PK activation drives genomic instability and therapy resistance in glioma stem cells.
SMARCAD1 Phosphorylation and Ubiquitination Are Required for Resection during DNA Double-Strand Break Repair.
Rad52 competes with Ku70/Ku86 for binding to S-region DSB ends to modulate antibody class-switch DNA recombination.
Long noncoding RNA LINP1 regulates repair of DNA double-strand breaks in triple-negative breast cancer.
A new phosphorylated form of Ku70 identified in resistant leukemic cells confers fast but unfaithful DNA repair in cancer cell lines.
The combination of Hsp90 inhibitor 17AAG and heavy-ion irradiation provides effective tumor control in human lung cancer cells.
SMCHD1 accumulates at DNA damage sites and facilitates the repair of DNA double-strand breaks.
Centrosomal localisation of the cancer/testis (CT) antigens NY-ESO-1 and MAGE-C1 is regulated by proteasome activity in tumour cells.
Characterization of estrogen response element binding proteins as biomarkers of breast cancer behavior.
Imatinib radiosensitizes bladder cancer by targeting homologous recombination.
A noncatalytic function of the ligation complex during nonhomologous end joining.
DNA damage-induced inhibition of rRNA synthesis by DNA-PK and PARP-1.
A new method for high-resolution imaging of Ku foci to decipher mechanisms of DNA double-strand break repair.
Ku80 functions as a tumor suppressor in hepatocellular carcinoma by inducing S-phase arrest through a p53-dependent pathway.
Concordant and opposite roles of DNA-PK and the "facilitator of chromatin transcription" (FACT) in DNA repair, apoptosis and necrosis after cisplatin.
A DNA-dependent stress response involving DNA-PK occurs in hypoxic cells and contributes to cellular adaptation to hypoxia.
The effect of DNA-dependent protein kinase on adeno-associated virus replication.
Involvement of p54(nrb), a PSF partner protein, in DNA double-strand break repair and radioresistance.
Hyperactivation of DNA-PK by double-strand break mimicking molecules disorganizes DNA damage response.
Ku proteins function as corepressors to regulate farnesoid X receptor-mediated gene expression.
Down regulation of BRCA2 causes radio-sensitization of human tumor cells in vitro and in vivo.
Cell-surface MMP-9 regulates the invasive capacity of leukemia blast cells with monocytic features.
Cell-surface MMP-9 regulates the invasive capacity of leukemia blast cells with monocytic features.
DNA-dependent conformational changes in the Ku heterodimer.
Transcription-dependent nucleolar cap localization and possible nuclear function of DExH RNA helicase RHAU.
Analytical biochemistry of DNA--protein assemblies from crude cell extracts.
Interplay between Cernunnos-XLF and nonhomologous end-joining proteins at DNA ends in the cell.
Long-term XPC silencing reduces DNA double-strand break repair.
Human cytomegalovirus disrupts both ataxia telangiectasia mutated protein (ATM)- and ATM-Rad3-related kinase-mediated DNA damage responses during lytic infection.
Interplay between Ku, Artemis, and the DNA-dependent protein kinase catalytic subunit at DNA ends.
Impaired DNA double strand break repair in cells from Nijmegen breakage syndrome patients.
Proteomic identification of Ku70/Ku80 autoantigen recognized by monoclonal antibody against hepatocellular carcinoma.
Human chronic lymphocytic leukemia B cells can escape DNA damage-induced apoptosis through the nonhomologous end-joining DNA repair pathway.
Cell cycle-dependent nuclear retention of p53 by E2F1 requires phosphorylation of p53 at Ser315.
The HoxC4 homeodomain protein mediates activation of the immunoglobulin heavy chain 3' hs1,2 enhancer in human B cells. Relevance to class switch DNA recombination.
The membrane form of the DNA repair protein Ku interacts at the cell surface with metalloproteinase 9.
Involvement of poly(ADP-ribose) polymerase-1 and XRCC1/DNA ligase III in an alternative route for DNA double-strand breaks rejoining.
Suppression of retroviral infection by the RAD52 DNA repair protein.
DNA damage modulates nucleolar interaction of the Werner protein with the AAA ATPase p97/VCP.
Distinct pathways of nonhomologous end joining that are differentially regulated by DNA-dependent protein kinase-mediated phosphorylation.
Possible anti-recombinogenic role of Bloom's syndrome helicase in double-strand break processing.
Possible anti-recombinogenic role of Bloom's syndrome helicase in double-strand break processing.
Evidence implicating Ku antigen as a structural factor in RNA polymerase II-mediated transcription.
Evidence implicating Ku antigen as a structural factor in RNA polymerase II-mediated transcription.
Coordinated assembly of Ku and p460 subunits of the DNA-dependent protein kinase on DNA ends is necessary for XRCC4-ligase IV recruitment.
Coordinated assembly of Ku and p460 subunits of the DNA-dependent protein kinase on DNA ends is necessary for XRCC4-ligase IV recruitment.
The radioprotective effect of the 24 kDa FGF-2 isoform in HeLa cells is related to an increased expression and activity of the DNA dependent protein kinase (DNA-PK) catalytic subunit.
Induction of protein translation by ADAR1 within living cell nuclei is not dependent on RNA editing.
Binding of inositol hexakisphosphate (IP6) to Ku but not to DNA-PKcs.
Hairpin opening and overhang processing by an Artemis/DNA-dependent protein kinase complex in nonhomologous end joining and V(D)J recombination.
Hairpin opening and overhang processing by an Artemis/DNA-dependent protein kinase complex in nonhomologous end joining and V(D)J recombination.
A method to detect particle-specific antibodies against Ku and the DNA-dependent protein kinase catalytic subunit in autoimmune sera.
Nuclear localization of Ku antigen is promoted independently by basic motifs in the Ku70 and Ku80 subunits.
Hypoxia-activated ligand HAL-1/13 is lupus autoantigen Ku80 and mediates lymphoid cell adhesion in vitro.
Hypoxia-activated ligand HAL-1/13 is lupus autoantigen Ku80 and mediates lymphoid cell adhesion in vitro.
