Antibody data
- Antibody Data
- Antigen structure
- References [11]
- Comments [0]
- Validations
- Immunocytochemistry [2]
- Immunoprecipitation [1]
- Immunohistochemistry [3]
- Flow cytometry [1]
- Chromatin Immunoprecipitation [2]
- Other assay [12]
Submit
Validation data
Reference
Comment
Report error
- Product number
- AHO0832 - Provider product page

- Provider
- Invitrogen Antibodies
- Product name
- STAT1 Monoclonal Antibody (STAT1-79)
- Antibody type
- Monoclonal
- Antigen
- Recombinant full-length protein
- Reactivity
- Human, Mouse, Rat
- Host
- Mouse
- Isotype
- IgG
- Antibody clone number
- STAT1-79
- Vial size
- 100 μg
- Concentration
- 0.5 mg/mL
- Storage
- -20°C
Submitted references Inhibition of the IFN-α JAK/STAT Pathway by MERS-CoV and SARS-CoV-1 Proteins in Human Epithelial Cells.
Metabolites Profiling of Melanoma Interstitial Fluids Reveals Uridine Diphosphate as Potent Immune Modulator Capable of Limiting Tumor Growth.
MiR-342 controls Mycobacterium tuberculosis susceptibility by modulating inflammation and cell death.
SARS-CoV-2 suppresses IFNβ production mediated by NSP1, 5, 6, 15, ORF6 and ORF7b but does not suppress the effects of added interferon.
PTPN14 aggravates inflammation through promoting proteasomal degradation of SOCS7 in acute liver failure.
STAT1 expression and HPV16 viral load predict cervical lesion progression.
MicroRNA-325-3p Facilitates Immune Escape of Mycobacterium tuberculosis through Targeting LNX1 via NEK6 Accumulation to Promote Anti-Apoptotic STAT3 Signaling.
Infant High-Grade Gliomas Comprise Multiple Subgroups Characterized by Novel Targetable Gene Fusions and Favorable Outcomes.
LncRNA PLAC2 down-regulates RPL36 expression and blocks cell cycle progression in glioma through a mechanism involving STAT1.
Endothelial IL-33 Expression Is Augmented by Adenoviral Activation of the DNA Damage Machinery.
PDlim2 selectively interacts with the PDZ binding motif of highly pathogenic avian H5N1 influenza A virus NS1.
Zhang Y, Gargan S, Roche FM, Frieman M, Stevenson NJ
Viruses 2022 Mar 23;14(4)
Viruses 2022 Mar 23;14(4)
Metabolites Profiling of Melanoma Interstitial Fluids Reveals Uridine Diphosphate as Potent Immune Modulator Capable of Limiting Tumor Growth.
Vecchio E, Caiazza C, Mimmi S, Avagliano A, Iaccino E, Brusco T, Nisticò N, Maisano D, Aloisio A, Quinto I, Renna M, Divisato G, Romano S, Tufano M, D'Agostino M, Vigliar E, Iaccarino A, Mignogna C, Andreozzi F, Mannino GC, Spiga R, Stornaiuolo M, Arcucci A, Mallardo M, Fiume G
Frontiers in cell and developmental biology 2021;9:730726
Frontiers in cell and developmental biology 2021;9:730726
MiR-342 controls Mycobacterium tuberculosis susceptibility by modulating inflammation and cell death.
Fu B, Lin X, Tan S, Zhang R, Xue W, Zhang H, Zhang S, Zhao Q, Wang Y, Feldman K, Shi L, Zhang S, Nian W, Chaitanya Pavani K, Li Z, Wang X, Wu H
EMBO reports 2021 Sep 6;22(9):e52252
EMBO reports 2021 Sep 6;22(9):e52252
SARS-CoV-2 suppresses IFNβ production mediated by NSP1, 5, 6, 15, ORF6 and ORF7b but does not suppress the effects of added interferon.
Shemesh M, Aktepe TE, Deerain JM, McAuley JL, Audsley MD, David CT, Purcell DFJ, Urin V, Hartmann R, Moseley GW, Mackenzie JM, Schreiber G, Harari D
PLoS pathogens 2021 Aug;17(8):e1009800
PLoS pathogens 2021 Aug;17(8):e1009800
PTPN14 aggravates inflammation through promoting proteasomal degradation of SOCS7 in acute liver failure.
Fu B, Yin S, Lin X, Shi L, Wang Y, Zhang S, Zhao Q, Li Z, Yang Y, Wu H
Cell death & disease 2020 Sep 25;11(9):803
Cell death & disease 2020 Sep 25;11(9):803
STAT1 expression and HPV16 viral load predict cervical lesion progression.
