Antibody data
- Antibody Data
- Antigen structure
- References [23]
- Comments [0]
- Validations
- Flow cytometry [1]
- Other assay [7]
Submit
Validation data
Reference
Comment
Report error
- Product number
- 14-0458-80 - Provider product page

- Provider
- Invitrogen Antibodies
- Product name
- CD45RA Monoclonal Antibody (HI100), eBioscience™
- Antibody type
- Monoclonal
- Antigen
- Other
- Description
- Description: The HI100 monoclonal antibody reacts with human CD45RA, a 220 kDa molecule expressed by subpopulations of CD4+ peripheral T lymphocytes, CD8+ peripheral T lymphocytes, and B cells. The CD45RA+ T cell populations are mainly naive/virgin allowing the use of HI100 mAb as a phenotypic marker to discriminate T cell subsets. Applications Reported: The HI100 antibody has been reported for use in flow cytometric analysis, and immunohistochemical staining. Applications Tested: The HI100 antibody has been tested by flow cytometric analysis of normal human peripheral blood cells. This can be used at less than or equal to 0.5 µg per test. A test is defined as the amount (µg) of antibody that will stain a cell sample in a final volume of 100 µL. Cell number should be determined empirically but can range from 10^5 to 10^8 cells/test. It is recommended that the antibody be carefully titrated for optimal performance in the assay of interest. Purity: Greater than 90%, as determined by SDS-PAGE. Aggregation: Less than 10%, as determined by HPLC. Filtration: 0.2 µm post-manufacturing filtered.
- Reactivity
- Human
- Host
- Mouse
- Isotype
- IgG
- Antibody clone number
- HI100
- Vial size
- 25 µg
- Concentration
- 0.5 mg/mL
- Storage
- 4° C
Submitted references Differences in Maturation Status and Immune Phenotypes of Circulating Helios(+) and Helios(-) Tregs and Their Disrupted Correlations With Monocyte Subsets in Autoantibody-Positive T1D Individuals.
ZEB1 promotes pathogenic Th1 and Th17 cell differentiation in multiple sclerosis.
Phase I Study of Ficlatuzumab and Cetuximab in Cetuximab-Resistant, Recurrent/Metastatic Head and Neck Cancer.
MicroRNA‑155 inhibits the proliferation of CD8+ T cells via upregulating regulatory T cells in vitiligo.
Intracellular delivery of mRNA to human primary T cells with microfluidic vortex shedding.
Targeting enhancer switching overcomes non-genetic drug resistance in acute myeloid leukaemia.
TIGIT expressing CD4+T cells represent a tumor-supportive T cell subset in chronic lymphocytic leukemia.
Targeting a CAR to the TRAC locus with CRISPR/Cas9 enhances tumour rejection.
The distribution and function of human memory T cell subsets in lung cancer.
Brief Report: IFIH1 Mutation Causes Systemic Lupus Erythematosus With Selective IgA Deficiency.
Efficient gene disruption in cultured primary human endothelial cells by CRISPR/Cas9.
Defining CD4 T cell memory by the epigenetic landscape of CpG DNA methylation.
PD-1(+) CD8(+) T cells are exhausted in tumours and functional in draining lymph nodes of colorectal cancer patients.
Microbe-specific unconventional T cells induce human neutrophil differentiation into antigen cross-presenting cells.
Autosomal dominant immune dysregulation syndrome in humans with CTLA4 mutations.
Monocytes and γδ T cells control the acute-phase response to intravenous zoledronate: insights from a phase IV safety trial.
Helios+ and Helios- cells coexist within the natural FOXP3+ T regulatory cell subset in humans.
Novel serial positive enrichment technology enables clinical multiparameter cell sorting.
Neutralizing IL-6 reduces human arterial allograft rejection by allowing emergence of CD161+ CD4+ regulatory T cells.
IL-15 transpresentation promotes both human T-cell reconstitution and T-cell-dependent antibody responses in vivo.
Cytomegalovirus-infected human endothelial cells can stimulate allogeneic CD4+ memory T cells by releasing antigenic exosomes.
TCR-induced downregulation of protein tyrosine phosphatase PEST augments secondary T cell responses.