Ku entry into DNA inhibits inward DNA transactions in vitro.
Interactions of the DNA ligase IV-XRCC4 complex with DNA ends and the DNA-dependent protein kinase.
Mutations in human ARF exon 2 disrupt its nucleolar localization and impair its ability to block nuclear export of MDM2 and p53.
Mutations in human ARF exon 2 disrupt its nucleolar localization and impair its ability to block nuclear export of MDM2 and p53.
Ku antigen-DNA conformation determines the activation of DNA-dependent protein kinase and DNA sequence-directed repression of mouse mammary tumor virus transcription.
Ku antigen-DNA conformation determines the activation of DNA-dependent protein kinase and DNA sequence-directed repression of mouse mammary tumor virus transcription.
Poly(ADP-ribose) polymerase and Ku autoantigen form a complex and synergistically bind to matrix attachment sequences.
Poly(ADP-ribose) polymerase and Ku autoantigen form a complex and synergistically bind to matrix attachment sequences.
The repressor which binds the -75 GATA motif of the GPB promoter contains Ku70 as the DNA binding subunit.
The repressor which binds the -75 GATA motif of the GPB promoter contains Ku70 as the DNA binding subunit.
Averbeck NB, Barent C, Jakob B, Syzonenko T, Durante M, Taucher-Scholz G
Cells 2022 Aug 17;11(16)
Cells 2022 Aug 17;11(16)
Genetic instability from a single S phase after whole-genome duplication.
Gemble S, Wardenaar R, Keuper K, Srivastava N, Nano M, Macé AS, Tijhuis AE, Bernhard SV, Spierings DCJ, Simon A, Goundiam O, Hochegger H, Piel M, Foijer F, Storchová Z, Basto R
Nature 2022 Apr;604(7904):146-151
Nature 2022 Apr;604(7904):146-151
Transcription-associated topoisomerase 2α (TOP2A) activity is a major effector of cytotoxicity induced by G-quadruplex ligands.
Bossaert M, Pipier A, Riou JF, Noirot C, Nguyên LT, Serre RF, Bouchez O, Defrancq E, Calsou P, Britton S, Gomez D
eLife 2021 Jun 28;10
eLife 2021 Jun 28;10
VAV2 is required for DNA repair and implicated in cancer radiotherapy resistance.
Liu W, Miao C, Zhang S, Liu Y, Niu X, Xi Y, Guo W, Chu J, Lin A, Liu H, Yang X, Chen X, Zhong C, Ma Y, Wang Y, Zhu S, Liu S, Tan W, Lin D, Wu C
Signal transduction and targeted therapy 2021 Aug 30;6(1):322
Signal transduction and targeted therapy 2021 Aug 30;6(1):322
Reciprocal regulation of RIG-I and XRCC4 connects DNA repair with RIG-I immune signaling.
Guo G, Gao M, Gao X, Zhu B, Huang J, Tu X, Kim W, Zhao F, Zhou Q, Zhu S, Wu Z, Yan Y, Zhang Y, Zeng X, Zhu Q, Yin P, Luo K, Sun J, Deng M, Lou Z
Nature communications 2021 Apr 12;12(1):2187
Nature communications 2021 Apr 12;12(1):2187
Systematic Characterization of Recurrent Genomic Alterations in Cyclin-Dependent Kinases Reveals Potential Therapeutic Strategies for Cancer Treatment.
Shan W, Yuan J, Hu Z, Jiang J, Wang Y, Loo N, Fan L, Tang Z, Zhang T, Xu M, Pan Y, Lu J, Long M, Tanyi JL, Montone KT, Fan Y, Hu X, Zhang Y, Zhang L
Cell reports 2020 Jul 14;32(2):107884
Cell reports 2020 Jul 14;32(2):107884
LINP1 facilitates DNA damage repair through non-homologous end joining (NHEJ) pathway and subsequently decreases the sensitivity of cervical cancer cells to ionizing radiation.
Wang X, Liu H, Shi L, Yu X, Gu Y, Sun X
Cell cycle (Georgetown, Tex.) 2018;17(4):439-447
Cell cycle (Georgetown, Tex.) 2018;17(4):439-447
Temporal DNA-PK activation drives genomic instability and therapy resistance in glioma stem cells.
Wang Y, Xu H, Liu T, Huang M, Butter PP, Li C, Zhang L, Kao GD, Gong Y, Maity A, Koumenis C, Fan Y
JCI insight 2018 Feb 8;3(3)
JCI insight 2018 Feb 8;3(3)
SMARCAD1 Phosphorylation and Ubiquitination Are Required for Resection during DNA Double-Strand Break Repair.
Chakraborty S, Pandita RK, Hambarde S, Mattoo AR, Charaka V, Ahmed KM, Iyer SP, Hunt CR, Pandita TK
iScience 2018 Apr 27;2:123-135
iScience 2018 Apr 27;2:123-135
Rad52 competes with Ku70/Ku86 for binding to S-region DSB ends to modulate antibody class-switch DNA recombination.
Zan H, Tat C, Qiu Z, Taylor JR, Guerrero JA, Shen T, Casali P
Nature communications 2017 Feb 8;8:14244
Nature communications 2017 Feb 8;8:14244
Long noncoding RNA LINP1 regulates repair of DNA double-strand breaks in triple-negative breast cancer.
Zhang Y, He Q, Hu Z, Feng Y, Fan L, Tang Z, Yuan J, Shan W, Li C, Hu X, Tanyi JL, Fan Y, Huang Q, Montone K, Dang CV, Zhang L
Nature structural & molecular biology 2016 Jun;23(6):522-30
Nature structural & molecular biology 2016 Jun;23(6):522-30
A new phosphorylated form of Ku70 identified in resistant leukemic cells confers fast but unfaithful DNA repair in cancer cell lines.
