Antibody data
- Antibody Data
- Antigen structure
- References [0]
- Comments [0]
- Validations
- Flow cytometry [1]
- Other assay [14]
Submit
Validation data
Reference
Comment
Report error
- Product number
- 48-0038-82 - Provider product page

- Provider
- Invitrogen Antibodies
- Product name
- CD3 Monoclonal Antibody (UCHT1), eFluor™ 450, eBioscience™
- Antibody type
- Monoclonal
- Antigen
- Other
- Description
- Description: The UCHT1 monoclonal antibody reacts with human CD3e, a 20 kDa subunit of the TCR complex. Along with the other CD3 subunits gamma and delta, the epsilon chain is required for proper assembly, trafficking and surface expression of the TCR complex. CD3 is expressed by thymocytes in a developmentally regulated manner and by all mature T cells. Crosslinking of TCR via immobilized UCHT1 initiates an intracellular biochemical pathway resulting in cellular activation and proliferation. Applications Reported: This UCHT1 antibody has been reported for use in flow cytometric analysis. Applications Tested: This UCHT1 antibody has been pre-titrated and tested by flow cytometric analysis of normal human peripheral blood cells. This can be used at 5 µL (0.5 µg) per test. A test is defined as the amount (µg) of antibody that will stain a cell sample in a final volume of 100 µL. Cell number should be determined empirically but can range from 10^5 to 10^8 cells/test. eFluor® 450 is a replacement for Pacific Blue®. eFluor® 450 emits at 445 nm and is excited with the Violet laser (405 nm). Please make sure that your instrument is capable of detecting this fluorochome. Excitation: 405 nm; Emission: 445 nm; Laser: Violet Laser. Filtration: 0.2 µm post-manufacturing filtered.
- Reactivity
- Human
- Host
- Mouse
- Isotype
- IgG
- Antibody clone number
- UCHT1
- Vial size
- 100 µg
- Concentration
- 0.2 mg/mL
- Storage
- 4° C, store in dark, DO NOT FREEZE!
No comments: Submit comment
Supportive validation
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
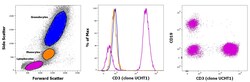

- Experimental details
- Staining of human peripheral blood mononuclear cells with CD45 PE, CD19 APC and CD3 eFluor 450. As expected based on known relative expression patterns, CD3 clone UCHT1 stains a subset of lymphocytes (pink), but not monocytes (orange) and granulocytes (blue).
Supportive validation
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
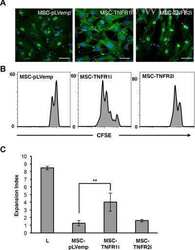
- Fig. 4 T/NK cell clusters in NPC. a t-SNE plot showing 10 clusters of 17,263 T/NK cells (indicated by colors). b t-SNE plot, color coding for the expression of the marker genes (gray to red) for the indicated cell subtypes. c Average expression of selected T cell function-associated genes of naive markers, inhibitory receptors, cytokines and effector molecules, co-stimulatory molecules, and Treg markers in each cell cluster. d Potential developmental trajectory of CD4 + T cells ( n = 5694) inferred by analysis with Monocle 2. Arrows show the increasing directions of certain CD4 + T cell properties annotated with the signatures shown in e . e Traceplots of (left) CD4 + T cell activation signature along activation component and (right) terminal differentiation signature along terminal differentiation component for the CD4 + T cells. Cells are projected along the component, with the blue line indicating the moving average of the expression of signatures (a sliding window of length equal to 5% of the total number of CD4 + T cells was used), and the shaded area displaying SEM. Signatures used are presented in Supplementary information, Table S9 . f Potential developmental trajectory of CD8 + T cells ( n = 6975) inferred by analysis with Monocle 2. Arrows show the increasing directions of certain CD8 + T cell properties annotated with the signatures shown in g . g Traceplots (as in e ) of (left) CD8 + T cell activation signature along activation component and (right) terminal diffe
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
