Antibody data
- Antibody Data
- Antigen structure
- References [23]
- Comments [0]
- Validations
- Flow cytometry [2]
- Other assay [6]
Submit
Validation data
Reference
Comment
Report error
- Product number
- 12-0458-41 - Provider product page

- Provider
- Invitrogen Antibodies
- Product name
- CD45RA Monoclonal Antibody (HI100), PE, eBioscience™
- Antibody type
- Monoclonal
- Antigen
- Other
- Description
- Description: The HI100 monoclonal antibody reacts with human CD45RA, a 220 kDa molecule expressed by subpopulations of CD4+ peripheral T lymphocytes, CD8+ peripheral T lymphocytes, and B cells. The CD45RA+ T cell populations are mainly naive/virgin allowing the use of HI100 mAb as a phenotypic marker to discriminate T cell subsets. Applications Reported: The HI100 antibody has been reported for use in flow cytometric analysis. Applications Tested: This HI100 antibody has been pre-titrated and tested by flow cytometric analysis of normal human peripheral blood cells. This can be used at 5 µL (0.06 µg) per test. A test is defined as the amount (µg) of antibody that will stain a cell sample in a final volume of 100 µL. Cell number should be determined empirically but can range from 10^5 to 10^8 cells/test. Excitation: 488-561 nm; Emission: 578 nm; Laser: Blue Laser, Green Laser, Yellow-Green Laser. Filtration: 0.2 µm post-manufacturing filtered.
- Reactivity
- Human
- Host
- Mouse
- Conjugate
- Yellow dye
- Isotype
- IgG
- Antibody clone number
- HI100
- Vial size
- 25 Tests
- Concentration
- 5 µL/Test
- Storage
- 4° C, store in dark, DO NOT FREEZE!
Submitted references Specific transcriptional programs differentiate ICOS from CD28 costimulatory signaling in human Naïve CD4(+) T cells.
Differences in Maturation Status and Immune Phenotypes of Circulating Helios(+) and Helios(-) Tregs and Their Disrupted Correlations With Monocyte Subsets in Autoantibody-Positive T1D Individuals.
Heterogeneous disease-propagating stem cells in juvenile myelomonocytic leukemia.
Single-Cell Analyses Reveal Megakaryocyte-Biased Hematopoiesis in Myelofibrosis and Identify Mutant Clone-Specific Targets.
TARGET-Seq: A Protocol for High-Sensitivity Single-Cell Mutational Analysis and Parallel RNA Sequencing.
The downregulated membrane expression of CD18 in CD34(+) cells defines a primitive population of human hematopoietic stem cells.
MicroRNA‑155 inhibits the proliferation of CD8+ T cells via upregulating regulatory T cells in vitiligo.
Targeting enhancer switching overcomes non-genetic drug resistance in acute myeloid leukaemia.
TIGIT expressing CD4+T cells represent a tumor-supportive T cell subset in chronic lymphocytic leukemia.
Targeting a CAR to the TRAC locus with CRISPR/Cas9 enhances tumour rejection.
The distribution and function of human memory T cell subsets in lung cancer.
Heterogeneous leukemia stem cells in myeloid blast phase chronic myeloid leukemia.
Circulating precursors of human CD1c+ and CD141+ dendritic cells.
Restricted dendritic cell and monocyte progenitors in human cord blood and bone marrow.
Skewed T-helper (Th)1/2- and Th17/T regulatory‑cell balances in patients with renal cell carcinoma.
The acute environment, rather than T cell subset pre-commitment, regulates expression of the human T cell cytokine amphiregulin.
Proportions of CD4+ memory T cells are altered in individuals chronically infected with Schistosoma haematobium.
Programmed death-1+ T cells and regulatory T cells are enriched in tumor-involved lymph nodes and associated with aggressive features in papillary thyroid cancer.
RUNX transcription factor-mediated association of Cd4 and Cd8 enables coordinate gene regulation.
Generation of bivalent chromatin domains during cell fate decisions.
IL-15 transpresentation promotes both human T-cell reconstitution and T-cell-dependent antibody responses in vivo.
Dendritic cells express hematopoietic prostaglandin D synthase and function as a source of prostaglandin D2 in the skin.
Naive CD4 t cell proliferation is controlled by mammalian target of rapamycin regulation of GRAIL expression.