Bouley J, Saad L, Grall R, Schellenbauer A, Biard D, Paget V, Morel-Altmeyer S, Guipaud O, Chambon C, Salles B, Maloum K, Merle-Béral H, Chevillard S, Delic J
Oncotarget 2015 Sep 29;6(29):27980-8000
Oncotarget 2015 Sep 29;6(29):27980-8000
The combination of Hsp90 inhibitor 17AAG and heavy-ion irradiation provides effective tumor control in human lung cancer cells.
Hirakawa H, Fujisawa H, Masaoka A, Noguchi M, Hirayama R, Takahashi M, Fujimori A, Okayasu R
Cancer medicine 2015 Mar;4(3):426-36
Cancer medicine 2015 Mar;4(3):426-36
SMCHD1 accumulates at DNA damage sites and facilitates the repair of DNA double-strand breaks.
Coker H, Brockdorff N
Journal of cell science 2014 May 1;127(Pt 9):1869-74
Journal of cell science 2014 May 1;127(Pt 9):1869-74
Centrosomal localisation of the cancer/testis (CT) antigens NY-ESO-1 and MAGE-C1 is regulated by proteasome activity in tumour cells.
Pagotto A, Caballero OL, Volkmar N, Devalle S, Simpson AJ, Lu X, Christianson JC
PloS one 2013;8(12):e83212
PloS one 2013;8(12):e83212
Characterization of estrogen response element binding proteins as biomarkers of breast cancer behavior.
Kruer TL, Cummins TD, Powell DW, Wittliff JL
Clinical biochemistry 2013 Nov;46(16-17):1739-46
Clinical biochemistry 2013 Nov;46(16-17):1739-46
Imatinib radiosensitizes bladder cancer by targeting homologous recombination.
Qiao B, Kerr M, Groselj B, Teo MT, Knowles MA, Bristow RG, Phillips RM, Kiltie AE
Cancer research 2013 Mar 1;73(5):1611-20
Cancer research 2013 Mar 1;73(5):1611-20
A noncatalytic function of the ligation complex during nonhomologous end joining.
Cottarel J, Frit P, Bombarde O, Salles B, Négrel A, Bernard S, Jeggo PA, Lieber MR, Modesti M, Calsou P
The Journal of cell biology 2013 Jan 21;200(2):173-86
The Journal of cell biology 2013 Jan 21;200(2):173-86
DNA damage-induced inhibition of rRNA synthesis by DNA-PK and PARP-1.
Calkins AS, Iglehart JD, Lazaro JB
Nucleic acids research 2013 Aug;41(15):7378-86
Nucleic acids research 2013 Aug;41(15):7378-86
A new method for high-resolution imaging of Ku foci to decipher mechanisms of DNA double-strand break repair.
Britton S, Coates J, Jackson SP
The Journal of cell biology 2013 Aug 5;202(3):579-95
The Journal of cell biology 2013 Aug 5;202(3):579-95
Ku80 functions as a tumor suppressor in hepatocellular carcinoma by inducing S-phase arrest through a p53-dependent pathway.
Wei S, Xiong M, Zhan DQ, Liang BY, Wang YY, Gutmann DH, Huang ZY, Chen XP
Carcinogenesis 2012 Mar;33(3):538-47
Carcinogenesis 2012 Mar;33(3):538-47
Concordant and opposite roles of DNA-PK and the "facilitator of chromatin transcription" (FACT) in DNA repair, apoptosis and necrosis after cisplatin.
Sand-Dejmek J, Adelmant G, Sobhian B, Calkins AS, Marto J, Iglehart DJ, Lazaro JB
Molecular cancer 2011 Jun 16;10:74
Molecular cancer 2011 Jun 16;10:74
A DNA-dependent stress response involving DNA-PK occurs in hypoxic cells and contributes to cellular adaptation to hypoxia.
Bouquet F, Ousset M, Biard D, Fallone F, Dauvillier S, Frit P, Salles B, Muller C
Journal of cell science 2011 Jun 1;124(Pt 11):1943-51
Journal of cell science 2011 Jun 1;124(Pt 11):1943-51
The effect of DNA-dependent protein kinase on adeno-associated virus replication.
Choi YK, Nash K, Byrne BJ, Muzyczka N, Song S
PloS one 2010 Dec 20;5(12):e15073
PloS one 2010 Dec 20;5(12):e15073
Involvement of p54(nrb), a PSF partner protein, in DNA double-strand break repair and radioresistance.
Li S, Kuhne WW, Kulharya A, Hudson FZ, Ha K, Cao Z, Dynan WS
Nucleic acids research 2009 Nov;37(20):6746-53
Nucleic acids research 2009 Nov;37(20):6746-53
Hyperactivation of DNA-PK by double-strand break mimicking molecules disorganizes DNA damage response.
Quanz M, Chassoux D, Berthault N, Agrario C, Sun JS, Dutreix M
PloS one 2009 Jul 21;4(7):e6298
PloS one 2009 Jul 21;4(7):e6298
Ku proteins function as corepressors to regulate farnesoid X receptor-mediated gene expression.
Ohno M, Kunimoto M, Nishizuka M, Osada S, Imagawa M
Biochemical and biophysical research communications 2009 Dec 18;390(3):738-42
Biochemical and biophysical research communications 2009 Dec 18;390(3):738-42
Down regulation of BRCA2 causes radio-sensitization of human tumor cells in vitro and in vivo.
Yu D, Sekine E, Fujimori A, Ochiya T, Okayasu R
Cancer science 2008 Apr;99(4):810-5
Cancer science 2008 Apr;99(4):810-5
Cell-surface MMP-9 regulates the invasive capacity of leukemia blast cells with monocytic features.
Paupert J, Mansat-De Mas V, Demur C, Salles B, Muller C
Cell cycle (Georgetown, Tex.) 2008 Apr 15;7(8):1047-53
Cell cycle (Georgetown, Tex.) 2008 Apr 15;7(8):1047-53
Cell-surface MMP-9 regulates the invasive capacity of leukemia blast cells with monocytic features.
Paupert J, Mansat-De Mas V, Demur C, Salles B, Muller C
Cell cycle (Georgetown, Tex.) 2008 Apr 15;7(8):1047-53
Cell cycle (Georgetown, Tex.) 2008 Apr 15;7(8):1047-53
DNA-dependent conformational changes in the Ku heterodimer.