Wu S, Wu Y, Lu Y, Yue Y, Cui C, Yu M, Wang S, Liu M, Zhao Y, Sun Z
Oncology letters 2020 Oct;20(4):28
Oncology letters 2020 Oct;20(4):28
MicroRNA-325-3p Facilitates Immune Escape of Mycobacterium tuberculosis through Targeting LNX1 via NEK6 Accumulation to Promote Anti-Apoptotic STAT3 Signaling.
Fu B, Xue W, Zhang H, Zhang R, Feldman K, Zhao Q, Zhang S, Shi L, Pavani KC, Nian W, Lin X, Wu H
mBio 2020 Jun 2;11(3)
mBio 2020 Jun 2;11(3)
Infant High-Grade Gliomas Comprise Multiple Subgroups Characterized by Novel Targetable Gene Fusions and Favorable Outcomes.
Clarke M, Mackay A, Ismer B, Pickles JC, Tatevossian RG, Newman S, Bale TA, Stoler I, Izquierdo E, Temelso S, Carvalho DM, Molinari V, Burford A, Howell L, Virasami A, Fairchild AR, Avery A, Chalker J, Kristiansen M, Haupfear K, Dalton JD, Orisme W, Wen J, Hubank M, Kurian KM, Rowe C, Maybury M, Crosier S, Knipstein J, Schüller U, Kordes U, Kram DE, Snuderl M, Bridges L, Martin AJ, Doey LJ, Al-Sarraj S, Chandler C, Zebian B, Cairns C, Natrajan R, Boult JKR, Robinson SP, Sill M, Dunkel IJ, Gilheeney SW, Rosenblum MK, Hughes D, Proszek PZ, Macdonald TJ, Preusser M, Haberler C, Slavc I, Packer R, Ng HK, Caspi S, Popović M, Faganel Kotnik B, Wood MD, Baird L, Davare MA, Solomon DA, Olsen TK, Brandal P, Farrell M, Cryan JB, Capra M, Karremann M, Schittenhelm J, Schuhmann MU, Ebinger M, Dinjens WNM, Kerl K, Hettmer S, Pietsch T, Andreiuolo F, Driever PH, Korshunov A, Hiddingh L, Worst BC, Sturm D, Zuckermann M, Witt O, Bloom T, Mitchell C, Miele E, Colafati GS, Diomedi-Camassei F, Bailey S, Moore AS, Hassall TEG, Lowis SP, Tsoli M, Cowley MJ, Ziegler DS, Karajannis MA, Aquilina K, Hargrave DR, Carceller F, Marshall LV, von Deimling A, Kramm CM, Pfister SM, Sahm F, Baker SJ, Mastronuzzi A, Carai A, Vinci M, Capper D, Popov S, Ellison DW, Jacques TS, Jones DTW, Jones C
Cancer discovery 2020 Jul;10(7):942-963
Cancer discovery 2020 Jul;10(7):942-963
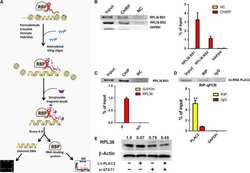
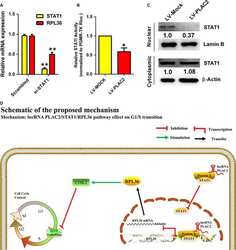
LncRNA PLAC2 down-regulates RPL36 expression and blocks cell cycle progression in glioma through a mechanism involving STAT1.
Hu YW, Kang CM, Zhao JJ, Nie Y, Zheng L, Li HX, Li X, Wang Q, Qiu YR
Journal of cellular and molecular medicine 2018 Jan;22(1):497-510
Journal of cellular and molecular medicine 2018 Jan;22(1):497-510
Endothelial IL-33 Expression Is Augmented by Adenoviral Activation of the DNA Damage Machinery.
Stav-Noraas TE, Edelmann RJ, Poulsen LC, Sundnes O, Phung D, Küchler AM, Müller F, Kamen AA, Haraldsen G, Kaarbø M, Hol J
Journal of immunology (Baltimore, Md. : 1950) 2017 Apr 15;198(8):3318-3325
Journal of immunology (Baltimore, Md. : 1950) 2017 Apr 15;198(8):3318-3325
PDlim2 selectively interacts with the PDZ binding motif of highly pathogenic avian H5N1 influenza A virus NS1.