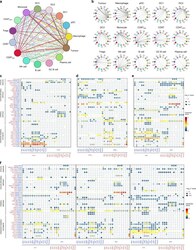
- Fig. 6 The dense network and multiple regulatory immune responses in the TME of NPC. a Capacity for intercellular communication between malignant cells and immune cells. Each line color indicates the ligands expressed by the cell population represented in the same color (labeled). The lines connect to the cell types that express the cognate receptors. The line thickness is proportional to the number of ligands when cognate receptors are present in the recipient cell type. The loops indicate autocrine circuits. The map quantifies potential communication but does not account for the anatomical locations or boundaries of the cell types. b Detailed view of the ligands expressed by each major cell type and the cells expressing the cognate receptors primed to receive the signal. Numbers indicate the quantity of ligand-receptor pairs for each intercellular link. c - f Overview of selected ligand-receptor interactions of tumor cells ( c ), dysfunctional CD8 + T cells ( d ), macrophages ( e ), and the three types of DCs ( f , DC1, DC2, and DC3). P values are indicated by circle size, with the scale to the right (permutation test). The means of the average expression levels of interacting molecule 1 in cluster 1 and interacting molecule 2 in cluster 2 are indicated by color. Assays were carried out at the mRNA level but were used to extrapolate protein interactions. CD4T conv , conventional CD4 + T cell; CD8T, CD8 + T cell; CD8T dys , dysfunctional CD8 + T cell; DC, dendritic cell; GCB
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- Figure 2 T cell ontogenesis in extremely premature infants with and without severe brain injury (A) Principal coordinates analysis (PCoA)--biplot of sequestered cytokine and chemokine composition. Silhouette scoring identified 3 main cyto-Clusters (as indicated by different symbols). The shade of the symbols (gray to black) is determined by days post-delivery--the older the infant the darker the symbol. The size of the symbols is determined by the Kidokoro Score as assessed at term-equivalent age via cMRI--the higher the score the larger the symbol. Significantly correlated (p < 0.05) cytokines/chemokines are plotted as arrows. In addition, box plots next to PCoA show the range of combined values for burst suppression and variance of cranial oxygenation (BS/100 x VAR-cSO 2 ), as well as the range of Kidokoro Scores in the respective cyto-clusters (cyto-cluster 1, CC1; cyto-cluster 2, CC2; cyto-cluster 3, CC3). (B) Blood cytokine/chemokine concentrations in infants with (red) and without (blue) severe BI (3 days post-delivery, d3; 7 days post-delivery, d7; 28 days post-delivery, d28; 32 weeks of gestational age, w32; term-equivalent age = term). (C) Representative images illustrating the gating strategy for FACS analysis. Differently colored gates mark gating for respective cell populations. Magenta, untargeted; blue, T helper cells; orange, cytotoxic T cells; green, T regulatory cells; pink, gammadelta T cells. For box plots in (D) and (F-J), darker shade represents data from
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
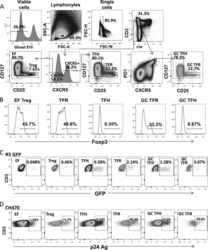
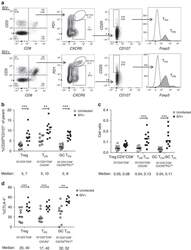
- Figure 2 T FR expansion in lymphoid tissues during chronic SIV infection. ( a ) Disaggregated lymph node and spleen cells from SIV uninfected ( n =9) or chronically SIV-infected rhesus macaques ( n =11) were analysed by flow cytometry. Representative examples of flow cytometry gating are shown. Of viable CD3 + CD8 - cells, follicular subsets were defined as CXCR5 + cells (F) and germinal centre subsets were defined as CXCR5 hi PD-1 hi cells (GC). Of these subsets, regulatory cells were defined as CD25 hi CD127 - . T FR (CXCR5 + CD25 hi CD127 - ) were Foxp3 + , whereas T FH (CXCR5 + CD25 lo/- ) were Foxp3 - . ( b ) The percentages of each rhesus macaque regulatory subset, as analysed in a are shown. ( c ) The ratios of each regulatory cell population to its non-regulatory cell counterpart are shown. ( d ) The percentage of total CTLA-4 expression is shown in SIV-uninfected ( n =9) and chronically SIV-infected ( n =8) rhesus macaques. The horizontal bars of each graph indicate the median value and are listed where appropriate for clarity. Statistical analyses were performed by Mann-Whitney (Wilcoxon) tests to compare unpaired, nonparametric values and significance is denoted by asterisks where * P