Lehman JA, Hoelz DJ, Turchi JJ
Biochemistry 2008 Apr 15;47(15):4359-68
Biochemistry 2008 Apr 15;47(15):4359-68
Transcription-dependent nucleolar cap localization and possible nuclear function of DExH RNA helicase RHAU.
Iwamoto F, Stadler M, Chalupníková K, Oakeley E, Nagamine Y
Experimental cell research 2008 Apr 1;314(6):1378-91
Experimental cell research 2008 Apr 1;314(6):1378-91
Analytical biochemistry of DNA--protein assemblies from crude cell extracts.
Hégarat N, Cardoso GM, Rusconi F, Francois JC, Praseuth D
Nucleic acids research 2007;35(13):e92
Nucleic acids research 2007;35(13):e92
Interplay between Cernunnos-XLF and nonhomologous end-joining proteins at DNA ends in the cell.
Wu PY, Frit P, Malivert L, Revy P, Biard D, Salles B, Calsou P
The Journal of biological chemistry 2007 Nov 2;282(44):31937-43
The Journal of biological chemistry 2007 Nov 2;282(44):31937-43
Long-term XPC silencing reduces DNA double-strand break repair.
Despras E, Pfeiffer P, Salles B, Calsou P, Kuhfittig-Kulle S, Angulo JF, Biard DS
Cancer research 2007 Mar 15;67(6):2526-34
Cancer research 2007 Mar 15;67(6):2526-34
Human cytomegalovirus disrupts both ataxia telangiectasia mutated protein (ATM)- and ATM-Rad3-related kinase-mediated DNA damage responses during lytic infection.
Luo MH, Rosenke K, Czornak K, Fortunato EA
Journal of virology 2007 Feb;81(4):1934-50
Journal of virology 2007 Feb;81(4):1934-50
Interplay between Ku, Artemis, and the DNA-dependent protein kinase catalytic subunit at DNA ends.
Drouet J, Frit P, Delteil C, de Villartay JP, Salles B, Calsou P
The Journal of biological chemistry 2006 Sep 22;281(38):27784-93
The Journal of biological chemistry 2006 Sep 22;281(38):27784-93
Impaired DNA double strand break repair in cells from Nijmegen breakage syndrome patients.
Howlett NG, Scuric Z, D'Andrea AD, Schiestl RH
DNA repair 2006 Feb 3;5(2):251-7
DNA repair 2006 Feb 3;5(2):251-7
Proteomic identification of Ku70/Ku80 autoantigen recognized by monoclonal antibody against hepatocellular carcinoma.
Luk JM, Su YC, Lam SC, Lee CK, Hu MY, He QY, Lau GK, Wong FW, Fan ST
Proteomics 2005 May;5(7):1980-6
Proteomics 2005 May;5(7):1980-6
Human chronic lymphocytic leukemia B cells can escape DNA damage-induced apoptosis through the nonhomologous end-joining DNA repair pathway.
Deriano L, Guipaud O, Merle-Béral H, Binet JL, Ricoul M, Potocki-Veronese G, Favaudon V, Maciorowski Z, Muller C, Salles B, Sabatier L, Delic J
Blood 2005 Jun 15;105(12):4776-83
Blood 2005 Jun 15;105(12):4776-83
Cell cycle-dependent nuclear retention of p53 by E2F1 requires phosphorylation of p53 at Ser315.
Fogal V, Hsieh JK, Royer C, Zhong S, Lu X
The EMBO journal 2005 Aug 3;24(15):2768-82
The EMBO journal 2005 Aug 3;24(15):2768-82
The HoxC4 homeodomain protein mediates activation of the immunoglobulin heavy chain 3' hs1,2 enhancer in human B cells. Relevance to class switch DNA recombination.
Kim EC, Edmonston CR, Wu X, Schaffer A, Casali P
The Journal of biological chemistry 2004 Oct 1;279(40):42258-69
The Journal of biological chemistry 2004 Oct 1;279(40):42258-69
The membrane form of the DNA repair protein Ku interacts at the cell surface with metalloproteinase 9.
Monferran S, Paupert J, Dauvillier S, Salles B, Muller C
The EMBO journal 2004 Oct 1;23(19):3758-68
The EMBO journal 2004 Oct 1;23(19):3758-68
Involvement of poly(ADP-ribose) polymerase-1 and XRCC1/DNA ligase III in an alternative route for DNA double-strand breaks rejoining.
Audebert M, Salles B, Calsou P
The Journal of biological chemistry 2004 Dec 31;279(53):55117-26
The Journal of biological chemistry 2004 Dec 31;279(53):55117-26
Suppression of retroviral infection by the RAD52 DNA repair protein.
Lau A, Kanaar R, Jackson SP, O'Connor MJ
The EMBO journal 2004 Aug 18;23(16):3421-9
The EMBO journal 2004 Aug 18;23(16):3421-9
DNA damage modulates nucleolar interaction of the Werner protein with the AAA ATPase p97/VCP.
Partridge JJ, Lopreiato JO Jr, Latterich M, Indig FE
Molecular biology of the cell 2003 Oct;14(10):4221-9
Molecular biology of the cell 2003 Oct;14(10):4221-9
Distinct pathways of nonhomologous end joining that are differentially regulated by DNA-dependent protein kinase-mediated phosphorylation.
Udayakumar D, Bladen CL, Hudson FZ, Dynan WS
The Journal of biological chemistry 2003 Oct 24;278(43):41631-5
The Journal of biological chemistry 2003 Oct 24;278(43):41631-5
Possible anti-recombinogenic role of Bloom's syndrome helicase in double-strand break processing.
Onclercq-Delic R, Calsou P, Delteil C, Salles B, Papadopoulo D, Amor-Guéret M
Nucleic acids research 2003 Nov 1;31(21):6272-82
Nucleic acids research 2003 Nov 1;31(21):6272-82
Possible anti-recombinogenic role of Bloom's syndrome helicase in double-strand break processing.