Yu J, Li X, Wang Y, Li B, Li H, Li Y, Zhou W, Zhang C, Wang Y, Rao Z, Bartlam M, Cao Y
PloS one 2011;6(5):e19511
PloS one 2011;6(5):e19511
No comments: Submit comment
Supportive validation
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
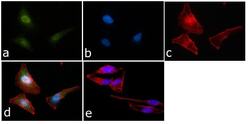
- Experimental details
- Immunofluorescent analysis of STAT-1 was done on 70% confluent log phase HeLa cells. The cells were fixed with 4% paraformaldehyde for 15 minutes, permeabilized with 0.25% Triton™ X-100 for 10 minutes, and blocked with 5% BSA for 1 hour at room temperature. The cells were labeled with STAT1 Antibody (STAT1-79) (Product # AHO0832) at 1:250 dilution in1% BSA and incubated for 3 hours at room temperature and then labeled with Alexa Fluor 488 Rabbit Anti-Mouse IgG Secondary Antibody (Product # A-11059) at a dilution of 1:400 for 45 minutes at room temperature (Panel a: green). Nuclei (Panel b: blue) were stained with SlowFade® Gold Antifade Mountant with DAPI (Product # S36938). F-actin (Panel c: red) was stained with Alexa Fluor 594 Phalloidin (Product # A12381). Panel d is a merged image showing cytoplasmic and Nuclear localization. Panel e is a no primary antibody control. The images were captured at 40X magnification.
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
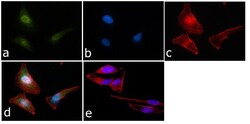
- Experimental details
- Immunofluorescent analysis of STAT-1 was done on 70% confluent log phase HeLa cells. The cells were fixed with 4% paraformaldehyde for 15 minutes, permeabilized with 0.25% Triton™ X-100 for 10 minutes, and blocked with 5% BSA for 1 hour at room temperature. The cells were labeled with STAT1 Antibody (STAT1-79) (Product # AHO0832) at 1:250 dilution in1% BSA and incubated for 3 hours at room temperature and then labeled with Alexa Fluor 488 Rabbit Anti-Mouse IgG Secondary Antibody (Product # A-11059) at a dilution of 1:400 for 45 minutes at room temperature (Panel a: green). Nuclei (Panel b: blue) were stained with SlowFade® Gold Antifade Mountant with DAPI (Product # S36938). F-actin (Panel c: red) was stained with Alexa Fluor 594 Phalloidin (Product # A12381). Panel d is a merged image showing cytoplasmic and Nuclear localization. Panel e is a no primary antibody control. The images were captured at 40X magnification.
Supportive validation
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
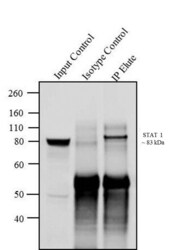
- Experimental details
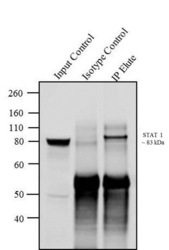
- Immunoprecipitation of STAT1 was performed with 5 µg of the STAT1 Mouse Monoclonal Antibody (Product # AHO0832) on cell extract from MCF7 using the Dynabeads® Protein A Immunoprecipitation Kit (10006D). Normal Mouse IgG was used as a negative IP control. Subsequently, western blot analysis was performed using Novex® NuPAGE® 4-12 % Bis-Tris gel (Product # NP0322BOX), XCell SureLock™ Electrophoresis System (Product # EI0002), Novex® Sharp Pre-Stained Protein Standard (LC5800). Proteins were transferred using iBlot® Dry Blotting System (IB21001) to a nitrocellulose membrane and blocked with 5% skim milk for 1 hour at room temperature on a rocking platform. STAT1 was detected at ~ 83 kDa using STAT1 Mouse Monoclonal Antibody (Product # AHO0832) at 0.5-1 µg/mL in 5 % skim milk for 1 hour at room temperature on a rocking platform. Goat Anti-Mouse IgG- HRP Secondary Antibody (G21234) at 1:5000 dilution was used and chemiluminescent detection was performed using Pierce™ ECL Western Blotting Substrate (Product # 32106). Lane 1 represents 10 % of the total cell extract (input), Lane 2 represents Mouse IgG as Isotype control and Lane 3 represents IP performed with STAT1 Mouse Monoclonal Antibody (Product # AHO0832).