PILAR is a novel modulator of human T-cell expansion.
Zhang Y, Zhang J, Shi Y, Shen M, Lv H, Chen S, Feng Y, Chen H, Xu X, Yang T, Xu K
Frontiers in immunology 2021;12:628504
Frontiers in immunology 2021;12:628504
ZEB1 promotes pathogenic Th1 and Th17 cell differentiation in multiple sclerosis.
Qian Y, Arellano G, Ifergan I, Lin J, Snowden C, Kim T, Thomas JJ, Law C, Guan T, Balabanov RD, Kaech SM, Miller SD, Choi J
Cell reports 2021 Aug 24;36(8):109602
Cell reports 2021 Aug 24;36(8):109602
Phase I Study of Ficlatuzumab and Cetuximab in Cetuximab-Resistant, Recurrent/Metastatic Head and Neck Cancer.
Bauman JE, Ohr J, Gooding WE, Ferris RL, Duvvuri U, Kim S, Johnson JT, Soloff AC, Wallweber G, Winslow J, Gaither-Davis A, Grandis JR, Stabile LP
Cancers 2020 Jun 11;12(6)
Cancers 2020 Jun 11;12(6)
MicroRNA‑155 inhibits the proliferation of CD8+ T cells via upregulating regulatory T cells in vitiligo.
Lv M, Li Z, Liu J, Lin F, Zhang Q, Li Z, Wang Y, Wang K, Xu Y
Molecular medicine reports 2019 Oct;20(4):3617-3624
Molecular medicine reports 2019 Oct;20(4):3617-3624
Intracellular delivery of mRNA to human primary T cells with microfluidic vortex shedding.
Jarrell JA, Twite AA, Lau KHWJ, Kashani MN, Lievano AA, Acevedo J, Priest C, Nieva J, Gottlieb D, Pawell RS
Scientific reports 2019 Mar 1;9(1):3214
Scientific reports 2019 Mar 1;9(1):3214
Targeting enhancer switching overcomes non-genetic drug resistance in acute myeloid leukaemia.
Bell CC, Fennell KA, Chan YC, Rambow F, Yeung MM, Vassiliadis D, Lara L, Yeh P, Martelotto LG, Rogiers A, Kremer BE, Barbash O, Mohammad HP, Johanson TM, Burr ML, Dhar A, Karpinich N, Tian L, Tyler DS, MacPherson L, Shi J, Pinnawala N, Yew Fong C, Papenfuss AT, Grimmond SM, Dawson SJ, Allan RS, Kruger RG, Vakoc CR, Goode DL, Naik SH, Gilan O, Lam EYN, Marine JC, Prinjha RK, Dawson MA
Nature communications 2019 Jun 20;10(1):2723
Nature communications 2019 Jun 20;10(1):2723
TIGIT expressing CD4+T cells represent a tumor-supportive T cell subset in chronic lymphocytic leukemia.
Catakovic K, Gassner FJ, Ratswohl C, Zaborsky N, Rebhandl S, Schubert M, Steiner M, Gutjahr JC, Pleyer L, Egle A, Hartmann TN, Greil R, Geisberger R
Oncoimmunology 2017;7(1):e1371399
Oncoimmunology 2017;7(1):e1371399
Targeting a CAR to the TRAC locus with CRISPR/Cas9 enhances tumour rejection.
Eyquem J, Mansilla-Soto J, Giavridis T, van der Stegen SJ, Hamieh M, Cunanan KM, Odak A, Gönen M, Sadelain M
Nature 2017 Mar 2;543(7643):113-117
Nature 2017 Mar 2;543(7643):113-117
The distribution and function of human memory T cell subsets in lung cancer.
Sheng SY, Gu Y, Lu CG, Zou JY, Hong H, Wang R
Immunologic research 2017 Jun;65(3):639-650
Immunologic research 2017 Jun;65(3):639-650
Brief Report: IFIH1 Mutation Causes Systemic Lupus Erythematosus With Selective IgA Deficiency.