Gigliotti CL, Boggio E, Favero F, Incarnato D, Santoro C, Oliviero S, Rojo JM, Zucchelli S, Persichetti F, Baldanzi G, Dianzani U, Corà D
Frontiers in immunology 2022;13:915963
Frontiers in immunology 2022;13:915963
Differences in Maturation Status and Immune Phenotypes of Circulating Helios(+) and Helios(-) Tregs and Their Disrupted Correlations With Monocyte Subsets in Autoantibody-Positive T1D Individuals.
Zhang Y, Zhang J, Shi Y, Shen M, Lv H, Chen S, Feng Y, Chen H, Xu X, Yang T, Xu K
Frontiers in immunology 2021;12:628504
Frontiers in immunology 2021;12:628504
Heterogeneous disease-propagating stem cells in juvenile myelomonocytic leukemia.
Louka E, Povinelli B, Rodriguez-Meira A, Buck G, Wen WX, Wang G, Sousos N, Ashley N, Hamblin A, Booth CAG, Roy A, Elliott N, Iskander D, de la Fuente J, Fordham N, O'Byrne S, Inglott S, Norfo R, Salio M, Thongjuea S, Rao A, Roberts I, Mead AJ
The Journal of experimental medicine 2021 Feb 1;218(2)
The Journal of experimental medicine 2021 Feb 1;218(2)
Single-Cell Analyses Reveal Megakaryocyte-Biased Hematopoiesis in Myelofibrosis and Identify Mutant Clone-Specific Targets.
Psaila B, Wang G, Rodriguez-Meira A, Li R, Heuston EF, Murphy L, Yee D, Hitchcock IS, Sousos N, O'Sullivan J, Anderson S, Senis YA, Weinberg OK, Calicchio ML, NIH Intramural Sequencing Center, Iskander D, Royston D, Milojkovic D, Roberts I, Bodine DM, Thongjuea S, Mead AJ
Molecular cell 2020 May 7;78(3):477-492.e8
Molecular cell 2020 May 7;78(3):477-492.e8
TARGET-Seq: A Protocol for High-Sensitivity Single-Cell Mutational Analysis and Parallel RNA Sequencing.
Rodriguez-Meira A, O'Sullivan J, Rahman H, Mead AJ
STAR protocols 2020 Dec 18;1(3):100125
STAR protocols 2020 Dec 18;1(3):100125
The downregulated membrane expression of CD18 in CD34(+) cells defines a primitive population of human hematopoietic stem cells.
Mesa-Núñez C, Leon-Rico D, Aldea M, Damián C, Sanchez-Baltasar R, Sanchez R, Alberquilla O, Segovia JC, Bueren JA, Almarza E
Stem cell research & therapy 2020 Apr 28;11(1):164
Stem cell research & therapy 2020 Apr 28;11(1):164
MicroRNA‑155 inhibits the proliferation of CD8+ T cells via upregulating regulatory T cells in vitiligo.
Lv M, Li Z, Liu J, Lin F, Zhang Q, Li Z, Wang Y, Wang K, Xu Y
Molecular medicine reports 2019 Oct;20(4):3617-3624
Molecular medicine reports 2019 Oct;20(4):3617-3624
Targeting enhancer switching overcomes non-genetic drug resistance in acute myeloid leukaemia.
Bell CC, Fennell KA, Chan YC, Rambow F, Yeung MM, Vassiliadis D, Lara L, Yeh P, Martelotto LG, Rogiers A, Kremer BE, Barbash O, Mohammad HP, Johanson TM, Burr ML, Dhar A, Karpinich N, Tian L, Tyler DS, MacPherson L, Shi J, Pinnawala N, Yew Fong C, Papenfuss AT, Grimmond SM, Dawson SJ, Allan RS, Kruger RG, Vakoc CR, Goode DL, Naik SH, Gilan O, Lam EYN, Marine JC, Prinjha RK, Dawson MA
Nature communications 2019 Jun 20;10(1):2723
Nature communications 2019 Jun 20;10(1):2723
TIGIT expressing CD4+T cells represent a tumor-supportive T cell subset in chronic lymphocytic leukemia.
Catakovic K, Gassner FJ, Ratswohl C, Zaborsky N, Rebhandl S, Schubert M, Steiner M, Gutjahr JC, Pleyer L, Egle A, Hartmann TN, Greil R, Geisberger R
Oncoimmunology 2017;7(1):e1371399
Oncoimmunology 2017;7(1):e1371399
Targeting a CAR to the TRAC locus with CRISPR/Cas9 enhances tumour rejection.
