MA1-2023
antibody from Invitrogen Antibodies
Targeting: TRIM28
KAP1, PPP1R157, RNF96, TF1B, TIF1B
 Western blot
Western blot Immunocytochemistry
Immunocytochemistry Immunoprecipitation
Immunoprecipitation Gel shift
Gel shift Chromatin Immunoprecipitation
Chromatin Immunoprecipitation Other assay
Other assayAntibody data
- Antibody Data
- Antigen structure
- References [5]
- Comments [0]
- Validations
- Western blot [3]
- Immunocytochemistry [2]
- Other assay [7]
Submit
Validation data
Reference
Comment
Report error
- Product number
- MA1-2023 - Provider product page

- Provider
- Invitrogen Antibodies
- Product name
- TRIM28 Monoclonal Antibody (20C1)
- Antibody type
- Monoclonal
- Antigen
- Recombinant full-length protein
- Description
- MA1-2023 detects human and mouse TIF1 beta. MA1-2023 has been used successfully in Western blot, and immunofluorescence procedures. By Western blot this antibody detects ~100 kDa protein representing TIF1 beta in gamma irradiated HeLa cell lysate. In immunofluorescence procedures, MA1-2023 recognizes TIF1 beta in gamma irradiated HeLa cells. The MA1-2023 immunogen is recombinant protein corresponding to residues 60-383 of the human TIF1 beta protein.
- Reactivity
- Human, Mouse
- Host
- Mouse
- Isotype
- IgG
- Antibody clone number
- 20C1
- Vial size
- 100 µg
- Concentration
- 1 mg/mL
- Storage
- -20° C, Avoid Freeze/Thaw Cycles
Submitted references The Emerging Role of E3 Ubiquitin Ligase SMURF2 in the Regulation of Transcriptional Co-Repressor KAP1 in Untransformed and Cancer Cells and Tissues.
The splicing factor U2AF1 contributes to cancer progression through a noncanonical role in translation regulation.
The L1TD1 protein interactome reveals the importance of post-transcriptional regulation in human pluripotency.
TRIM28 is required by the mouse KRAB domain protein ZFP568 to control convergent extension and morphogenesis of extra-embryonic tissues.
TIF1beta functions as a coactivator for C/EBPbeta and is required for induced differentiation in the myelomonocytic cell line U937.
Shah PA, Boutros-Suleiman S, Emanuelli A, Paolini B, Levy-Cohen G, Blank M
Cancers 2022 Mar 22;14(7)
Cancers 2022 Mar 22;14(7)
The splicing factor U2AF1 contributes to cancer progression through a noncanonical role in translation regulation.
Palangat M, Anastasakis DG, Fei DL, Lindblad KE, Bradley R, Hourigan CS, Hafner M, Larson DR
Genes & development 2019 May 1;33(9-10):482-497
Genes & development 2019 May 1;33(9-10):482-497
The L1TD1 protein interactome reveals the importance of post-transcriptional regulation in human pluripotency.
Emani MR, Närvä E, Stubb A, Chakroborty D, Viitala M, Rokka A, Rahkonen N, Moulder R, Denessiouk K, Trokovic R, Lund R, Elo LL, Lahesmaa R
Stem cell reports 2015 Mar 10;4(3):519-28
Stem cell reports 2015 Mar 10;4(3):519-28
TRIM28 is required by the mouse KRAB domain protein ZFP568 to control convergent extension and morphogenesis of extra-embryonic tissues.
Shibata M, Blauvelt KE, Liem KF Jr, García-García MJ
Development (Cambridge, England) 2011 Dec;138(24):5333-43
Development (Cambridge, England) 2011 Dec;138(24):5333-43
TIF1beta functions as a coactivator for C/EBPbeta and is required for induced differentiation in the myelomonocytic cell line U937.