Supportive validation
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
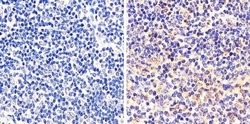
- Experimental details
- Immunohistochemistry analysis of STAT1 showing staining in the cytoplasm and nucleus of paraffin-embedded mouse spleen tissue (right) compared to a negative control without primary antibody (left). To expose target proteins, antigen retrieval was performed using 10mM sodium citrate (pH 6.0), microwaved for 8-15 min. Following antigen retrieval, tissues were blocked in 3% H2O2-methanol for 15 min at room temperature, washed with ddH2O and PBS, and then probed with a STAT1 monoclonal antibody (Product # AHO0832) diluted in 3% BSA-PBS at a dilution of 1:20 overnight at 4ºC in a humidified chamber. Tissues were washed extensively in PBST and detection was performed using an HRP-conjugated secondary antibody followed by colorimetric detection using a DAB kit. Tissues were counterstained with hematoxylin and dehydrated with ethanol and xylene to prep for mounting.
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
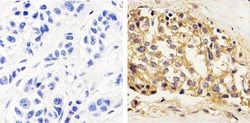
- Experimental details
- Immunohistochemistry analysis of STAT1 showing staining in the cytoplasm and nucleus of paraffin-embedded human breast carcinoma tissue (right) compared to a negative control without primary antibody (left). To expose target proteins, antigen retrieval was performed using 10mM sodium citrate (pH 6.0), microwaved for 8-15 min. Following antigen retrieval, tissues were blocked in 3% H2O2-methanol for 15 min at room temperature, washed with ddH2O and PBS, and then probed with a STAT1 monoclonal antibody (Product # AHO0832) diluted in 3% BSA-PBS at a dilution of 1:20 overnight at 4ºC in a humidified chamber. Tissues were washed extensively in PBST and detection was performed using an HRP-conjugated secondary antibody followed by colorimetric detection using a DAB kit. Tissues were counterstained with hematoxylin and dehydrated with ethanol and xylene to prep for mounting.
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
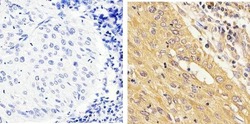
- Experimental details
- Immunohistochemistry analysis of STAT1 showing staining in the cytoplasm and nucleus of paraffin-embedded human cervical carcinoma tissue (right) compared to a negative control without primary antibody (left). To expose target proteins, antigen retrieval was performed using 10mM sodium citrate (pH 6.0), microwaved for 8-15 min. Following antigen retrieval, tissues were blocked in 3% H2O2-methanol for 15 min at room temperature, washed with ddH2O and PBS, and then probed with a STAT1 monoclonal antibody (Product # AHO0832) diluted in 3% BSA-PBS at a dilution of 1:20 overnight at 4ºC in a humidified chamber. Tissues were washed extensively in PBST and detection was performed using an HRP-conjugated secondary antibody followed by colorimetric detection using a DAB kit. Tissues were counterstained with hematoxylin and dehydrated with ethanol and xylene to prep for mounting.
Supportive validation
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
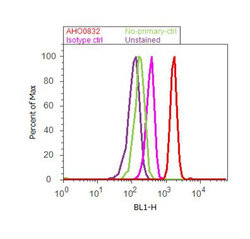
- Experimental details
- Flow cytometry analysis of STAT1 was done on HeLa cells. Cells were fixed with 70% ethanol for 10 minutes, permeabilized with 0.25% Triton™ X-100 for 20 minutes, and blocked with 5% BSA for 30 minutes at room temperature. Cells were labeled with STAT1 Mouse Monoclonal Antibody (AHO0832, red histogram) or with mouse isotype control (pink histogram) at 3-5 ug/million cells in 2.5% BSA. After incubation at room temperature for 2 hours, the cells were labeled with Alexa Fluor® 488 Rabbit Anti-Mouse Secondary Antibody (A11059) at a dilution of 1:400 for 30 minutes at room temperature. The representative 10,000 cells were acquired and analyzed for each sample using an Attune® Acoustic Focusing Cytometer. The purple histogram represents unstained control cells and the green histogram represents no-primary-antibody control.