Van Eyck L, De Somer L, Pombal D, Bornschein S, Frans G, Humblet-Baron S, Moens L, de Zegher F, Bossuyt X, Wouters C, Liston A
Arthritis & rheumatology (Hoboken, N.J.) 2015 Jun;67(6):1592-7
Arthritis & rheumatology (Hoboken, N.J.) 2015 Jun;67(6):1592-7
Efficient gene disruption in cultured primary human endothelial cells by CRISPR/Cas9.
Abrahimi P, Chang WG, Kluger MS, Qyang Y, Tellides G, Saltzman WM, Pober JS
Circulation research 2015 Jul 3;117(2):121-8
Circulation research 2015 Jul 3;117(2):121-8
Defining CD4 T cell memory by the epigenetic landscape of CpG DNA methylation.
Komori HK, Hart T, LaMere SA, Chew PV, Salomon DR
Journal of immunology (Baltimore, Md. : 1950) 2015 Feb 15;194(4):1565-79
Journal of immunology (Baltimore, Md. : 1950) 2015 Feb 15;194(4):1565-79
PD-1(+) CD8(+) T cells are exhausted in tumours and functional in draining lymph nodes of colorectal cancer patients.
Wu X, Zhang H, Xing Q, Cui J, Li J, Li Y, Tan Y, Wang S
British journal of cancer 2014 Sep 23;111(7):1391-9
British journal of cancer 2014 Sep 23;111(7):1391-9
Microbe-specific unconventional T cells induce human neutrophil differentiation into antigen cross-presenting cells.
Davey MS, Morgan MP, Liuzzi AR, Tyler CJ, Khan MWA, Szakmany T, Hall JE, Moser B, Eberl M
Journal of immunology (Baltimore, Md. : 1950) 2014 Oct 1;193(7):3704-3716
Journal of immunology (Baltimore, Md. : 1950) 2014 Oct 1;193(7):3704-3716
Autosomal dominant immune dysregulation syndrome in humans with CTLA4 mutations.
Schubert D, Bode C, Kenefeck R, Hou TZ, Wing JB, Kennedy A, Bulashevska A, Petersen BS, Schäffer AA, Grüning BA, Unger S, Frede N, Baumann U, Witte T, Schmidt RE, Dueckers G, Niehues T, Seneviratne S, Kanariou M, Speckmann C, Ehl S, Rensing-Ehl A, Warnatz K, Rakhmanov M, Thimme R, Hasselblatt P, Emmerich F, Cathomen T, Backofen R, Fisch P, Seidl M, May A, Schmitt-Graeff A, Ikemizu S, Salzer U, Franke A, Sakaguchi S, Walker LSK, Sansom DM, Grimbacher B
Nature medicine 2014 Dec;20(12):1410-1416
Nature medicine 2014 Dec;20(12):1410-1416
Monocytes and γδ T cells control the acute-phase response to intravenous zoledronate: insights from a phase IV safety trial.
Welton JL, Morgan MP, Martí S, Stone MD, Moser B, Sewell AK, Turton J, Eberl M
Journal of bone and mineral research : the official journal of the American Society for Bone and Mineral Research 2013 Mar;28(3):464-71
Journal of bone and mineral research : the official journal of the American Society for Bone and Mineral Research 2013 Mar;28(3):464-71
Helios+ and Helios- cells coexist within the natural FOXP3+ T regulatory cell subset in humans.
Himmel ME, MacDonald KG, Garcia RV, Steiner TS, Levings MK
Journal of immunology (Baltimore, Md. : 1950) 2013 Mar 1;190(5):2001-8
Journal of immunology (Baltimore, Md. : 1950) 2013 Mar 1;190(5):2001-8
Novel serial positive enrichment technology enables clinical multiparameter cell sorting.
Stemberger C, Dreher S, Tschulik C, Piossek C, Bet J, Yamamoto TN, Schiemann M, Neuenhahn M, Martin K, Schlapschy M, Skerra A, Schmidt T, Edinger M, Riddell SR, Germeroth L, Busch DH
PloS one 2012;7(4):e35798
PloS one 2012;7(4):e35798
Neutralizing IL-6 reduces human arterial allograft rejection by allowing emergence of CD161+ CD4+ regulatory T cells.
