Antibody data
- Antibody Data
- Antigen structure
- References [3]
- Comments [0]
- Validations
- Flow cytometry [1]
- Other assay [3]
Submit
Validation data
Reference
Comment
Report error
- Product number
- 47-0457-41 - Provider product page

- Provider
- Invitrogen Antibodies
- Product name
- CD45RO Monoclonal Antibody (UCHL1), APC-eFluor™ 780, eBioscience™
- Antibody type
- Monoclonal
- Antigen
- Other
- Description
- Description: The UCHL1 monoclonal antibody reacts with human CD45RO, a 180 kDa isoform of CD45. CD45RO is expressed by most thymocytes, activated memory T cells, granulocytes and monocytes. CD22 is a ligand for CD45RO. Expression of CD45RO and CD45RA is used commonly to discriminate subsets of peripheral T cells. Applications Reported: This UCHL1 antibody has been reported for use in flow cytometric analysis. Applications Tested: This UCHL1 antibody has been pre-titrated and tested by flow cytometric analysis of normal human peripheral blood cells. This can be used at 5 µL (0.5 µg) per test. A test is defined as the amount (µg) of antibody that will stain a cell sample in a final volume of 100 µL. Cell number should be determined empirically but can range from 10^5 to 10^8 cells/test. APC-eFluor® 780 emits at 780 nm and is excited with the Red laser (633 nm). Please make sure that your instrument is capable of detecting this fluorochome. Light sensitivity: This tandem dye is sensitive to photo-induced oxidation. Please protect this vial and stained samples from light. Fixation: Samples can be stored in IC Fixation Buffer (Product # 00-8222) (100 µL of cell sample + 100 µL of IC Fixation Buffer) or 1-step Fix/Lyse Solution (Product # 00-5333) for up to 3 days in the dark at 4°C with minimal impact on brightness and FRET efficiency/compensation. Some generalizations regarding fluorophore performance after fixation can be made, but clone specific performance should be determined empirically. Excitation: 633-647 nm; Emission: 780 nm; Laser: Red Laser. Filtration: 0.2 µm post-manufacturing filtered.
- Reactivity
- Human
- Host
- Mouse
- Isotype
- IgG
- Antibody clone number
- UCHL1
- Vial size
- 25 Tests
- Concentration
- 5 µL/Test
- Storage
- 4° C, store in dark, DO NOT FREEZE!
Submitted references Longitudinal analysis of subtype C envelope tropism for memory CD4(+) T cell subsets over the first 3 years of untreated HIV-1 infection.
Blocking the recruitment of naive CD4(+) T cells reverses immunosuppression in breast cancer.
Functional expression of CD137 (4-1BB) on T helper follicular cells.
Gartner MJ, Gorry PR, Tumpach C, Zhou J, Dantanarayana A, Chang JJ, Angelovich TA, Ellenberg P, Laumaea AE, Nonyane M, Moore PL, Lewin SR, Churchill MJ, Flynn JK, Roche M
Retrovirology 2020 Aug 6;17(1):24
Retrovirology 2020 Aug 6;17(1):24
Blocking the recruitment of naive CD4(+) T cells reverses immunosuppression in breast cancer.
Su S, Liao J, Liu J, Huang D, He C, Chen F, Yang L, Wu W, Chen J, Lin L, Zeng Y, Ouyang N, Cui X, Yao H, Su F, Huang JD, Lieberman J, Liu Q, Song E
Cell research 2017 Apr;27(4):461-482
Cell research 2017 Apr;27(4):461-482
Functional expression of CD137 (4-1BB) on T helper follicular cells.
Alfaro C, Echeveste JI, Rodriguez-Ruiz ME, Solorzano JL, Perez-Gracia JL, Idoate MA, Lopez-Picazo JM, Sanchez-Paulete AR, Labiano S, Rouzaut A, Oñate C, Aznar A, Lozano MD, Melero I
Oncoimmunology 2015 Dec;4(12):e1054597
Oncoimmunology 2015 Dec;4(12):e1054597
No comments: Submit comment
Supportive validation
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- Staining of normal human peripheral blood cells with Anti-Human CD45RA FITC (Product # 11-0458-42) and Mouse IgG2a K Isotype Control APC-eFluor® 780 (Product # 47-4724) (left) or Anti-Human CD45RO APC-eFluor® 780 (right). Cells in the lymphocyte gate were used for analysis.