Eyquem J, Mansilla-Soto J, Giavridis T, van der Stegen SJ, Hamieh M, Cunanan KM, Odak A, Gönen M, Sadelain M
Nature 2017 Mar 2;543(7643):113-117
Nature 2017 Mar 2;543(7643):113-117
The distribution and function of human memory T cell subsets in lung cancer.
Sheng SY, Gu Y, Lu CG, Zou JY, Hong H, Wang R
Immunologic research 2017 Jun;65(3):639-650
Immunologic research 2017 Jun;65(3):639-650
Heterogeneous leukemia stem cells in myeloid blast phase chronic myeloid leukemia.
Kinstrie R, Karamitros D, Goardon N, Morrison H, Hamblin M, Robinson L, Clark RE, Copland M, Vyas P
Blood advances 2016 Dec 27;1(3):160-169
Blood advances 2016 Dec 27;1(3):160-169
Circulating precursors of human CD1c+ and CD141+ dendritic cells.
Breton G, Lee J, Zhou YJ, Schreiber JJ, Keler T, Puhr S, Anandasabapathy N, Schlesinger S, Caskey M, Liu K, Nussenzweig MC
The Journal of experimental medicine 2015 Mar 9;212(3):401-13
The Journal of experimental medicine 2015 Mar 9;212(3):401-13
Restricted dendritic cell and monocyte progenitors in human cord blood and bone marrow.
Lee J, Breton G, Oliveira TY, Zhou YJ, Aljoufi A, Puhr S, Cameron MJ, Sékaly RP, Nussenzweig MC, Liu K
The Journal of experimental medicine 2015 Mar 9;212(3):385-99
The Journal of experimental medicine 2015 Mar 9;212(3):385-99
Skewed T-helper (Th)1/2- and Th17/T regulatory‑cell balances in patients with renal cell carcinoma.
Li L, Yang C, Zhao Z, Xu B, Zheng M, Zhang C, Min Z, Guo J, Rong R
Molecular medicine reports 2015 Feb;11(2):947-53
Molecular medicine reports 2015 Feb;11(2):947-53
The acute environment, rather than T cell subset pre-commitment, regulates expression of the human T cell cytokine amphiregulin.
Qi Y, Operario DJ, Georas SN, Mosmann TR
PloS one 2012;7(6):e39072
PloS one 2012;7(6):e39072
Proportions of CD4+ memory T cells are altered in individuals chronically infected with Schistosoma haematobium.
Nausch N, Bourke CD, Appleby LJ, Rujeni N, Lantz O, Trottein F, Midzi N, Mduluza T, Mutapi F
Scientific reports 2012;2:472
Scientific reports 2012;2:472
Programmed death-1+ T cells and regulatory T cells are enriched in tumor-involved lymph nodes and associated with aggressive features in papillary thyroid cancer.
French JD, Kotnis GR, Said S, Raeburn CD, McIntyre RC Jr, Klopper JP, Haugen BR
The Journal of clinical endocrinology and metabolism 2012 Jun;97(6):E934-43
The Journal of clinical endocrinology and metabolism 2012 Jun;97(6):E934-43
RUNX transcription factor-mediated association of Cd4 and Cd8 enables coordinate gene regulation.
Collins A, Hewitt SL, Chaumeil J, Sellars M, Micsinai M, Allinne J, Parisi F, Nora EP, Bolland DJ, Corcoran AE, Kluger Y, Bosselut R, Ellmeier W, Chong MM, Littman DR, Skok JA
Immunity 2011 Mar 25;34(3):303-14
Immunity 2011 Mar 25;34(3):303-14
Generation of bivalent chromatin domains during cell fate decisions.
De Gobbi M, Garrick D, Lynch M, Vernimmen D, Hughes JR, Goardon N, Luc S, Lower KM, Sloane-Stanley JA, Pina C, Soneji S, Renella R, Enver T, Taylor S, Jacobsen SE, Vyas P, Gibbons RJ, Higgs DR
Epigenetics & chromatin 2011 Jun 6;4(1):9
Epigenetics & chromatin 2011 Jun 6;4(1):9
IL-15 transpresentation promotes both human T-cell reconstitution and T-cell-dependent antibody responses in vivo.
