Antibody data
- Antibody Data
- Antigen structure
- References [4]
- Comments [0]
- Validations
- Western blot [5]
- Immunocytochemistry [3]
- Immunohistochemistry [3]
- Other assay [4]
Submit
Validation data
Reference
Comment
Report error
- Product number
- PA5-34941 - Provider product page

- Provider
- Invitrogen Antibodies
- Product name
- GRP78 Polyclonal Antibody
- Antibody type
- Polyclonal
- Antigen
- Synthetic peptide
- Description
- Recommended positive controls: 293T, A431, HeLa, HepG2, NIH-3T-3, JC, BCL-1, C2C12, Raw264.7, Rat heart, Rat Lung, Rat Liver, Rat kidney.
- Concentration
- 0.139 mg/mL
Submitted references Effect of Propionic Acid on Diabetes-Induced Impairment of Unfolded Protein Response Signaling and Astrocyte/Microglia Crosstalk in Rat Ventromedial Nucleus of the Hypothalamus.
Specificity of AMPylation of the human chaperone BiP is mediated by TPR motifs of FICD.
Nuclear receptor binding factor 2 (NRBF2) is required for learning and memory.
EIF3D promotes sunitinib resistance of renal cell carcinoma by interacting with GRP78 and inhibiting its degradation.
Natrus LV, Osadchuk YS, Lisakovska OO, Labudzinskyi DO, Klys YG, Chaikovsky YB
Neural plasticity 2022;2022:6404964
Neural plasticity 2022;2022:6404964
Specificity of AMPylation of the human chaperone BiP is mediated by TPR motifs of FICD.
Fauser J, Gulen B, Pogenberg V, Pett C, Pourjafar-Dehkordi D, Krisp C, Höpfner D, König G, Schlüter H, Feige MJ, Zacharias M, Hedberg C, Itzen A
Nature communications 2021 Apr 23;12(1):2426
Nature communications 2021 Apr 23;12(1):2426
Nuclear receptor binding factor 2 (NRBF2) is required for learning and memory.
Ouyang X, Ahmad I, Johnson MS, Redmann M, Craver J, Wani WY, Benavides GA, Chacko B, Li P, Young M, Jegga AG, Darley-Usmar V, Zhang J
Laboratory investigation; a journal of technical methods and pathology 2020 Sep;100(9):1238-1251
Laboratory investigation; a journal of technical methods and pathology 2020 Sep;100(9):1238-1251
EIF3D promotes sunitinib resistance of renal cell carcinoma by interacting with GRP78 and inhibiting its degradation.
Huang H, Gao Y, Liu A, Yang X, Huang F, Xu L, Danfeng X, Chen L
EBioMedicine 2019 Nov;49:189-201
EBioMedicine 2019 Nov;49:189-201
No comments: Submit comment
Supportive validation
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
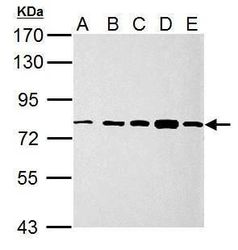
- Experimental details
- Western blot analysis of GRP78/HSPA5 using 30 µg of A) 293T (B) A431 (C) HeLa (D) HepG2 and E) A375 lysate. Samples were loaded onto a 7.5% SDS-PAGE gel and probed with a GRP78/HSPA5 polyclonal antibody (Product # PA5-34941) at a dilution of 1:10,000.
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
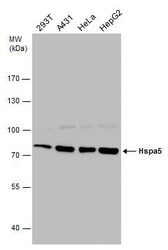
- Experimental details
- Western Blot analysis of GRP78 was performed by separating 30 µg of various whole cell extracts by 7.5% SDS-PAGE. Proteins were transferred to a membrane and probed with a GRP78 Polyclonal Antibody (Product # PA5-34941) at a dilution of 1:10,000.
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
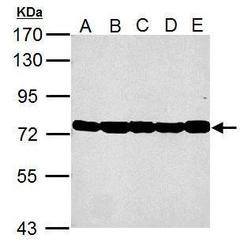
- Experimental details
- Western Blot using GRP78 Polyclonal Antibody (Product # PA5-34941). Sample (30 µg of whole cell lysate). Lane A: NIH-3T-3. Lane B: JC. Lane C: BCL-1. Lane D: C2C12. Lane E: Raw264.7 . 7.5% SDS PAGE. GRP78 Polyclonal Antibody (Product # PA5-34941) diluted at 1:10,000.
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
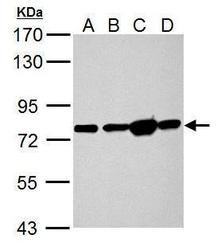
- Experimental details
- Hspa5 antibody detects Hspa5 protein by Western blot analysis. A. 50 µg Rat heart lysate/extract. B. 50 µg Rat Lung lysate/extract. C. 50 µg Rat Liver lysate/extract. D. 50 µg Rat kidney lysate/extract.7.5 % SDS-PAGE. Hspa5 antibody GRP78 Polyclonal Antibody (Product # PA5-34941) dilution: 1:10,000.