Supportive validation
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
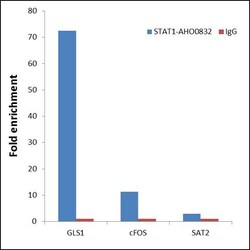
- Experimental details
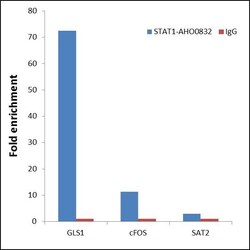
- ChIP- qPCR analysis of STAT1 was performed with 5 µg of the STAT1 Mouse Monoclonal antibody (Product # AHO0832) on sheared chromatin from 2 million HeLa cells treated with 100 ng/mL of IFN gamma for 45 minutes using the MAGnify™ Chromatin Immunoprecipitation System (Product # 49-2024). Normal Mouse IgG was used as a negative IP control. The purified DNA from each ChIP sample was analyzed by StepOnePlus™ Real-Time PCR System (Product # 4376600) with primers for the promoter of active GLS1, cFOS gene, used as positive control target, and the SAT2 used as negative control target. Data is presented as fold enrichment of the antibody signal versus the negative control IgG using the comparative CT method.
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
- ChIP- qPCR analysis of STAT1 was performed with 5 µg of the STAT1 Mouse Monoclonal antibody (Product # AHO0832) on sheared chromatin from 2 million HeLa cells treated with 100 ng/mL of IFN gamma for 45 minutes using the MAGnify™ Chromatin Immunoprecipitation System (Product # 49-2024). Normal Mouse IgG was used as a negative IP control. The purified DNA from each ChIP sample was analyzed by StepOnePlus™ Real-Time PCR System (Product # 4376600) with primers for the promoter of active GLS1, cFOS gene, used as positive control target, and the SAT2 used as negative control target. Data is presented as fold enrichment of the antibody signal versus the negative control IgG using the comparative CT method.
Supportive validation
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
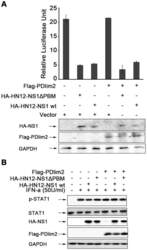
- Figure 6 Effects of the PDlim2 and HN12-NS1 interaction on NF-kappaB activity or STAT1 phosphorylation. (A) HeLa cells was transfected with the reporter plasmid pNF-kappaB-luc and ptk-RL, together with the indicated plasmids. After 24-hour post-transfection, cells were stimulated by LPS (10 ug/ml) for 5 hours. Cells were harvested and firefly and renilla fluorescence unit were measured by luminomitor. The relative luciferase activity (Y-axis) = the unit of firefly luciferase activity / the unit of Renilla luciferase activity (to indicat transfection efficiency, and is presented as the mean calculated from three independent experiments. The expression of NS1 and PDlim2 were analyzed by Western blot with indicated antibodies (lower panels). (B) HeLa cells were transfected with the combination of the plasmids as indicated at the top of the figure and treated by IFN-alpha at 50 U/ml for 4 hours. The cell lysates were analyzed by SDS-PAGE and Western blot, with antibodies specific for phosphorylated STAT1, STAT1, HA-tag, Flag-tag, and GAPDH as loading control.
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
- Immunoprecipitation of STAT1 was performed with 5 µg of the STAT1 Mouse Monoclonal Antibody (Product # AHO0832) on cell extract from MCF7 using the Dynabeads® Protein A Immunoprecipitation Kit (10006D). Normal Mouse IgG was used as a negative IP control. Subsequently, western blot analysis was performed using Novex® NuPAGE® 4-12 % Bis-Tris gel (Product # NP0322BOX), XCell SureLock™ Electrophoresis System (Product # EI0002), Novex® Sharp Pre-Stained Protein Standard (LC5800). Proteins were transferred using iBlot® Dry Blotting System (IB21001) to a nitrocellulose membrane and blocked with 5% skim milk for 1 hour at room temperature on a rocking platform. STAT1 was detected at ~ 83 kDa using STAT1 Mouse Monoclonal Antibody (Product # AHO0832) at 0.5-1 µg/mL in 5 % skim milk for 1 hour at room temperature on a rocking platform. Goat Anti-Mouse IgG- HRP Secondary Antibody (G21234) at 1:5000 dilution was used and chemiluminescent detection was performed using Pierce™ ECL Western Blotting Substrate (Product # 32106). Lane 1 represents 10 % of the total cell extract (input), Lane 2 represents Mouse IgG as Isotype control and Lane 3 represents IP performed with STAT1 Mouse Monoclonal Antibody (Product # AHO0832).