Fogal B, Yi T, Wang C, Rao DA, Lebastchi A, Kulkarni S, Tellides G, Pober JS
Journal of immunology (Baltimore, Md. : 1950) 2011 Dec 15;187(12):6268-80
Journal of immunology (Baltimore, Md. : 1950) 2011 Dec 15;187(12):6268-80
IL-15 transpresentation promotes both human T-cell reconstitution and T-cell-dependent antibody responses in vivo.
Huntington ND, Alves NL, Legrand N, Lim A, Strick-Marchand H, Mention JJ, Plet A, Weijer K, Jacques Y, Becker PD, Guzman C, Soussan P, Kremsdorf D, Spits H, Di Santo JP
Proceedings of the National Academy of Sciences of the United States of America 2011 Apr 12;108(15):6217-22
Proceedings of the National Academy of Sciences of the United States of America 2011 Apr 12;108(15):6217-22
Cytomegalovirus-infected human endothelial cells can stimulate allogeneic CD4+ memory T cells by releasing antigenic exosomes.
Walker JD, Maier CL, Pober JS
Journal of immunology (Baltimore, Md. : 1950) 2009 Feb 1;182(3):1548-59
Journal of immunology (Baltimore, Md. : 1950) 2009 Feb 1;182(3):1548-59
TCR-induced downregulation of protein tyrosine phosphatase PEST augments secondary T cell responses.
Arimura Y, Vang T, Tautz L, Williams S, Mustelin T
Molecular immunology 2008 Jun;45(11):3074-84
Molecular immunology 2008 Jun;45(11):3074-84
PILAR is a novel modulator of human T-cell expansion.
Huarte E, Cubillos-Ruiz JR, Nesbeth YC, Scarlett UK, Martinez DG, Engle XA, Rigby WF, Pioli PA, Guyre PM, Conejo-Garcia JR
Blood 2008 Aug 15;112(4):1259-68
Blood 2008 Aug 15;112(4):1259-68
No comments: Submit comment
Supportive validation
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
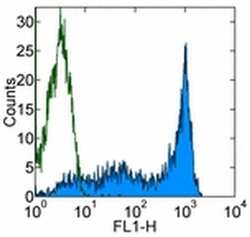
- Experimental details
- Staining of normal human peripheral blood cells with 0.25 µg of Mouse IgG2b Isotype Control Purified (Product # 14-4732-82) (open histogram) or 0.25 µg of Anti-Human CD45RA Purified (filled histogram) followed by Anti-Mouse IgG FITC (Product # 11-4011-85). Cells in the lymphocyte gate were used for analysis.
Supportive validation
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
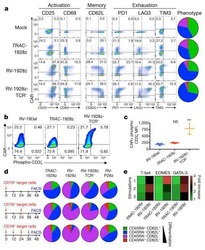
- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
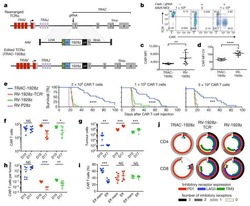
- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
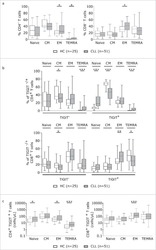
- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- Figure 6 T cell activation marker expression is unaffected in cells processed via uVS . Flow cytometry was used to quantify the surface level expression of key T cell activation markers (CD69, CD154, CD44, CCR7, CD45RA and CD25) in T cells processed with uVS (processed cells, red dashed) compared to non-processed cells (handling control, blue solid) 24 hours post transfection. Histogram overlays represent expression levels as a percent of maximum value of each marker for a representative sample for each condition. Experiment was performed in triplicate.