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
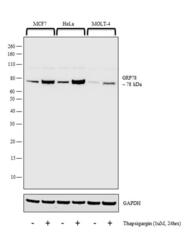
- Experimental details
- Western blot analysis was performed on whole cell extracts (30 µg lysate) of MCF7 (Lane 1), MCF7 treated with Thapsigargin (1uM for 24 hrs) (Lane 2), HeLa (Lane 3), HeLa treated with Thapsigargin (1uM for 24 hrs) (Lane 4), MOLT-4 (Lane 5) and MOLT-4 treated with Thapsigargin (1uM for 24 hrs) (Lane 6). The blot was probed with Anti-GRP78 Polyclonal Antibody (Product # PA5-34941, 1:2000 dilution) and detected by chemiluminescence using Goat anti-Rabbit IgG (H+L) Superclonal™ Secondary Antibody, HRP conjugate (Product # A27036, 0.25 µg/ml, 1:4000 dilution). A 78 kDa band corresponding to GRP78 was observed in all the cell lines tested and enhanced upon treatment with thapsigargin.
Supportive validation
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
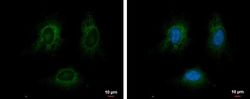
- Experimental details
- Immunofluorescent analysis of GRP78/HSPA5 showing staining in the endoplasmic reticulum of HeLa cells. HeLa cells were fixed in ice-cold MeOH for 5 min and stained using a GRP78/HSPA5 polyclonal antibody (Product # PA5-34941) diluted at 1:500. Blue: Hoechst 33342 staining.
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
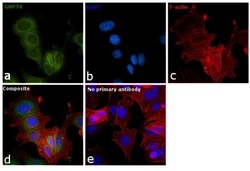
- Experimental details
- Immunofluorescence analysis of GRP78 was performed using 70% confluent log phase MCF7 cells. The cells were fixed with 4% paraformaldehyde for 10 minutes, permeabilized with 0.1% Triton™ X-100 for 15 minutes, and blocked with 1% BSA for 1 hour at room temperature. The cells were labeled with GRP78 Rabbit Polyclonal Antibody (Product # PA5-34941) at 1:200 dilution in 0.1% BSA, incubated at 4 degree celsius overnight and then labeled with Goat anti-Rabbit IgG (H+L) Superclonal™ Secondary Antibody, Alexa Fluor® 488 conjugate (Product # A27034) at a dilution of 1:2000 for 45 minutes at room temperature (Panel a: green). Nuclei (Panel b: blue) were stained with ProLong™ Diamond Antifade Mountant with DAPI (Product # P36962). F-actin (Panel c: red) was stained with Rhodamine Phalloidin (Product # R415, 1:300). Panel d represents the merged image showing Endoplasmic Reticulum localization. Panel e represents control cells with no primary antibody to assess background. The images were captured at 60X magnification.
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
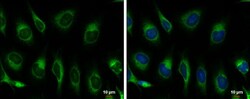
- Experimental details
- Immunocytochemistry-Immunofluorescence analysis of GRP78 was performed in HeLa cells fixed in ice-cold MeOH for 5 min. Green: GRP78 Polyclonal Antibody (Product # PA5-34941) diluted at 1:500. Blue: Hoechst 33342 staining. Scale bar = 10 µm.
Supportive validation
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
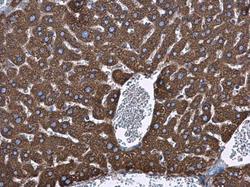
- Experimental details
- Hspa5 antibody detects Hspa5 protein at cytoplasm in mouse liver by immunohistochemical analysis. Sample: Paraffin-embedded mouse liver. Hspa5 antibody (Product # PA5-34941) diluted at 1:500. Antigen Retrieval: Citrate buffer, pH 6.0, 15 min.
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
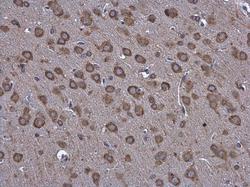
- Experimental details
- Hspa5 antibody detects Hspa5 protein at cytoplasm in rat brain by immunohistochemical analysis. Sample: Paraffin-embedded rat brain. Hspa5 antibody (Product # PA5-34941) diluted at 1:500. Antigen Retrieval: Citrate buffer, pH 6.0, 15 min.
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
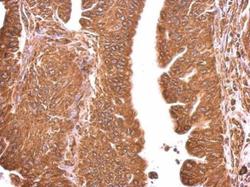
- Experimental details
- Hspa5 antibody detects Hspa5 protein at cytosol on human gastric cancer by immunohistochemical analysis. Sample: Paraffin-embedded gastric cancer. Hspa5 antibody (Product # PA5-34941) dilution: 1:500. Antigen Retrieval: EDTA based buffer, pH 8.0, 15 min.