Onclercq-Delic R, Calsou P, Delteil C, Salles B, Papadopoulo D, Amor-Guéret M
Nucleic acids research 2003 Nov 1;31(21):6272-82
Nucleic acids research 2003 Nov 1;31(21):6272-82
Evidence implicating Ku antigen as a structural factor in RNA polymerase II-mediated transcription.
Bertinato J, Tomlinson JJ, Schild-Poulter C, Haché RJ
Gene 2003 Jan 2;302(1-2):53-64
Gene 2003 Jan 2;302(1-2):53-64
Evidence implicating Ku antigen as a structural factor in RNA polymerase II-mediated transcription.
Bertinato J, Tomlinson JJ, Schild-Poulter C, Haché RJ
Gene 2003 Jan 2;302(1-2):53-64
Gene 2003 Jan 2;302(1-2):53-64
Coordinated assembly of Ku and p460 subunits of the DNA-dependent protein kinase on DNA ends is necessary for XRCC4-ligase IV recruitment.
Calsou P, Delteil C, Frit P, Drouet J, Salles B
Journal of molecular biology 2003 Feb 7;326(1):93-103
Journal of molecular biology 2003 Feb 7;326(1):93-103
Coordinated assembly of Ku and p460 subunits of the DNA-dependent protein kinase on DNA ends is necessary for XRCC4-ligase IV recruitment.
Calsou P, Delteil C, Frit P, Drouet J, Salles B
Journal of molecular biology 2003 Feb 7;326(1):93-103
Journal of molecular biology 2003 Feb 7;326(1):93-103
The radioprotective effect of the 24 kDa FGF-2 isoform in HeLa cells is related to an increased expression and activity of the DNA dependent protein kinase (DNA-PK) catalytic subunit.
Ader I, Muller C, Bonnet J, Favre G, Cohen-Jonathan E, Salles B, Toulas C
Oncogene 2002 Sep 19;21(42):6471-9
Oncogene 2002 Sep 19;21(42):6471-9
Induction of protein translation by ADAR1 within living cell nuclei is not dependent on RNA editing.
Herbert A, Wagner S, Nickerson JA
Molecular cell 2002 Nov;10(5):1235-46
Molecular cell 2002 Nov;10(5):1235-46
Binding of inositol hexakisphosphate (IP6) to Ku but not to DNA-PKcs.
Ma Y, Lieber MR
The Journal of biological chemistry 2002 Mar 29;277(13):10756-9
The Journal of biological chemistry 2002 Mar 29;277(13):10756-9
Hairpin opening and overhang processing by an Artemis/DNA-dependent protein kinase complex in nonhomologous end joining and V(D)J recombination.
Ma Y, Pannicke U, Schwarz K, Lieber MR
Cell 2002 Mar 22;108(6):781-94
Cell 2002 Mar 22;108(6):781-94
Hairpin opening and overhang processing by an Artemis/DNA-dependent protein kinase complex in nonhomologous end joining and V(D)J recombination.
Ma Y, Pannicke U, Schwarz K, Lieber MR
Cell 2002 Mar 22;108(6):781-94
Cell 2002 Mar 22;108(6):781-94
A method to detect particle-specific antibodies against Ku and the DNA-dependent protein kinase catalytic subunit in autoimmune sera.
Jafri F, Hardin JA, Dynan WS
Journal of immunological methods 2001 May 1;251(1-2):53-61
Journal of immunological methods 2001 May 1;251(1-2):53-61
Nuclear localization of Ku antigen is promoted independently by basic motifs in the Ku70 and Ku80 subunits.
Bertinato J, Schild-Poulter C, Haché RJ
Journal of cell science 2001 Jan;114(Pt 1):89-99
Journal of cell science 2001 Jan;114(Pt 1):89-99
Hypoxia-activated ligand HAL-1/13 is lupus autoantigen Ku80 and mediates lymphoid cell adhesion in vitro.
Lynch EM, Moreland RB, Ginis I, Perrine SP, Faller DV
American journal of physiology. Cell physiology 2001 Apr;280(4):C897-911
American journal of physiology. Cell physiology 2001 Apr;280(4):C897-911
Hypoxia-activated ligand HAL-1/13 is lupus autoantigen Ku80 and mediates lymphoid cell adhesion in vitro.
Lynch EM, Moreland RB, Ginis I, Perrine SP, Faller DV
American journal of physiology. Cell physiology 2001 Apr;280(4):C897-911
American journal of physiology. Cell physiology 2001 Apr;280(4):C897-911
Ku entry into DNA inhibits inward DNA transactions in vitro.
Frit P, Li RY, Arzel D, Salles B, Calsou P
The Journal of biological chemistry 2000 Nov 17;275(46):35684-91
The Journal of biological chemistry 2000 Nov 17;275(46):35684-91
Interactions of the DNA ligase IV-XRCC4 complex with DNA ends and the DNA-dependent protein kinase.
Chen L, Trujillo K, Sung P, Tomkinson AE
The Journal of biological chemistry 2000 Aug 25;275(34):26196-205
The Journal of biological chemistry 2000 Aug 25;275(34):26196-205
Mutations in human ARF exon 2 disrupt its nucleolar localization and impair its ability to block nuclear export of MDM2 and p53.
Zhang Y, Xiong Y
Molecular cell 1999 May;3(5):579-91
Molecular cell 1999 May;3(5):579-91
Mutations in human ARF exon 2 disrupt its nucleolar localization and impair its ability to block nuclear export of MDM2 and p53.
Zhang Y, Xiong Y
Molecular cell 1999 May;3(5):579-91
Molecular cell 1999 May;3(5):579-91
Ku antigen-DNA conformation determines the activation of DNA-dependent protein kinase and DNA sequence-directed repression of mouse mammary tumor virus transcription.
Giffin W, Gong W, Schild-Poulter C, Haché RJ
Molecular and cellular biology 1999 Jun;19(6):4065-78
Molecular and cellular biology 1999 Jun;19(6):4065-78
Ku antigen-DNA conformation determines the activation of DNA-dependent protein kinase and DNA sequence-directed repression of mouse mammary tumor virus transcription.