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
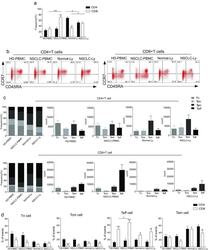
- Fig. 1 The distribution of CD4+ and CD8+ T cells subsets in human lung cancer. PBMCs were isolated from the blood of lung cancer patients and healthy donors and analyzed by flow cytometry. a The frequency of the CD3+CD4+ T cells and CD3+CD8+ T cells in the HD-PBMC, PBMCs from healthy donors; NSCLC-PBMC, PBMCs from non-small lung cancer patients, Normal-Ly, from healthy lymph node, NSCLC-Ly, tumor infiltrated lymph node from non-small lung cancer patients. b Representative flow cytometric analyses of CD45RA and CCR7 expression in CD3+CD4+ T cells and CD3+CD8+ T cells, indicating naive T cells (CD45RA+/CD45RO-CCR7+, top right quadrant ), terminal effector T cells (CD45RA+/CD45RO-CCR7-, bottom right quadrant ), central memory T cells (Tcm, CD45RO+/CD45RA-CCR7+, top left quadrant ), and effector memory T cells (Tem, CD45RO+/CD45RA-CCR7-, bottom left quadrant ), gated on the forward and side scatter of the lymphocyte populations. c The frequency and absolute number of the CD4+ ( top ) and CD8+ ( bottom ), Tn ( middle gray ), Teff ( black ), Tcm ( grey ), and Tem ( dark grey ) cell subsets in the blood from the non small cell lung cancer patients and healthy donors. d The events of Tn, Teff, Tcm and Tem cell subsets of CD4+ and CD8+ cells in the blood from non small cell lung cancer patients and healthy donors, expressed as the mean +- SEM. * p < 0.05; ** p < 0.005; *** p < 0.001; Mann-Whitney test (two-tailed) and non-paired Student's t-test
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
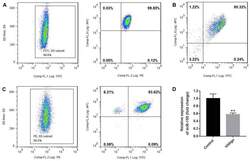
- Figure 1. Purity of CD3 + CD4 + CD45RA + T cells, CD3 + CD8 + T cells and CD4 + CD25 + FoxP3 + Treg cells. CD3 + CD4 + CD45RA + T cells and CD3 + CD8 + T cells were purified by magnetic cell sorting, and their purity was determined by flow cytometry. (A) The purity of CD3 + CD4 + CD45RA + T cells was 99.45% (CD3 + T cells, 99.6%; CD4 + CD45RA + T cells, 99.85%). (B) The purity of CD3 + CD8 + T cells was 95.32%. (C) The purity of CD4 + CD25 + FoxP3 + Treg cells was 93.15% (CD4 + T cells, 99.5%; CD25 + FoxP3 + T cells, 93.62%). (D) miR-155 expression in T cells of the patients with vitiligo and healthy donor was detected by reverse transcription quantitative polymerase chain reaction. **P
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
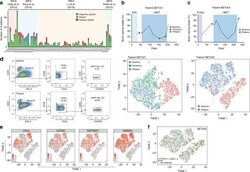
- Fig. 1 Non-genetic adaptation drives clinical resistance in AML. a Meta-analysis from four independent studies analysing either the whole genome or whole exome of AML patients at diagnosis and relapse. Mutations are defined as non-synonymous changes within the coding sequence of any gene. Shared mutations are mutations present at both diagnosis and relapse. Whole exome sequencing data from Li et al. (REF 4 ) was analysed to access the mutations in known AML genes, as defined by the authors. b Schematic of the treatment regime and bone marrow blast percentage for patient BET001 over the clinical trial treatment course (top panel). t-SNE analysis of 7360 individual blast cells isolated from patient BET001 at baseline, remission and relapse (bottom panel). scRNA-seq and genomic DNA sampling points are highlighted on the schematic. c Schematic of treatment regime and bone marrow blast percentage for patient BET002 over the clinical trial treatment course (top panel). t-SNE analysis of 6349 single blast cells isolated from patient BET002 at baseline and relapse (bottom panel). scRNA-seq and genomic DNA sampling points are highlighted on the schematic. d Flow cytometry analysis of cells from patient BET002 at baseline and relapse identifies enrichment for LMPP-like LSCs at relapse based on CD34 + CD38-CD90-CD45RA + expression. Gating strategy is defined by boxes. e Expression analysis of selected LSC signature genes (defined in REF 15 ) in blast cells from patient BET00
 Explore
Explore Validate
Validate Learn
Learn Immunohistochemistry
Immunohistochemistry Flow cytometry
Flow cytometry