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
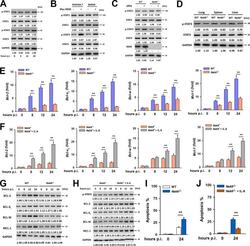
- FIG 5 NEK6 inhibits apoptosis through the activation of STAT3. (A) Immunoblot analysis of the phosphorylated STAT1 (p-STAT1), STAT1, phosphorylated STAT3 (p-STAT3), and STAT3 in RAW 264.7 cells stimulated with 10 mug/mL of gamma-irradiated M. tuberculosis . (B) Immunoblot analysis of the p-STAT1, STAT1, p-STAT3, and STAT3 in RAW 264.7 cells and BMDMs transfected with Myc-NEK6. (C) BMDMs from WT and NEK6-deficient ( Nek6 -/- ) mice were stimulated with 10 mug/mL of gamma-irradiated M. tuberculosis for 24 h. The expressions of p-STAT1, STAT1, p-STAT3, STAT3, and NEK6 were analyzed by Western blotting. (D) Immunoblot analysis of the p-STAT3 and STAT3 in lungs, spleens, and livers from WT and Nek6 -/- mice at 7 dpi. (E and G) The expression levels of BCL-2, BCL-X L , BCL-W, and MCL-1 in 10 mug/mL of gamma-irradiated M. tuberculosis -stimulated BMDMs from WT and Nek6 -/- mice were calculated by qRT-PCR at the indicated times (E) and by Western blotting (G). (F and H) BMDMs from Nek6 -/- mice were pretreated with 20 ng/mL IL-6 for 4 h h, then cells were stimulated with 10 mug/mL of gamma-irradiated M. tuberculosis . The expression levels of BCL-2, BCL-X L , BCL-W, and MCL-1 were calculated by qRT-PCR (F) and Western blotting (H) at the indicated times. (I) BMDMs from WT and Nek6 -/- mice were stimulated with 10 mug/mL of gamma-irradiated M. tuberculosis for 24 h, and the apoptosis rates were detected by flow cytometry. (J) BMDMs from Nek6 -/- mice were pretreated with 20 ng/mL of I
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
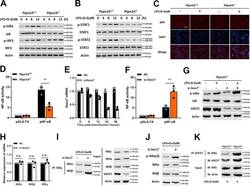
- Fig. 4 SOCS7 blocks the NF-kappaB pathway by interfering with IKK complex assembly. a , b Wild-type, and PTPN14-deficient Hep G2 cells were stimulated with 1 mug/mL LPS and 5 mM D-GalN. Cell lysates were collected at time points as indicated and western blotting was used to detect the phosphorylation levels of IkappaB, IRF-3 ( a ), and STAT1, STAT2 ( b ). c Wild-type and PTPN14-deficient Hep G2 cells were stimulated with 1 mug/mL LPS and 5 mM D-GalN for 8 h. The p65 location was identified by immunofluorescence. Scale bar = 50 mum. d The activation of the NF-kappaB pathway in wild-type and PTPN14-deficient BMDMs. e The knockdown efficiency of Socs7 in PTPN14-deficient BMDMs stimulated with 1 mug/mL LPS and 5 mM D-GalN. f , g PTPN14-deficient BMDMs were transfected with Socs7 siRNA and were stimulated with 1 mug/mL LPS and 5 mM D-GalN for 24 h. The activation of the NF-kappaB pathway ( f ) and the phosphorylation level of IkappaB ( g ) were detected. h PTPN14-deficient BMDMs were transfected with Socs7 siRNA and were stimulated with 1 mug/mL LPS and 5 mM D-GalN for 24 h. The mRNA levels of IKKalpha, IKKbeta, and IKKgamma were tested using qRT-PCR. i , j PTPN14-deficient BMDMs were transfected with Socs7 siRNA and were stimulated with 1 mug/mL LPS and 5 mM D-GalN for 24 h. The interaction between IKKgamma and IKKalpha or IKKbeta ( h ) and the phosphorylation level of IKKalpha/beta ( i ) were detected. k Wild-type and PTPN14-deficient BMDMs were stimulated with 1 mug/mL LPS and
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
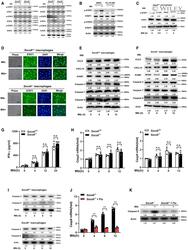
- EV3 Figure SOCS6 has no effects on IFN-gamma, caspase 2, and caspase 9 A Phosphorylation states of STAT family members in response to Mtb stimulation (4 h) in Socs6 +/+ or Socs6 -/- BMDMs were examined by Western blotting. Representative blots from n = 3 biological replicates are shown. B Phosphorylation states of STAT1 and STAT3 in response to Mtb stimulation (4 h) in Socs6 +/+ BMDMs were examined by Western blotting. Fludarabine treatment concentration was 10 muM, and the treatment time was 24 h. Representative blots from n = 3 biological replicates are shown. C Phosphorylation states of STAT1 in response to Mtb stimulation in Socs6 +/+ BMDMs were examined by Western blotting. Representative blots from n = 3 biological replicates are shown. D Intracellular localization of STAT1 in Mtb-stimulated Socs6 +/+ and Socs6 -/- BMDMs were detected by immunofluorescence. Representative images from n = 3 biological replicates are shown. Scale bar = 100 mum. E, F Relative expressions of chemokines CCL5, CXCL10, ICAM1, and caspase 3, caspase 7, caspase 8 in Mtb-stimulated Socs6 +/+ (E) or Socs6 -/- (F) BMDMs were detected by Western blotting. Representative blots from n = 3 biological replicates are shown. G ELISA was performed to detect the secretion of IFN-gamma in Socs6 +/+ and Socs6 -/- BMDMs during Mtb stimulation. Data are shown as the mean +- SEM of n = 3 biological replicates. H, I caspase 2, caspase 9 in Mtb-stimulated Socs6 +/+ or Socs6 -/- BMDMs were detected by qRT-PCR (H, d