Huntington ND, Alves NL, Legrand N, Lim A, Strick-Marchand H, Mention JJ, Plet A, Weijer K, Jacques Y, Becker PD, Guzman C, Soussan P, Kremsdorf D, Spits H, Di Santo JP
Proceedings of the National Academy of Sciences of the United States of America 2011 Apr 12;108(15):6217-22
Proceedings of the National Academy of Sciences of the United States of America 2011 Apr 12;108(15):6217-22
Dendritic cells express hematopoietic prostaglandin D synthase and function as a source of prostaglandin D2 in the skin.
Shimura C, Satoh T, Igawa K, Aritake K, Urade Y, Nakamura M, Yokozeki H
The American journal of pathology 2010 Jan;176(1):227-37
The American journal of pathology 2010 Jan;176(1):227-37
Naive CD4 t cell proliferation is controlled by mammalian target of rapamycin regulation of GRAIL expression.
Lin JT, Lineberry NB, Kattah MG, Su LL, Utz PJ, Fathman CG, Wu L
Journal of immunology (Baltimore, Md. : 1950) 2009 May 15;182(10):5919-28
Journal of immunology (Baltimore, Md. : 1950) 2009 May 15;182(10):5919-28
No comments: Submit comment
Supportive validation
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
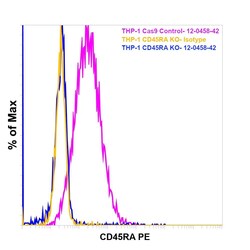
- Experimental details
- Knockout of CD45RA was achieved by CRISPR-Cas9 genome editing using LentiArray™ Lentiviral sgRNA (Product # A32042, Assay ID CRISPR664203_LV) and LentiArray Cas9 Lentivirus (Product # A32064). Flow cytometry analysis of CD45RA was performed by staining THP-1 CD45RA Knock out cells with 0.06 µg Mouse IgG2b kappa Isotype Control (eBMG2b), PE, eBioscience™ (Product # 12-4732-81, yellow histogram) or 0.06 µg CD45RA Monoclonal Antibody (HI100), PE, eBioscience™ (Product # 12-0458-42, blue histogram) in the presence of Fc Receptor Binding Inhibitor Polyclonal Antibody, eBioscience™ (Product # 14-9161-73). THP-1 Cas9 control cells were also stained similarly with0.06 µg CD45RA Monoclonal Antibody (HI100), PE, eBioscience™ (Product # 12-0458-42, pink histogram). Lossof signal was observed in the CD45RA KOcells stained with CD45RA antibody clone HI100 but not in the control Cas9cells. Viable cells were used for analysis, as determined by Fixable Viability DyeeFluor™780 (Product # 65-0865-18).
- Conjugate
- Yellow dye
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- Staining of normal human peripheral blood cells with Anti-Human CD45RO PerCP-eFluor® 710 (Product # 46-0457-42) and Mouse IgG2b K Isotype Control PE (Product # 12-4732-81) (left) or Anti-Human CD45RA PE (right). Cells in the lymphocyte gate were used for analysis.