Giffin W, Gong W, Schild-Poulter C, Haché RJ
Molecular and cellular biology 1999 Jun;19(6):4065-78
Molecular and cellular biology 1999 Jun;19(6):4065-78
Poly(ADP-ribose) polymerase and Ku autoantigen form a complex and synergistically bind to matrix attachment sequences.
Galande S, Kohwi-Shigematsu T
The Journal of biological chemistry 1999 Jul 16;274(29):20521-8
The Journal of biological chemistry 1999 Jul 16;274(29):20521-8
Poly(ADP-ribose) polymerase and Ku autoantigen form a complex and synergistically bind to matrix attachment sequences.
Galande S, Kohwi-Shigematsu T
The Journal of biological chemistry 1999 Jul 16;274(29):20521-8
The Journal of biological chemistry 1999 Jul 16;274(29):20521-8
The repressor which binds the -75 GATA motif of the GPB promoter contains Ku70 as the DNA binding subunit.
Camara-Clayette V, Thomas D, Rahuel C, Barbey R, Cartron JP, Bertrand O
Nucleic acids research 1999 Apr 1;27(7):1656-63
Nucleic acids research 1999 Apr 1;27(7):1656-63
The repressor which binds the -75 GATA motif of the GPB promoter contains Ku70 as the DNA binding subunit.
Camara-Clayette V, Thomas D, Rahuel C, Barbey R, Cartron JP, Bertrand O
Nucleic acids research 1999 Apr 1;27(7):1656-63
Nucleic acids research 1999 Apr 1;27(7):1656-63
No comments: Submit comment
Supportive validation
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- Western blot of Ku (P80) using Ku (P80) Monoclonal Antibody (Product # MA5-12933) on HeLa Cells.
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
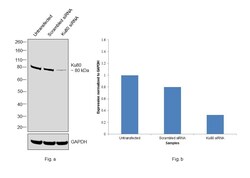
- Experimental details
- Knockdown of Ku80 was achieved by transfecting HeLa with Ku80 specific siRNAs (Silencer® select Product # s14952, s14953). Western blot analysis (Fig. a) was performed using whole cell extracts from the Ku80 knockdown cells (Lane 3), non-specific scrambled siRNA transfected cells (Lane 2) and untransfected cells (Lane 1). The blot was probed with Ku80 Monoclonal Antibody (111) (Product # MA5-12933, 0.4 ug/ml concentration) and Goat anti-Mouse IgG (H+L), Superclonal™ Recombinant Secondary Antibody, HRP (Product # A28177, 0.25ug/ml, 1:4000 dilution). Densitometric analysis of this western blot is shown in histogram (Fig. b). Decrease in signal upon siRNA mediated knock down confirms that antibody is specific to Ku80.
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
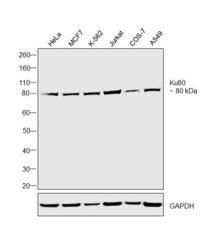

- Experimental details
- Western blot was performed using Anti-Ku80 Monoclonal Antibody (111), (Product # MA5-12933) and a 80 kDa band corresponding to Ku80 was observed in the cell lines tested. Modified whole cell extracts (1% SDS) (30 µg lysate) of HeLa (Lane 1), MCF7 (Lane 2), K-562 (Lane 3), Jurkat (Lane 4), COS-7 (Lane 5) and A549 (Lane 6) were electrophoresed using Novex® NuPAGE® 4-12 % Bis-Tris gel (Product # NP0321BOX). Resolved proteins were then transferred onto a nitrocellulose membrane (Product # IB23001) by iBlot® 2 Dry Blotting System (Product # IB21001). The blot was probed with the primary antibody (1:500 dilution) and detected by chemiluminescence with Goat anti-Mouse IgG (H+L), Superclonal™ Recombinant Secondary Antibody, HRP conjugate (Product # A28177, 1:4000 dilution) using the iBright FL 1000 (Product # A32752). Chemiluminescent detection was performed using Novex® ECL Chemiluminescent Substrate Reagent Kit (Product # WP20005).
Supportive validation
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
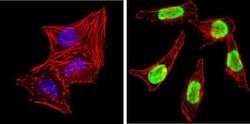
- Experimental details
- Immunofluorescent analysis of Ku (p80) (green) showing staining in the nucleus of HeLa cells. Formalin-fixed cells were permeabilized with 0.1% Triton X-100 in TBS for 5-10 minutes and blocked with 3% BSA-PBS for 30 minutes at room temperature. Cells were probed with a Ku (p80) monoclonal antibody (Product # MA5-12933) in 3% BSA-PBS at a dilution of 1:150 and incubated overnight at 4 ºC in a humidified chamber. Cells were washed with PBST and incubated with a DyLight-conjugated secondary antibody in PBS at room temperature in the dark. F-actin (red) was stained with a fluorescent red phalloidin and nuclei (blue) were stained with Hoechst or DAPI. Images were taken at a magnification of 100x.
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
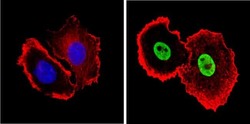
- Experimental details
- Immunofluorescent analysis of Ku (p80) (green) showing staining in the nucleus of MCF-7 cells. Formalin-fixed cells were permeabilized with 0.1% Triton X-100 in TBS for 5-10 minutes and blocked with 3% BSA-PBS for 30 minutes at room temperature. Cells were probed with a Ku (p80) monoclonal antibody (Product # MA5-12933) in 3% BSA-PBS at a dilution of 1:150 and incubated overnight at 4 ºC in a humidified chamber. Cells were washed with PBST and incubated with a DyLight-conjugated secondary antibody in PBS at room temperature in the dark. F-actin (red) was stained with a fluorescent red phalloidin and nuclei (blue) were stained with Hoechst or DAPI. Images were taken at a magnification of 100x.