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
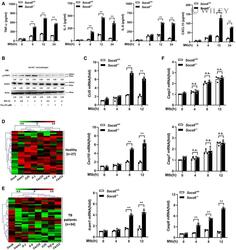
- 5 Figure SOCS6 regulates the expression of chemokines via modulating JAK/STAT signaling A ELISA was performed to detect the secretion of cytokines TNF-alpha, IL-1, IL-6, and CXCL15 in Socs6 +/+ and Socs6 -/- BMDMs during Mtb stimulation. Data are shown as the mean +- SEMof n = 3 biological replicates. B Phosphorylation states of STAT1 in response to Mtb stimulation in Socs6 -/- BMDMs were examined by Western blotting. Fludarabine treatment concentration was 10 muM, and the treatment time was 24 h. Representative blots from n = 3 biological replicates are shown. C-F Relative expressions of chemokines CCL5, CXCL10, ICAM1 (C), and caspase 3, caspase 7, caspase 8 (F) in Mtb-stimulated Socs6 +/+ or Socs6 -/- BMDMs were detected by qRT-PCR. Data are shown as the mean +- SEM of n = 3 biological replicates. Cluster analysis of clinical samples. Heat maps showed relatively high (red) or low (green) expression levels of Socs6 and inflammatory factors in PBMCs from healthy individuals ( n = 27) (D) and TB patients ( n = 34) (E). Data information: ANOVA followed by Bonferroni post hoc test (A, C, F) was used for data analysis. ** P < 0.01. Abbreviation: n .s., not significant. Source data are available online for this figure.
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
- 10.1371/journal.ppat.1009800.g008 Fig 8 SARS-CoV-2 infection reduces total STAT1 levels and subsequent phosphorylation and nuclear translocation of STAT1. Vero cells were infected with SARS-CoV-2 for 24 hours before IFNalpha was exogenously added for 30 minutes prior to analysis. (A) Immunofluorescence (IF) analysis of total STAT1 (green in panels i, iv, vii and x) in mock- (panels i-iii and vii-ix) and CoV-2-infected cells (panels iv-vi and x-xii) detected with anti-dsRNA antibodies (red in panels v and xi) in the absence (panels i-vi) or presence (panels vii-xii) of IFNalpha. (B) Quantification of total STAT1 (whole cell) assessed by IF analysis. Number of cells used are indicated from replicate experiments. Average measurement of fluorescence +/- SEM is depicted. Significance was assessed by students t -test: * p
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
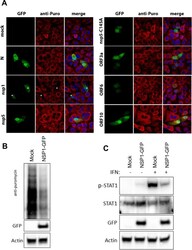
- 10.1371/journal.ppat.1009800.g010 Fig 10 SARS-CoV-2 NSP1 attenuates host cell protein translation restricting the abundance of STAT1. (A) Vero cells were transfected with cDNA expression plasmids encoding vector only (panels i-iii), or the indicated SARS-CoV-2 protein co-expressed with GFP (panels i, iv, vii, x, xiii, xvi, xix, xxii); N (panels iv-vi), NSP1 (panels vii-ix), NSP5 (panels x-xii) NSP5-C145A (panels xiii-xv), ORF3a (panels xvi-xviii), ORF6 (panels xix-xxi) and ORF10 (panels xxii-xxiv). 24 hours after transfection puromycin was exogenously added for 20 minutes prior to detection by anti-puromycin antibodies and AF594 (panels ii, v, viii, xi, xiv, xvii, xx and xxiii) by IF analysis. Merged images are provided in panels iii, vi, ix, xii, xv, xviii, xxi and xxiv. Stars *--NSP1-transfected cells (green) demonstrate a sharp reduction in puromycin signal (red) (B) Western blot showing a decrease in puromycin incorporation during expression of SARS-CoV-2 NSP1 for 24 hours in Vero cells. The incorporated puromycin was detected using anti-puromycin antibodies. Transfected cells were detected using anti-GFP antibodies and anti-actin antibodies served as a loading control. (C) Western blot depicting the abundance of both STAT1 and pSTAT1 in mock- and NSP1-transfected Vero cells after exogenous addition of IFNalpha for 30 minutes. Transfected cells were detected using anti-GFP antibodies. Anti-actin antibodies served as loading control.