Rooney JW, Calame KL
Genes & development 2001 Nov 15;15(22):3023-38
Genes & development 2001 Nov 15;15(22):3023-38
No comments: Submit comment
Supportive validation
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- Western blot analysis was performed on nuclear enriched extracts (30 µg lysate) of HEK 293 treated with Hydrogen peroxide (100 µm Hydrogen peroxide for 10 min) (Lane 1), HEK 293 (Lane 2), A549 (Lane 3), HEL 92.1.7 (Lane 4), PC-3 (Lane 5), and Caco-2 (Lane 6). The blots were probed with Anti-TRIM28 Mouse Monoclonal Antibody (Product # MA1-2023, 1 µg/mL) and detected by chemiluminescence using Goat anti-Mouse IgG (H+L) Superclonal™ Secondary Antibody, HRP conjugate (Product # A28177, 0.25 µg/mL, 1:4000 dilution). A 110 kDa band corresponding to TRIM28 was observed across the cell lines tested. Known quantity of protein samples were electrophoresed using Novex® NuPAGE® 4-12 % Bis-Tris gel (Product # NP0321BOX), XCell SureLock™ Electrophoresis System (Product # EI0002) and Novex® Sharp Pre-Stained Protein Standard (Product # LC5800). Resolved proteins were then transferred onto a nitrocellulose membrane with iBlot® 2 Dry Blotting System (Product # IB21001). The membrane was probed with the relevant primary and secondary Antibody following blocking with 5 % skimmed milk. Chemiluminescent detection was performed using Pierce™ ECL Western Blotting Substrate (Product # 32106).
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- Western blot analysis of TRIM28 was performed by loading 10 g of whole cell human BJ fibroblast protein lysate and run on a 4-12% BTE gel. Proteins were transferred to PVDF membrane. Membrane was blocked in 5% non-fat milk in TBST. TRIM28 was detected at approximately 90 kDa using a TRIM28 monoclonal antibody (Product # MA1-2023) at a dilution of 1:500 (2 µg/mL) in 5% milk, followed by a 1:10,000 dilution of anti-mouse HRP. Chemiluminescent detection was performed using SuperSignal West Pico PLUS substrate (Product # 34580). Data courtesy of Antibody Data Exchange Program.
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
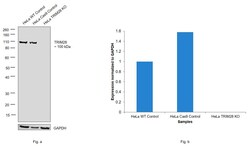
- Experimental details
- Knockout of TRIM28 was achieved by CRISPR-Cas9 genome editing using LentiArray™ Lentiviral sgRNA (Product # A32042, AssayID CRISPR801077_LV) and LentiArray Cas9 Lentivirus (Product # A32064). Western blot analysis of TRIM28 was performed by loading 30 µg of HeLa wild type (Lane 1), HeLa CAS9 (Lane 2) and HeLa TRIM28 KO (Lane 3) modified whole cell extracts. The samples were electrophoresed using Novex® NuPAGE® 4-12% Bis-Tris Protein Gel (Product # NP0321BOX). Resolved proteins were then transferred onto a nitrocellulose membrane (Product # IB23001) by iBlot® 2 Dry Blotting System (Product # IB21001). The blot was probed with Anti-TRIM28 Monoclonal Antibody (20C1) (Product # MA1-2023) at a dilution of 1 µg/mL and and Goat anti-Mouse IgG (H+L), Superclonal™ Recombinant Secondary Antibody, HRP (Product # A28177, 1:4,000 dilution) using the iBright FL 1000 (Product # A32752). Chemiluminescent detection was performed using Novex® ECL Chemiluminescent Substrate Reagent Kit (Product # WP20005). Loss of signal upon CRISPR mediated knockout (KO) using the LentiArray™ CRISPR product line confirms that antibody is specific to TRIM28.
Supportive validation
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
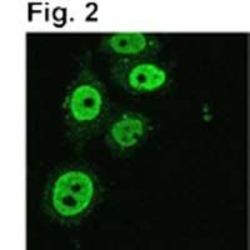
- Experimental details
- Immunofluorescence of TIF1 beta in gamma irradiated HeLa cells using Product # MA1-2023.
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
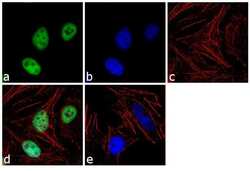
- Experimental details
- Immunofluorescence analysis of TIF1 beta was performed using 70% confluent log phase HeLa cells. The cells were fixed with 4% paraformaldehyde for 10 minutes, permeabilized with 0.1% Triton™ X-100 for 10 minutes, and blocked with 1% BSA for 1 hour at room temperature. The cells were labeled with TRIM28 (20C1) Mouse Monoclonal Antibody (Product # MA1-2023) at 2 µg/mL in 0.1% BSA and incubated for 3 hours at room temperature and then labeled with Goat anti-Mouse IgG (H+L) Superclonal™ Secondary Antibody, Alexa Fluor® 488 conjugate (Product # A28175) at a dilution of 1:2000 for 45 minutes at room temperature (Panel a: green). Nuclei (Panel b: blue) were stained with SlowFade® Gold Antifade Mountant with DAPI (Product # S36938). F-actin (Panel c: red) was stained with Rhodamine Phalloidin (Product # R415, 1:300). Panel d represents the merged image showing nuclear localization. Panel e shows the no primary antibody control. The images were captured at 60X magnification.