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
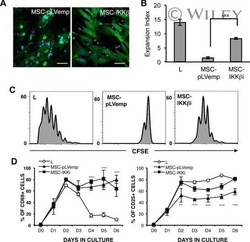
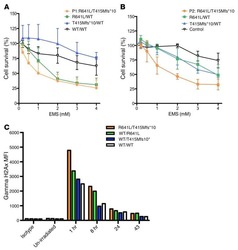
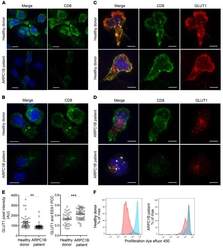
- Figure 7 Absence of ARPC1B alters surface expression of CD8 and GLUT1 in hCTLs. (A-D) HD and ARPC1B-deficient patient hCTLs were fixed in PFA for 20 minutes, permeabilized, and stained with an antibody against CD8 alone (green) ( A and B ) or in combination with anti-GLUT1 (red) and anti-EEA1 (white) antibodies ( C and D ). Images are 3D reconstructions of Z -stack. Scale bars: 4 mum. ( E ) Measurement of the mean intensity of GLUT1 expressed in AU and the degree of colocalization with EEA1 expressed as PCC (see Methods) in HD and ARPC1B-deficient patient hCTLs based on images as sampled in C and D . HD, n = 41 cells; ARPC1B-deficient patient, n = 38 cells. P < 0.005 (unpaired t test). ** P < 0.0013; *** P < 0.0002. ( F ) Flow cytometry analysis of the proliferation capacity of HD and ARPC1B-deficient patient hCTLs (gated on live CD8 + cells) in the absence (blue) or presence (red) of plate-bound anti-CD3 stimulation (1 mug/mL). All data are representative of 3 independent experiments.
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
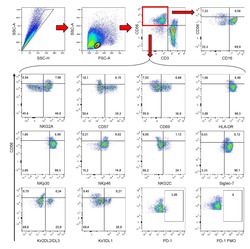
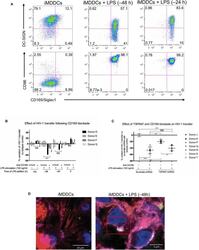
- Figure 5 CD169, as an HIV-1 receptor, mostly impacts transfer from mature MDDCs rather than immature MDDCs. (A) Flow cytometry plots showing CD86, DC-SIGN, and CD169 expression levels on MDDCs (pre-gated on SSC FSC, living cells, CD3 - cells and singlets). Panels show the expression of these proteins in iMDDCs (left panel) and MDDCs with LPS pretreatment at 100 ng/ml for 48 or 24 h before co-culture (middle and right panels, respectively). (B) Percentage of variation of HIV-1 transfer when using iMDDCs or LPS-treated MDDCs (100 ng/ml LPS for different lengths of time) incubated with a blocking antibody against CD169 as compared to an isotype control for each condition. Results are displayed for 4 different blood donors with the mean +- SD of technical triplicates. (C) Percent of variation in HIV-1 transfer to assess the impact of blocking CD169 and TSPAN7 knockdown as compared to scramble shRNA on MDDCs matured with LPS for 48 h treated by an isotype control. Mean +- SD of seven different blood donors in 4 experiments. (B,C) NS, not significant. ** p < 0.01; *** p < 0.001. (D) Confocal microscopy images of iMDDCs (left panel) and mature MDDCs (mMDDCs) right panel, to assess the degree of colocalization between CD169 (magenta) and incoming X4-HIV-1-Gag-iGFP (green). Actin filaments and nuclei were stained with phalloidin (red) and DAPI (blue). Four hundred nanometers of Z-stacks were taken 40 h after the start of the co-culture with CD4 + T cells and X4-HIV-1-Gag-iGFP. The pic
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
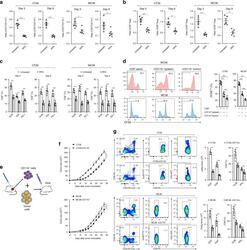