Supportive validation
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
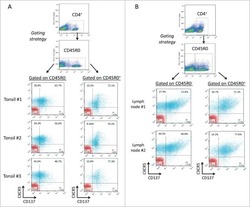
- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
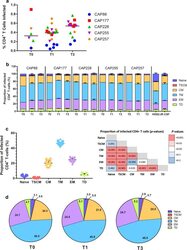
- Fig. 3 Transitional memory and effector memory cells were most frequently infected by C-HIV Envs. a Each data point represents the percentage of infected CD4 + T cells with one pseudovirus (averaged from four independent seronegative blood bank donors). The Env donor is indicated as follows; CAP88 (blue circles), CAP177 (red squares), CAP228 (green triangles), CAP255 (purple inverted triangles) and CAP257 (orange diamonds). Black lines represents the median of all pseudoviruses within each time point. Comparisons were made using a Kruskal-Wallis test with Dunn's post hoc test for multiple comparisons. b Stacked bar graphs represent the contribution of each T cell subset to the pool of infected CD4 + T cells. Values represent the median percentage of infected CD4 + T-cells (averaged across four HIV-seronegative PBMC donors) that belong to the indicated subset [naive; dark blue, T stem cell memory (TSCM); red, central memory (CM); yellow, transitional memory (TM); light blue, effector memory (EM); purple and terminally differentiated (TD); green], and are stratified by participant and time point. Error bars represent the interquartile range. c Dot plot representing the proportion of each T cell subset contributing to the total pool of infected cells for all Env-pseudoviruses. Each point represents a single virus averaged across four seronegative donors, lines represent median and error bars represent interquartile range. Comparisons were made using a Kruskal-Wallis test with Du
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
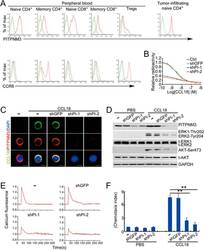
- Figure 5 PITPNM3 is a CCL18 receptor on naive CD4 + T cells. (A) Representative flow cytometry staining for PITPNM3 and CCR8, potential CCL18 receptors, on gated PB T cell subsets and paired TI naive CD4 + T cells of a breast cancer patient. Cells were gated on CD3 + CD45RA + CD45RO - CD25 - CD4 + /CD8 + for naive CD4 + /CD8 + T cells, CD3 + CD45RA - CD45RO + CD25 - CD4 + /CD8 + for memory CD4 + /CD8 + T cells and CD3 + CD4 + CD25 + for Tregs). Quantitation of PITPNM3 and CCR8 expression on T cell subsets for eight breast cancer patients is provided in Supplementary information, Figure S8A . (B-F) Knockdown of PITPNM3 in naive CD4 + T cells inhibits CCL18 binding, signaling and chemotaxis. (B) Binding of 125 I-CCL18 to naive CD4 + T cells, knocked down or not for PITPNM3 (shPI-1,2) in the presence of increasing concentrations of unlabeled CCL18. Shown are the representative assays for three independent experiments using PB T cells from three normal donors. (C) Representative fluorescence microscopy images of CCL18 binding to naive CD4 + T cells, knocked down or not for PITPNM3 , stained for PITPNM3 and CCL18 3 h after adding CCL18. Scale bar, 5 mum. Shown are the representative images for three independent experiments using PB T cells from three normal donors. (D) Immunoblot of CCL18-treated naive CD4 + T cells, knocked down or not for PITPNM3 , showing expression of PITPNM3 and phosphorylated/total (t-) Erk1/2 and Akt, relative to GAPDH as a loading control. Blots are repres
 Explore
Explore Validate
Validate Learn
Learn Flow cytometry
Flow cytometry