Supportive validation
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
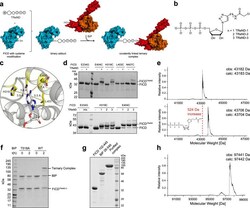
- Experimental details
- Fig. 1 Binary adduct formation and complex formation. a Schematic representation of covalently tethering FICD to BiP via TReNDs. First, the binary adduct is produced with TReNDs. Subsequently, a covalent ternary complex is formed with BiP in an AMPylation reaction. The ribose is displayed as pentagon, the triazole as circle, and the electrophilic moiety as encircled ""+."" b Thiol-reactive nucleotide derivatives (TReNDs) that are used in this study. c Selection of residues (yellow) within FICD suitable for cysteine replacement (based on the structure of FICD E234G:ATP, PDB: 4U07 ). d Reactivity of TReNDs toward FICD 102-458 E234G bearing cysteine substitutions resolved by Phos-Tag TM SDS-PAGE. TReND-mix consists of TReND-1, TReND-2, and TReND-3 at equimolar concentrations. Representative gels are shown from three independent experiments. e Intact mass spectrometry indicating successful reaction of TReND-2 with FICD 102-458 E234G E404C. The mass deviation of unreacted FICD and FICD TReND-2 is -1 and +2 Da, respectively. Intact mass spectrometric data for all binary adducts of FICD 102-458 E234G E404C is shown in Supplementary Fig. 1 . f SDS-PAGE displaying the formation of a covalently linked ternary complex of FICD 102-445 E234G E404C TReND-2 and BiP 19-654. A representative gel is shown from three independent experiments. g SDS-PAGE of the purified ternary complex (using BiP 28-549 T229A and FICD 102-445 T168A T183A E234G L258D E404C) submitted for crystallography. The coval
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
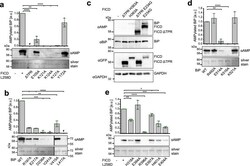
- Fig. 3 Confirmation of the TPR-NBD interface of the FICD TReND -BiP complex and biochemical elucidation of Fic domain-SBD interaction. a AMPylation of BiP 19-654 WT by FICD 102-458 L258D bearing distinct alanine substitutions within the TPR motif. The reaction was stopped after 30 min and the extent of AMPylation quantified via western blot with an alphaAMP-antibody. The experiment was performed in three independent replicates. Data are presented as mean values + /- standard deviation. Compared to FICD L258D, the p values of DeltaTPR, E105A, K121A, K124A, K127A, and Y172A are
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
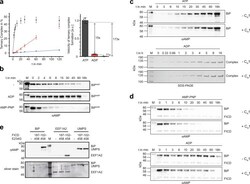
- Fig. 4 Effect of nucleotides, unfolded proteins, and TPR motifs on FICD-mediated AMPylation and deAMPylation. a Ternary complex formation of FICD TReND-2 (102-445 E234G E404C) and BiP 19-654 as readout for the AMPylation of BiP in the presence of ATP (gray) and ADP (red) (both at 0.5 mM) or the absence of any nucleotide (-, blue). The reaction progress was monitored via SDS-PAGE and quantified by densitometric analysis. The experiment was performed in three independent replicates. Data are presented as mean values + /- standard deviation The bar chart represents the relative mean value + /- standard deviation of the linear slope. Full gels are shown in Supplementary Fig. 11 . b FICD 102-458 WT mediated deAMPylation of BiP AMP 19-654 in the presence of ATP and ADP (both at 1 mM) or the absence of any nucleotide (-) was monitored via western blotting with an alphaAMP antibody. c AMPylation of BiP with (+C H 1) and without (-C H 1) the unfolded protein substrate C H 1. AMPylation of BiP in the ATP-bound state of BiP was monitored in an AMPylation assay and analyzed via western blot using the alphaAMP antibody (upper panel). AMPylation of BiP 19-654 WT in its ADP-bound state was monitored via ternary complex formation using FICD 102-445 E234G E404C (lower panel). The full SDS-PAGE gels and the silver-stained western blots are deposited in Supplementary Fig. 13 . d DeAMPylation of BiP was performed with purified BiP AMP in a deAMPylation assay in the presence of 200 uM AMP-PNP (up
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
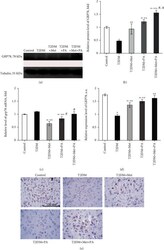
- Figure 6 Effects of administration of metformin and PA on the level of unfolded protein response regulator GRP78/BiP. (a) Immunoblotting analysis of GRP78 in rat VMH: representative immunoblots are shown and quantified using tubulin as a loading control for hypothalamus lysates. (b) The bar graphs of GRP78 are presented as means +- SEM ( n = 6/group). (c) Quantitative polymerase chain reaction (PCR) of Grp78 in rat VMH: data were normalized to beta -actin and pooled from three independent experiments ( n = 6 rats/group). Immunocytochemical analysis of GRP78-positive VMH cells: representative (d) histogram and (e) images are shown. DAB staining was used to visualize GRP78-positive cells; Hematoxylin Gill was used for nuclear staining. Scale bars indicate 150 mu m (magnification x400). All data are shown as means +- SEM; * p < 0.05 vs. control, ** p < 0.05 vs. T2DM, # p < 0.05 vs. metformin administration, and @ p < 0.05 vs. PA administration.
 Explore
Explore Validate
Validate Learn
Learn Western blot
Western blot