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
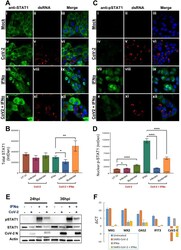
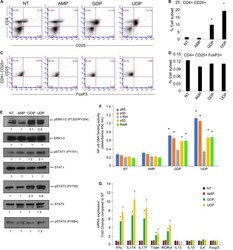
- FIGURE 2 GDP and UDP modulate immune response of CD4 + T-cells. (A) PBMCs (1 x 10 6 ) were treated with 30 muM of indicated metabolites for 24 h and were stained with CD4-FITC and CD25-PE. Each plot represents 20,000 events of a representative experiment. (B) Percentage of CD4 + CD25 + population in the different conditions. Values (mean +- SE, n = 6) are shown. The asterisk indicates a statistically significant difference compared to untreated control, according to Student's t -test ( p < 0.01). (C) PBMCs stained with CD4-FITC and CD25-PE were permeabilized and next stained with FoxP3-APC. Each plot represents 20,000 events of a representative experiment. (D) Percentage of CD4 + CD25 + FoxP3 + population in the different conditions. Values (mean +- SE, n = 6) are shown. The asterisk indicates a statistically significant difference compared to untreated control, according to Student's t -test ( p < 0.01). (E) CD4 + T-cells, isolated from PBMCs of healthy donors (5 x 10 6 ), were treated with 30 muM of indicated metabolites for 2 h and then were lysed. Total protein extracts (30 mug) were separated by 4-12% gradient NuPAGE and analyzed by Western blotting with anti-pERK1/2 (phospho-tyrosine 202/204), anti-ERK1/2, anti-pSTAT1 (phospho-tyrosine 701), anti-STAT1, anti-pSTAT3 (phospho-tyrosine 705), anti-STAT3, and anti-pSTAT5 (phospho-tyrosine 694). (F) CD4 + T-cells, isolated from PBMCs of healthy donors (5 x 10 6 ), were treated with 30 muM of indicated metabolites for 2 h, and
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
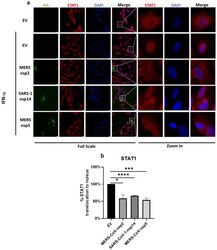
- Figure 6 MERS-CoV-nsp2, SARS-CoV-1-nsp14, and MERS-CoV-nsp5 expression block IFN-alpha-mediated STAT1 nuclear translocation of BEAS-2B cells. ( a ) Confocal microscopy images of BEAS-2B cells transfected with Empty Vector (EV) or HA-tagged MERS-CoV-nsp2, SARS-CoV-1-nsp14, or MERS-CoV-nsp5. After 24 h, cells were stimulated for 30 min with IFN-alpha (1000 U/mL) then fixed and permeabilised. Anti-HA and anti-STAT1 were probed as primary antibodies, and anti-CF(tm) 568 and anti-Alexa Fluor 647 were probed as secondary antibodies to monitor HA expression and STAT1 subcellular localisation. Nuclei were stained by DAPI. Each full scaled image was then zoomed in for presenting single transfected cells. ( b ) Quantification of nuclear translocation of STAT1 in control, MERS-CoV-nsp2, SARS-CoV-1-nsp14, and MERS-CoV-nsp5-positive cells from three fields of view on each coverslip collected from three independent experiments. The level of STAT1 translocation to nucleus for IFN-alpha-treated EV was normalised to 100%. Scale bar, 20 mum. All graphs are the mean +- SEM of three independent experiments * p < 0.05, *** p < 0.01, **** p < 0.0001 (Student's t test).
 Explore
Explore Validate
Validate Learn
Learn Western blot
Western blot Immunocytochemistry
Immunocytochemistry