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
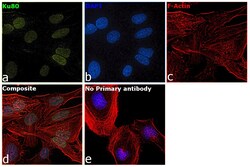
- Experimental details
- Immunofluorescence analysis of Ku80 was performed using 70% confluent log phase HeLa cells. The cells were fixed with 4% paraformaldehyde for 10 minutes, permeabilized with 0.1% Triton™ X-100 for 15 minutes, and blocked with 2% BSA for 1 hour at room temperature. The cells were labeled with Ku80 Monoclonal Antibody (111) (Product # MA5-12933) at 1:200 dilution in 0.1% BSA, incubated at 4 degree celsius overnight and then with Goat anti-Mouse IgG (H+L), Superclonal™ Recombinant Secondary Antibody, Alexa Fluor 488 conjugate (Product # A28175) at a dilution of 1:2000 for 45 minutes at room temperature (Panel a: green). Nuclei (Panel b: blue) were stained with SlowFade® Gold Antifade Mountant with DAPI (Product # S36938). F-actin (Panel c: red) was stained with Rhodamine Phalloidin (Product # R415, 1:300). Panel d represents the merged image showing staining in nucleus. Panel e represents control cells with no primary antibody to assess background. The images were captured at 60X magnification.
Supportive validation
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
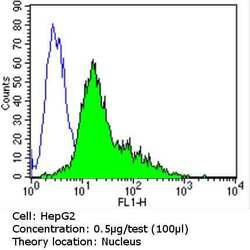
- Experimental details
- Flow cytometry analysis of Ku (p80) in HepG2 cells (green) compared to an isotype control (blue). Cells were harvested, adjusted to a concentration of 1-5x10^6 cells/mL, fixed with 2% paraformaldehyde and washed with PBS. Cells were blocked with a 2% solution of BSA-PBS for 30 min at room temperature and incubated with a Ku (p80) monoclonal antibody (Product # MA5-12933) at a dilution of 0.5 µg/test for 60 min at room temperature. Cells were then incubated for 40 min at room temperature in the dark using a Dylight 488-conjugated goat anti-mouse IgG (H+L) secondary antibody and re-suspended in PBS for FACS analysis.
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
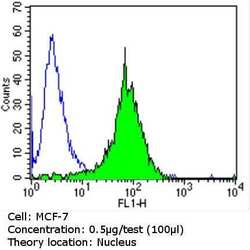
- Experimental details
- Flow cytometry analysis of Ku (p80) in MCF-7 cells (green) compared to an isotype control (blue). Cells were harvested, adjusted to a concentration of 1-5x10^6 cells/mL, fixed with 2% paraformaldehyde and washed with PBS. Cells were blocked with a 2% solution of BSA-PBS for 30 min at room temperature and incubated with a Ku (p80) monoclonal antibody (Product # MA5-12933) at a dilution of 0.5 µg/test for 60 min at room temperature. Cells were then incubated for 40 min at room temperature in the dark using a Dylight 488-conjugated goat anti-mouse IgG (H+L) secondary antibody and re-suspended in PBS for FACS analysis.
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
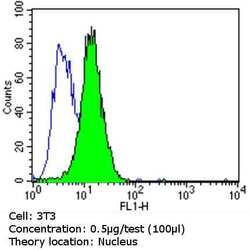
- Experimental details
- Flow cytometry analysis of Ku (p80) in NIH-3T3 cells (green) compared to an isotype control (blue). Cells were harvested, adjusted to a concentration of 1-5x10^6 cells/mL, fixed with 2% paraformaldehyde and washed with PBS. Cells were blocked with a 2% solution of BSA-PBS for 30 min at room temperature and incubated with a Ku (p80) monoclonal antibody (Product # MA5-12933) at a dilution of 0.5 µg/test for 60 min at room temperature. Cells were then incubated for 40 min at room temperature in the dark using a Dylight 488-conjugated goat anti-mouse IgG (H+L) secondary antibody and re-suspended in PBS for FACS analysis.
Supportive validation
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- Immunoprecipitation of Ku (P80) using Ku (P80) Monoclonal Antibody (Product # MA5-12933) on Native Human T47D Cells.
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- Immunoprecipitation of Ku (P80) using Ku (P80) Monoclonal Antibody (Product # MA5-12933) on denatured Human BT474 Cells.
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
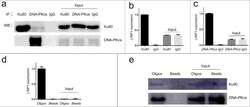
- Experimental details
- Figure 2. LINP1 associates with NHEJ pathway proteins Ku80 and DNA-PKcs in Hela S3 cells a. Immunoprecipitation efficiency of Ku80 and DNA-PKcs specific antibodies detected by western blots. b and c. Results from RNA-IP. QRT-PCR assays showing the relative level of LINP1 in the complex immunoprecipitated by Ku80- and DNA-PKcs-specific antibodies. d. QRT-PCR assays showing the enrichment of LINP1 with specific biotinylated oligos. e. Western blots showing the Ku80 and DNA-PKcs protein levels in the complexes pulled down by LINP1.