Supportive validation
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- Figure 3 SMURF2 positively regulates KAP1 protein levels in tumor cells. ( A ) Western blot analysis showing that overexpression of SMURF2WT, but not its E3 ligase-inactive mutant (SMURF2 CG ), enhances the protein abundance of KAP1 in osteosarcoma U2OS cells. ( B ) Immunoblot analysis showing diminished protein levels of KAP1 in SMURF2 knockdown U2OS cells. Two different siRNAs targeting SMURF2 mRNA expression at either 3'UTR or coding sequence (CDS) were used. NS--non-silencing siRNA. ( C ) Immunoblot analysis of KAP1 expression in SMURF2 CRISPR knockdown U2OS cells ( SMURF2 CR , left panels) and in cells following SMURF2 reconstitution (right panels). SMURF2 CR cells were reconstituted with FLAG-SMURF2. Cells transduced with an empty FLAG vector were used as a control. ( D ) Western blot analysis showing the effect of SMURF2 knockdown on KAP1 protein levels in different types of human cancer cells: cervix carcinoma HeLa cells, prostate carcinoma DU-145 cells, and breast carcinoma MDA-MB-468 cells. Non-silencing siNS and shLuc were used as controls for siRNA and shRNA experiments, respectively. ( E , F ) Confocal microscopy analysis of KAP1 expression in SMURF2 knockdown MDA-MB-468 cells. Scale bars: 20 um. Quantification of the results (panel F) is shown as mean +- SD. n--number of cells quantified for each group from 10 different fields. ( G ) Western blot analysis of KAP1 protein expression in SMURF2 knockdown U2OS cells treated with the proteasomal inhibitor MG-132 (5 u
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- Figure 4 SMURF2 exerts a negative impact on KAP1 expression in untransformed human cells and normal mouse tissues. ( A ) Western blot analyses of KAP1 expression in IMR-90 and BJ-1 cells knocked down of SMURF2. NS--non-silencing siRNA. ( B ) Immunoblot analysis of KAP1 expression in SMURF2 knock-out ( SMURF2 CR ) BJ1 cells. Two different SMURF2 CRISPR clones ( SMURF2 CR C1 and SMURF2 CR C2) were examined. ( C , D ) Confocal microscopy analysis of KAP1 protein expression in SMURF2 CR BJ1 cells. Scale bars: 10 um. Quantification of the results (panel D) is presented as mean +-S D. n--number of cells quantified for each group from at least 10 different fields. ( E ) Western blot analysis of KAP1 expression in Smurf2 KO ( Smurf2 -/- ) and littermate control wild-type ( Smurf2 +/+ ) mouse tissues. ( F ) Quantification of data shown in ( E ) obtained in different pairs of mice. Data are mean +- SD. N--number of animals. ( G ) IHC staining of KAP1 (brown) in Smurf2 WT and KO mouse tissues. The nuclei were counterstained with hematoxylin (blue). Scale bars: 50 um. The uncropped Western blot images can be found in Figure S6 .
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- Analysis of the SMURF2-KAP1 relationship in human normal and cancer tissues. ( A ) Summary of the quantification analysis of SMURF2 and KAP1 staining intensity and percentage of positive cells in a panel of human normal tissues (FDA999m). Note, some of the tissues in this TMA are cancer-adjacent normal tissues. The full details on this TMA are available at https://www.biomax.us/FDA999m , accessed on 22 February 2022. ( B ) SMURF2 and KAP1 expression in different tissue types. Data are mean +- SD. ( C ) Representative IHC images of SMURF2 and KAP1-stained normal tissues. A9 exemplifies tissues in which KAP1 and SMURF2 were equally scored (i.e., cerebellum), and G2 and I3 show differentially scored samples: low KAP1 (intensity score = 2) in stomach, and high KAP1 and low SMURF2 in prostate tissue samples. The nuclei were counterstained with hematoxylin (blue). Scale bars: 50 um. ( D ) Summary of the quantification analysis of SMURF2 and KAP1 staining intensity and %positive cells in human normal and breast cancer TMA (BR804a). ( E ) Representative images of IHC staining of SMURF2 and KAP1 in breast TMA. D3 and D4 are the coordinates of the samples in the tissue array. Scale bars: 50 um. Adj.Normal--adjacent normal tissue. ( F ) The relationship between scoring values of SMURF2 and KAP1 in breast TMA. Data are mean +- SD obtained from the analysis of 40 tumor and corresponding normal tissues [].
 Explore
Explore Validate
Validate Learn
Learn