- Conjugate
- Yellow dye
Supportive validation
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
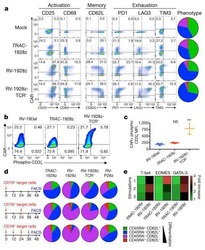
- Experimental details
- NULL
- Conjugate
- Yellow dye
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
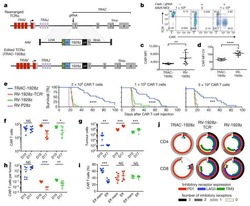
- Experimental details
- NULL
- Conjugate
- Yellow dye
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
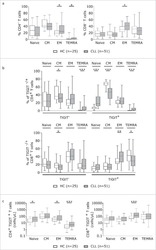
- Experimental details
- NULL
- Conjugate
- Yellow dye
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
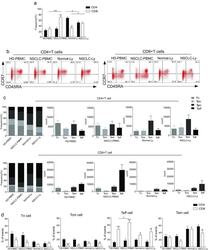
- Experimental details
- Fig. 1 The distribution of CD4+ and CD8+ T cells subsets in human lung cancer. PBMCs were isolated from the blood of lung cancer patients and healthy donors and analyzed by flow cytometry. a The frequency of the CD3+CD4+ T cells and CD3+CD8+ T cells in the HD-PBMC, PBMCs from healthy donors; NSCLC-PBMC, PBMCs from non-small lung cancer patients, Normal-Ly, from healthy lymph node, NSCLC-Ly, tumor infiltrated lymph node from non-small lung cancer patients. b Representative flow cytometric analyses of CD45RA and CCR7 expression in CD3+CD4+ T cells and CD3+CD8+ T cells, indicating naive T cells (CD45RA+/CD45RO-CCR7+, top right quadrant ), terminal effector T cells (CD45RA+/CD45RO-CCR7-, bottom right quadrant ), central memory T cells (Tcm, CD45RO+/CD45RA-CCR7+, top left quadrant ), and effector memory T cells (Tem, CD45RO+/CD45RA-CCR7-, bottom left quadrant ), gated on the forward and side scatter of the lymphocyte populations. c The frequency and absolute number of the CD4+ ( top ) and CD8+ ( bottom ), Tn ( middle gray ), Teff ( black ), Tcm ( grey ), and Tem ( dark grey ) cell subsets in the blood from the non small cell lung cancer patients and healthy donors. d The events of Tn, Teff, Tcm and Tem cell subsets of CD4+ and CD8+ cells in the blood from non small cell lung cancer patients and healthy donors, expressed as the mean +- SEM. * p < 0.05; ** p < 0.005; *** p < 0.001; Mann-Whitney test (two-tailed) and non-paired Student's t-test
- Conjugate
- Yellow dye
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
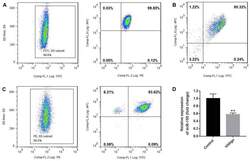
- Experimental details
- Figure 1. Purity of CD3 + CD4 + CD45RA + T cells, CD3 + CD8 + T cells and CD4 + CD25 + FoxP3 + Treg cells. CD3 + CD4 + CD45RA + T cells and CD3 + CD8 + T cells were purified by magnetic cell sorting, and their purity was determined by flow cytometry. (A) The purity of CD3 + CD4 + CD45RA + T cells was 99.45% (CD3 + T cells, 99.6%; CD4 + CD45RA + T cells, 99.85%). (B) The purity of CD3 + CD8 + T cells was 95.32%. (C) The purity of CD4 + CD25 + FoxP3 + Treg cells was 93.15% (CD4 + T cells, 99.5%; CD25 + FoxP3 + T cells, 93.62%). (D) miR-155 expression in T cells of the patients with vitiligo and healthy donor was detected by reverse transcription quantitative polymerase chain reaction. **P
- Conjugate
- Yellow dye
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
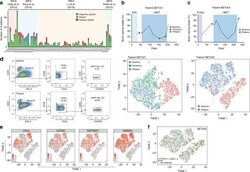
- Experimental details
- Fig. 1 Non-genetic adaptation drives clinical resistance in AML. a Meta-analysis from four independent studies analysing either the whole genome or whole exome of AML patients at diagnosis and relapse. Mutations are defined as non-synonymous changes within the coding sequence of any gene. Shared mutations are mutations present at both diagnosis and relapse. Whole exome sequencing data from Li et al. (REF 4 ) was analysed to access the mutations in known AML genes, as defined by the authors. b Schematic of the treatment regime and bone marrow blast percentage for patient BET001 over the clinical trial treatment course (top panel). t-SNE analysis of 7360 individual blast cells isolated from patient BET001 at baseline, remission and relapse (bottom panel). scRNA-seq and genomic DNA sampling points are highlighted on the schematic. c Schematic of treatment regime and bone marrow blast percentage for patient BET002 over the clinical trial treatment course (top panel). t-SNE analysis of 6349 single blast cells isolated from patient BET002 at baseline and relapse (bottom panel). scRNA-seq and genomic DNA sampling points are highlighted on the schematic. d Flow cytometry analysis of cells from patient BET002 at baseline and relapse identifies enrichment for LMPP-like LSCs at relapse based on CD34 + CD38-CD90-CD45RA + expression. Gating strategy is defined by boxes. e Expression analysis of selected LSC signature genes (defined in REF 15 ) in blast cells from patient BET00
- Conjugate
- Yellow dye
 Explore
Explore Validate
Validate Learn
Learn Flow cytometry
Flow cytometry