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
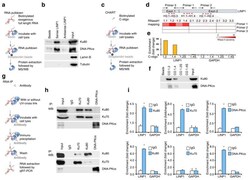
- Experimental details
- Figure 3 LINP1 RNA associates with Ku80 and DNA-PKcs ( a ) A schematic representation of the RNA pull-down assay. ( b ) Western analysis of Ku80 and DNA-PKcs levels in the protein complexes pulled down by either LINP1, antisense LINP1, or empty beads from whole cell extracts of MDA-MB-231 cells. ""Input"" refers to 5% input of whole cell lysates. ( c ) A schematic representation of CHART assay. ( d ) A schematic diagram of LINP1 RNA, the eleven C-oligos (red bars) designed for CHART assay, and the three primer sets (convergent arrows) used in the RNase-H sensitivity assay. RNase-H sensitivity was indicated with blocks with different shades of red, with darker shade indicating higher sensitivity. ( e ) Enrichment of LINP1 with C-oligo 1.2 and 1.4, as measured by qRT-PCR. GAPDH was used as negative control. ( f ) Presence of Ku80 and DNA-PKcs in C-oligo 1.2 and 1.4 enriched complexes, detected by Western analysis. C-oligo 1.2S and 1.4S were used as non-specific controls. ( g ) A schematic representation of an RNA-immunoprecipitation (RNA-IP) assay. ( h ) Results from native RNA-IP (upper panel) and CLIP (lower panel) with Ku80, Ku70, and DNA-PKcs specific antibodies. Western blots showing the immunoprecipitation efficiency of Ku80, Ku70 and DNA-PKcs. ( i ) Results from native RNA-IP (upper panel) and CLIP (lower panel) with Ku80, Ku70, and DNA-PKcs specific antibodies. qRT-PCR assays showing the level of LINP1 and GAPDH (non-specific control) in the co-precipitates. Ku80 (left)
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
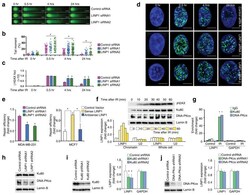
- Experimental details
- LINP1 serves as a modular scaffold in the NHEJ pathway (a) IR-induced DNA damage in control and LINP1 knockdown MDA-MB-231 cells, measured by the comet assay. Scale bar indicates 10 µm. (b) Levels of IR-induced DNA damage, quantified by the tail moment in the comet assay. Two-tailed Student’s t-test; * indicates p-value
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
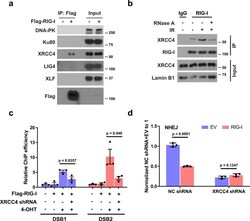
- Experimental details
- Fig. 2 XRCC4 is required for the recruitment of RIG-I to DSB sites. a HEK293T cells were transfected with empty vector or Flag-RIG-I. The cells were then lysed and immunoprecipitated with anti-Flag agarose beads. The beads were boiled and probed with indicated antibodies. b A549 cells were treated with IR (10 Gy, 1-2 h). Cells were lysed, and nuclear fractions were immunoprecipitated with anti-RIG-I antibody. The beads were treated with RNase A, boiled and blotted with indicated antibodies. c Control and XRCC4 knockdown ER-AsiSI U2OS cells were transfected with Flag-RIG-I, and then treated with 4-OHT to induce DSBs. Flag-RIG-I accumulation at DNA damage sites generated by AsiSI was detected by ChIP-qPCR. Data are presented as mean values +- SEM from three independent experiments. P values are determined by unpaired two-sided t -test. d Control and XRCC4 knockdown HEK293T cells overexpressing RIG-I were transfected with NHEJ reporter, and then cells were harvested for the NHEJ assay. Data are presented as mean values +- SEM from three independent experiments. P values are determined by unpaired two-sided t -test.
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
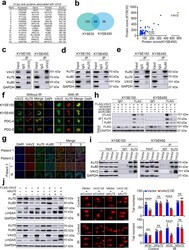
- Fig. 4 VAV2 is necessary in Ku70 and Ku80-mediated DNA NHEJ repair. a Fourteen top proteins potentially associate with VAV2 identified by mass spectrometry in ESCC cells ectopically overexpressing FLAG-tagged VAV2. Cell lysates were immunoprecipitated with antibody against FLAG and antibody against IgG was used as negative control. b Proteins potentially associate with VAV2 in KYSE30 and KYSE450 cells. Left panel is a Venn diagram showing 68 proteins identified in both cell lines. Right panel shows score of each protein. c - e Western blot detection of VAV2, Ku70 and Ku80 proteins by reciprocal immunoprecipitation with antibody against VAV2 ( c ), Ku70 ( d ) or Ku80 ( e ) in KYSE150 and KYSE450 cells. IgG was used as control. f Immunofluorescence analysis of VAV2 and Ku70 co-staining in KYSE150, KYSE450, PDC-4 and PDC-5 cells with or without IR (4 Gy), showing colocalization of VAV2 and Ku70. DAPI was used to label the nucleus. Scale bars, 20 um. g Multiple immunofluorescences analysis of VAV2, Ku70, and Ku80 in clinical ESCC tumor tissues. All tumors (T) show strong colocalization signal of the 3 proteins while the corresponding non-tumor tissues (N) from patient 3, which had low VAV2, show very weak colocalization signal. Merge 1 represents the merge of VAV2 and Ku70, Merge 2 represents the merge of VAV2 and Ku80 and Merge 3 represents the merge of VAV2, Ku70, and Ku80. Scale bars, 100 um. h Western blot analysis of FLAG-VAV2, Ku70 and Ku80 in ESCC cells ectopically express
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
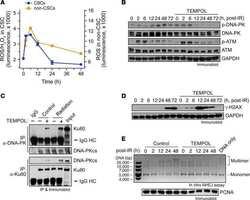
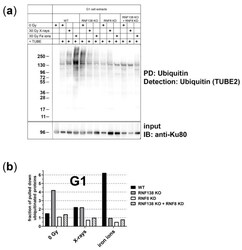
- Protein-pulldown experiments point out that the E3 ubiquitin ligases RNF8 and RNF138 are required for radiation-related ubiquitination in G1 cells . ( a ) HeLa.S-Fucci cells proficient (wt) or deficient for RNF138 (RNF138 KO) and/or RNF8 (RNF8 KD) were irradiated either with 30 Gy X-rays or 30 Gy high-energy iron ions. 1 h after irradiation G1 cells were isolated via flow cytometry, their proteins extracted, and ubiquitinated proteins pulled down and visualized with TUBE2. PD: pull down; IB: immunoblot. ( b ) For quantification of ubiquitinated proteins, western-blot detection of Ku80 in the input-protein extracts (whole-cell extracts that were used for PD, see ( a ) was used for normalization to equal loading. For all conditions, the extracts for the pull-down reactions were performed from 10 6 cells each. Differences in the protein concentration of these extracts are based on the success of cell lysis and reflected in the Ku80 signal. n = 1.
 Explore
Explore Validate
Validate Learn
Learn Western blot
Western blot Immunoprecipitation
Immunoprecipitation