- Fig. 5 Infiltrating myeloid cells inhibit T cell functionality in residual tumors. a Flow cytometric analysis of CD11b + , CD3 + and the ratio of CD3 + /CD11b + in the untreated and iRFA-treated CT26 and MC38 tumors on day 3 and day 9 ( n = 5). b Flow cytometric analysis of CD8 + and CD4 + FoxP3 + cells and the ratio of CD8 + /CD4 + FoxP3 + in the untreated and iRFA-treated CT26 and MC38 tumors on day 3 and day 9 ( n = 5). c Flow cytometric analysis and quantification of granzyme B, IFN-gammaand PD-1 expression in CD8 + cells in the untreated and iRFA-treated CT26 and MC38 tumors on day 3 and day 9 ( n = 5). d In vitro suppressive activity of tumor-infiltrating CD11b + cells purified from spleen or CT26 residual tumors on day 3 after iRFA. Representative histograms of CD8 + T cell proliferation at a ratio of 1:1 CD8 + to CD11b + T cells and percent CD8 + T cell proliferation ( n = 5). e - g CD11b + cells isolated from the residual tumor were mixed with CT26 or MC38 tumor cells and transferred into recipient mice. e Adoptive transfer method. f Growth curve of tumor (one-sided ANOVA test, *** P < 0.001, n = 5). g Flow cytometric analysis and quantification of CD3 + CD8 + cells (gate on single live cells) and Granzyme B expression and IFN-gamma on CD8 + cells ( n = 5). Data represent cumulative results from 1/2 independent experiments with 5 mice/group. The data are represented as mean +- SEM. Statistical differences between pairs of groups
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
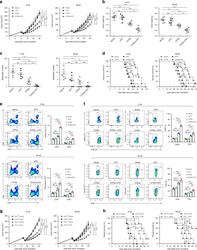
- Fig. 8 CCL2/CCR2 blockade inhibits tumor progression and overcomes resistance to anti-PD-1 therapy. a - f iRFA treatment was performed in CT26 and MC38 colon cancer models as shown in Fig. 2a . Anti-PD-1 mAb (200 mug, clone: J43) was administered through intraperitoneal injection to mice every 3 days for a total of four times. The CCR2 antagonist (CCR2a) (RS504393, Tocris) was given subcutaneously at a dose of 5 mg/kg twice per day for 9 days. a Growth curve of the CT26 and MC38 residual tumor (one-sided ANOVA test, n = 8). b The weight of the residual CT26 and MC38 tumor examined on day 14 after iRFA by dissection of the mice ( n = 6). c The number of metastases examined on day 14 after iRFA by dissection the mice ( n = 6). d Kaplan-Meier survival curves are shown, and the log-rank test was performed ( n = 8). e Flow cytometric analysis and quantification of CD3 + and CD8 + infiltration (gate on single live cells) in residual CT26 tumors. f Granzyme B and IFNgamma expression on CD8 + cells in residual CT26 tumors. (gate on CD8 + cells) ( n = 5). g , h iRFA treatment was performed in mice bearing wild type and CCL2 -/- CT26 or MC38 tumor. g Growth curve of the CT26 and MC38 residual tumor (one-sided ANOVA test, n = 5). h Kaplan-Meier survival curves are shown, and the log-rank test was performed ( n = 8). Data represent results from 1/2 independent experiments. The data are represented as mean +- SEM. Statistical differences between pairs o
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
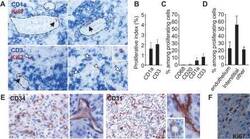
- Figure 1 Proliferating Cells in LCH Granuloma are Mostly Endothelial Cells, Fibroblasts, and T Cells Paraffin-embedded and frozen sections were stained with antibodies against Ki-67 (which label proliferating cells), CD1a (LCs), CD3 (T cells), CD20 (B cells), CD68, CD31, and CD34 (endothelial cells). (A) Double immunostaining of paraffin-embedded section from LCH eosinophilic granulomas with anti-Ki-67 Ab, (brown nuclear staining) and with anti-CD1a Ab (upper images, blue staining) or anti-CD3 Ab (lower images, blue staining). Open arrowheads indicate double-stained cells, black arrowheads indicate Ki-67 + cells with an endothelial morphology. (B) Histogram represents percentage of CD1a + cells and of CD3 + cells labeled with Ki-67 ( n = 15). (C) Histogram represents percentage of proliferating cells (Ki-67 + ) that express CD1a, CD3, CD20, or CD68 ( n = 15). (D) Histogram represents percentage of proliferating cells (Ki-67 + ) that are endothelial cells, interstitial cells (fibroblasts), and other types based on morphological examination. (E) Immunolabeling of blood vessels on paraffin-embedded section from LCH eosinophilic granulomas with CD34 (left) and CD31 (right) antibodies. (F) Proliferating Ki-67 + cells (brown nuclear staining) with a fibroblast-cell morphology in an eosinophilic granuloma.
 Explore
Explore Validate
Validate Learn
Learn Flow cytometry
Flow cytometry